

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137077.1 - phase: 0
(1555 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
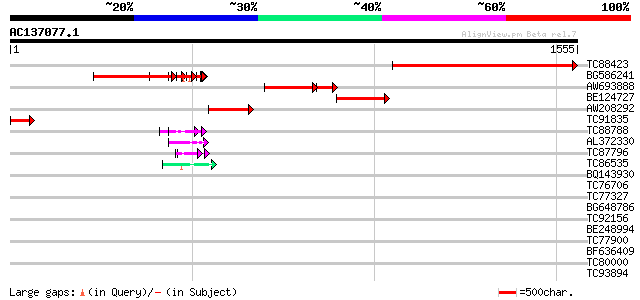
Score E
Sequences producing significant alignments: (bits) Value
TC88423 similar to GP|9758777|dbj|BAB09075.1 contains similarity... 1010 0.0
BG586241 similar to GP|9758776|dbj dbj|BAA90625.1~gene_id:MNJ7.7... 477 e-164
AW693888 weakly similar to GP|9758777|db contains similarity to ... 285 e-104
BE124727 weakly similar to GP|9758777|db contains similarity to ... 293 5e-79
AW208292 257 2e-68
TC91835 similar to GP|9758776|dbj|BAB09074.1 dbj|BAA90625.1~gene... 140 5e-33
TC88788 weakly similar to SP|P05790|FBOH_BOMMO Fibroin heavy cha... 62 2e-09
AL372330 similar to GP|21322752|dbj cold shock protein-1 {Tritic... 45 2e-04
TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear prot... 44 5e-04
TC86535 similar to GP|9758076|dbj|BAB08520.1 contains similarity... 43 8e-04
BQ143930 43 0.001
TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 42 0.001
TC77327 homologue to GP|16224254|gb|AAL15651.1 dehydrin-like pro... 42 0.002
BG648786 weakly similar to SP|P10495|GRP1_ Glycine-rich cell wal... 40 0.005
TC92156 similar to GP|15724296|gb|AAL06541.1 AT5g61660/k11j9_180... 40 0.007
BE248994 similar to GP|16604364|gb| AT5g39570/MIJ24_40 {Arabidop... 40 0.009
TC77900 similar to GP|9759182|dbj|BAB09797.1 ATP-dependent Clp p... 39 0.021
BF636409 37 0.046
TC80000 weakly similar to GP|15451136|gb|AAK96839.1 periaxin-lik... 37 0.060
TC93894 37 0.060
>TC88423 similar to GP|9758777|dbj|BAB09075.1 contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (15%)
Length = 1640
Score = 1010 bits (2612), Expect = 0.0
Identities = 505/505 (100%), Positives = 505/505 (100%)
Frame = +1
Query: 1051 AIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKTGRA 1110
AIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKTGRA
Sbjct: 1 AIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKTGRA 180
Query: 1111 PEVETWKQLVLALEERIRTHQQGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTS 1170
PEVETWKQLVLALEERIRTHQQGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTS
Sbjct: 181 PEVETWKQLVLALEERIRTHQQGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTS 360
Query: 1171 SQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSE 1230
SQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSE
Sbjct: 361 SQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSE 540
Query: 1231 PDIGRSPRQESPSPDAQGKVQVSGGASRFSRFGFGSQLLQKTVGLVLRSGKQAKLGEKNK 1290
PDIGRSPRQESPSPDAQGKVQVSGGASRFSRFGFGSQLLQKTVGLVLRSGKQAKLGEKNK
Sbjct: 541 PDIGRSPRQESPSPDAQGKVQVSGGASRFSRFGFGSQLLQKTVGLVLRSGKQAKLGEKNK 720
Query: 1291 FYYDEKLKRWVEEGAEVPAEEAALPPPPPTTAAFQNGSADYNLKSALKTEGLTPNEFSST 1350
FYYDEKLKRWVEEGAEVPAEEAALPPPPPTTAAFQNGSADYNLKSALKTEGLTPNEFSST
Sbjct: 721 FYYDEKLKRWVEEGAEVPAEEAALPPPPPTTAAFQNGSADYNLKSALKTEGLTPNEFSST 900
Query: 1351 RTSSPELSPGMPPIPPSSNQFSARSRLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKPAL 1410
RTSSPELSPGMPPIPPSSNQFSARSRLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKPAL
Sbjct: 901 RTSSPELSPGMPPIPPSSNQFSARSRLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKPAL 1080
Query: 1411 PANAKFFIPAPVPSSSEQNMEAIAESNLEDSAANENPSTSSTNDWSYHPPKHAQTMTMQR 1470
PANAKFFIPAPVPSSSEQNMEAIAESNLEDSAANENPSTSSTNDWSYHPPKHAQTMTMQR
Sbjct: 1081 PANAKFFIPAPVPSSSEQNMEAIAESNLEDSAANENPSTSSTNDWSYHPPKHAQTMTMQR 1260
Query: 1471 FPSAGNISKQGQTDGNESHFSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFM 1530
FPSAGNISKQGQTDGNESHFSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFM
Sbjct: 1261 FPSAGNISKQGQTDGNESHFSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFM 1440
Query: 1531 PDPSSLMQGPTRSGSFGEDLQEVEL 1555
PDPSSLMQGPTRSGSFGEDLQEVEL
Sbjct: 1441 PDPSSLMQGPTRSGSFGEDLQEVEL 1515
>BG586241 similar to GP|9758776|dbj dbj|BAA90625.1~gene_id:MNJ7.7~similar to
unknown protein {Arabidopsis thaliana}, partial (6%)
Length = 852
Score = 477 bits (1228), Expect(2) = e-164
Identities = 227/227 (100%), Positives = 227/227 (100%)
Frame = +3
Query: 230 QYQGVQGYDTSFDNHTGKQGDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSSSQNWED 289
QYQGVQGYDTSFDNHTGKQGDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSSSQNWED
Sbjct: 3 QYQGVQGYDTSFDNHTGKQGDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSSSQNWED 182
Query: 290 LYPGWKYDHITGQWYQIEDYNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQSVAG 349
LYPGWKYDHITGQWYQIEDYNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQSVAG
Sbjct: 183 LYPGWKYDHITGQWYQIEDYNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQSVAG 362
Query: 350 TLAETGTTESVSSWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSV 409
TLAETGTTESVSSWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSV
Sbjct: 363 TLAETGTTESVSSWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSV 542
Query: 410 HGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSG 456
HGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSG
Sbjct: 543 HGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSG 683
Score = 121 bits (303), Expect(2) = e-164
Identities = 55/55 (100%), Positives = 55/55 (100%)
Frame = +2
Query: 458 HGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 512
HGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH
Sbjct: 686 HGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 850
Score = 114 bits (285), Expect = 3e-25
Identities = 52/55 (94%), Positives = 53/55 (95%)
Frame = +2
Query: 485 HGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 539
HGV+QAGNY GS GVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH
Sbjct: 686 HGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 850
Score = 92.4 bits (228), Expect = 1e-18
Identities = 45/52 (86%), Positives = 47/52 (89%), Gaps = 1/52 (1%)
Frame = +2
Query: 435 SQAGNHV-SQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSH 485
SQAGN+ S GVGSQAVNGSWSGSHGV+QAGNY GS GVGSQAVNGSWSGSH
Sbjct: 695 SQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 850
Score = 63.9 bits (154), Expect = 5e-10
Identities = 28/31 (90%), Positives = 29/31 (93%)
Frame = +2
Query: 512 HGVNQAGNYGGSQGVGSQAVNGSWSGSHGVN 542
HGV+QAGNY GS GVGSQAVNGSWSGSHGVN
Sbjct: 686 HGVSQAGNYDGSHGVGSQAVNGSWSGSHGVN 778
Score = 59.7 bits (143), Expect = 9e-09
Identities = 50/169 (29%), Positives = 68/169 (39%), Gaps = 14/169 (8%)
Frame = +3
Query: 383 YPDWYYDTIAQEWRSLATYNSSVQSSVHGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVS 442
YP W YD I +W + YN++ S N T+ ++ +D + S QA V+
Sbjct: 186 YPGWKYDHITGQWYQIEDYNATTTSQQTSEAN--TAVDWAAASDGKTEISYLQQAAQSVA 359
Query: 443 QGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGS--------QAVNGSWSGSHGVNQA---- 490
+ S S + VSQ N H V + W N +
Sbjct: 360 GTLAETGTTESVSSWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQSS 539
Query: 491 --GNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSG 537
G G + + N S +QAGN+ SQGVGSQAVNGSWSG
Sbjct: 540 VHGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHV-SQGVGSQAVNGSWSG 683
>AW693888 weakly similar to GP|9758777|db contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (7%)
Length = 611
Score = 285 bits (728), Expect(2) = e-104
Identities = 144/144 (100%), Positives = 144/144 (100%)
Frame = +1
Query: 700 GKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQSF 759
GKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQSF
Sbjct: 1 GKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQSF 180
Query: 760 PGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFG 819
PGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFG
Sbjct: 181 PGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFG 360
Query: 820 TDTILKENDAPESAVAKLFASAKV 843
TDTILKENDAPESAVAKLFASAKV
Sbjct: 361 TDTILKENDAPESAVAKLFASAKV 432
Score = 114 bits (285), Expect(2) = e-104
Identities = 54/56 (96%), Positives = 55/56 (97%)
Frame = +2
Query: 842 KVNGTEFTQYGMPSHCLQNFPSEEQMKAIASEMQNLLVSGKKMEALQRAQEGQLWG 897
+ NGTEFTQYGMPSHCLQNFPSEEQMKAIASEMQNLLVSGKKMEALQRAQEGQLWG
Sbjct: 428 RCNGTEFTQYGMPSHCLQNFPSEEQMKAIASEMQNLLVSGKKMEALQRAQEGQLWG 595
>BE124727 weakly similar to GP|9758777|db contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (10%)
Length = 439
Score = 293 bits (749), Expect = 5e-79
Identities = 143/145 (98%), Positives = 144/145 (98%)
Frame = +3
Query: 897 GPALVLASQLGEQFYVDTVRQMALRQLVAGSPLRTLCLLIAGRPNDVFPTEETSISGHPG 956
GPALVLASQLGEQFYVDTVRQMALRQ+VAGSPLRTLCLLIAGRPNDVFPTEETSISGHPG
Sbjct: 3 GPALVLASQLGEQFYVDTVRQMALRQVVAGSPLRTLCLLIAGRPNDVFPTEETSISGHPG 182
Query: 957 AVGMPQQSEQAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKREITAAHICY 1016
AV MPQQSEQAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKREITAAHICY
Sbjct: 183 AVRMPQQSEQAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKREITAAHICY 362
Query: 1017 LIAEVNFSSYSDATRLCLIGADHWT 1041
LIAEVNFSSYSDATRLCLIGADHWT
Sbjct: 363 LIAEVNFSSYSDATRLCLIGADHWT 437
>AW208292
Length = 373
Score = 257 bits (657), Expect = 2e-68
Identities = 122/123 (99%), Positives = 123/123 (99%)
Frame = +1
Query: 545 QGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSSFGSVALNNKGSFEPKAFVPH 604
+GFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSSFGSVALNNKGSFEPKAFVPH
Sbjct: 4 RGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSSFGSVALNNKGSFEPKAFVPH 183
Query: 605 RDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQFSNVFAENQNSHSY 664
RDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQFSNVFAENQNSHSY
Sbjct: 184 RDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQFSNVFAENQNSHSY 363
Query: 665 SQQ 667
SQQ
Sbjct: 364 SQQ 372
>TC91835 similar to GP|9758776|dbj|BAB09074.1
dbj|BAA90625.1~gene_id:MNJ7.7~similar to unknown protein
{Arabidopsis thaliana}, partial (2%)
Length = 618
Score = 140 bits (352), Expect = 5e-33
Identities = 67/67 (100%), Positives = 67/67 (100%)
Frame = +2
Query: 1 MASNPPFQMEDQTDEDFFDKLVEDDDDVGPVKPVVIKSGNDEGNDSDDANAFANLSISDV 60
MASNPPFQMEDQTDEDFFDKLVEDDDDVGPVKPVVIKSGNDEGNDSDDANAFANLSISDV
Sbjct: 404 MASNPPFQMEDQTDEDFFDKLVEDDDDVGPVKPVVIKSGNDEGNDSDDANAFANLSISDV 583
Query: 61 DAGAFDN 67
DAGAFDN
Sbjct: 584 DAGAFDN 604
>TC88788 weakly similar to SP|P05790|FBOH_BOMMO Fibroin heavy chain
precursor (Fib-H) (H-fibroin). [Silk moth] {Bombyx
mori}, partial (2%)
Length = 782
Score = 62.0 bits (149), Expect = 2e-09
Identities = 43/105 (40%), Positives = 60/105 (56%)
Frame = +3
Query: 435 SQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYG 494
S AG G GS + +GS SGS G S AG+ GSH GS+A GS +GS ++AG+Y
Sbjct: 228 SGAGAGSGSGSGSASTSGSGSGS-GSSGAGSEAGSHA-GSRA--GSGAGSGAGSEAGSYA 395
Query: 495 GSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 539
GS + GS +GS + AG++ GS + + GS++GSH
Sbjct: 396 GSHAGSGSSGAGSEAGSEAGSYAGSHAGSGSSRAGSEAGSYAGSH 530
Score = 58.5 bits (140), Expect = 2e-08
Identities = 39/109 (35%), Positives = 58/109 (52%)
Frame = +3
Query: 411 GLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSH 470
G +G S STS + S+AG+H GS+A GS +GS S+AG+Y GSH
Sbjct: 243 GSGSGSGSASTSGSGSGSGSSGAGSEAGSHA----GSRA--GSGAGSGAGSEAGSYAGSH 404
Query: 471 GVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGN 519
+ GS +GS + AG++ GS + + GS++GSH + +GN
Sbjct: 405 AGSGSSGAGSEAGSEAGSYAGSHAGSGSSRAGSEAGSYAGSHAGSGSGN 551
>AL372330 similar to GP|21322752|dbj cold shock protein-1 {Triticum
aestivum}, partial (33%)
Length = 435
Score = 45.4 bits (106), Expect = 2e-04
Identities = 34/111 (30%), Positives = 46/111 (40%)
Frame = +2
Query: 435 SQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYG 494
S +G + S + S G +G +G Y GS G + G G +G +G YG
Sbjct: 38 SSSGGYGGSSSSSYPASSS-GGGYGGGSSGGYGGSSGGYGGSSGGYGGGGYGGGASGGYG 214
Query: 495 GSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNHQQ 545
G G G + G G +G G YGG G G G G+ G N Q+
Sbjct: 215 GG-GYGGGSGGGYGGGGYG---GGGYGGGGGGG---YGGDKMGALGANLQK 346
Score = 40.0 bits (92), Expect = 0.007
Identities = 33/90 (36%), Positives = 39/90 (42%), Gaps = 2/90 (2%)
Frame = +2
Query: 453 SWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 512
S SG +G S +G Y GS S + S SG G GGS G G +G + GS
Sbjct: 14 SSSGGYGGSSSGGYGGS---SSSSYPASSSGG------GYGGGSSG-GYGGSSGGYGGSS 163
Query: 513 GVNQAGNYGG--SQGVGSQAVNGSWSGSHG 540
G G YGG S G G G G +G
Sbjct: 164 GGYGGGGYGGGASGGYGGGGYGGGSGGGYG 253
Score = 39.3 bits (90), Expect = 0.012
Identities = 23/68 (33%), Positives = 31/68 (44%)
Frame = +2
Query: 480 SWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 539
S SG +G + +G YGGS A S G +G +G YGGS G + G G +
Sbjct: 14 SSSGGYGGSSSGGYGGSSSSSYPA--SSSGGGYGGGSSGGYGGSSGGYGGSSGGYGGGGY 187
Query: 540 GVNHQQGF 547
G G+
Sbjct: 188 GGGASGGY 211
>TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear protein.
[strain B95-8 Human herpesvirus 4] {Epstein-barr
virus}, partial (23%)
Length = 431
Score = 43.9 bits (102), Expect = 5e-04
Identities = 26/72 (36%), Positives = 31/72 (42%)
Frame = +1
Query: 456 GSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVN 515
G G G G+HGVG GS G+ G G+ GG G G G G+HG
Sbjct: 34 GGSGGGGGGGAGGAHGVG----YGSGGGTGGGYGGGSPGGGGGGGGSGGGGGGGGAHGGA 201
Query: 516 QAGNYGGSQGVG 527
G GG +G G
Sbjct: 202 YGGGIGGGEGGG 237
Score = 43.1 bits (100), Expect = 8e-04
Identities = 32/93 (34%), Positives = 38/93 (40%), Gaps = 3/93 (3%)
Frame = +1
Query: 459 GVSQAGNYDGSHGVGSQAVNGSWSGSHGV--NQAGNYGGSQGVGSQAVNGSWSGSHGVNQ 516
G G Y G G G G+ G+HGV G GG G GS G GS G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGA-GGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGG 177
Query: 517 AGN-YGGSQGVGSQAVNGSWSGSHGVNHQQGFD 548
G +GG+ G G + G G HG GF+
Sbjct: 178 GGGAHGGAYGGG---IGGGEGGGHGGYEP*GFN 267
>TC86535 similar to GP|9758076|dbj|BAB08520.1 contains similarity to RNA
binding protein~gene_id:MNF13.1 {Arabidopsis thaliana},
partial (48%)
Length = 1555
Score = 43.1 bits (100), Expect = 8e-04
Identities = 44/155 (28%), Positives = 59/155 (37%), Gaps = 8/155 (5%)
Frame = +1
Query: 420 STSSFNDDNSLYSE--YSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDG------SHG 471
S+ +ND S Y Y A + G G S SGS + G Y G +G
Sbjct: 793 SSKRYNDSRSSYGSGGYGDAYDGFGGGFGGVGGGYSRSGSAYGGRGGPYGGYGSEFAGYG 972
Query: 472 VGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAV 531
+ A+ G + G + AG YGGS S+ S +YG G G+ A
Sbjct: 973 GYAGAMGGPYRGDPSLAYAGRYGGS-------FRSSYDLSGYGGPGESYGAYGGAGAGAA 1131
Query: 532 NGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSG 566
G S +Q G+D AS G A+SG
Sbjct: 1132AGGGGSSGAGAYQSGYD-----ASLGGGYGGAASG 1221
Score = 37.0 bits (84), Expect = 0.060
Identities = 35/140 (25%), Positives = 50/140 (35%), Gaps = 8/140 (5%)
Frame = +1
Query: 415 GHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSH---- 470
G S S S++ Y Y G + A+ G + G ++ AG Y GS
Sbjct: 889 GGYSRSGSAYGGRGGPYGGYGS--EFAGYGGYAGAMGGPYRGDPSLAYAGRYGGSFRSSY 1062
Query: 471 ---GVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGS-QGV 526
G G + G G A GGS G G+ + + + G YGG+ G
Sbjct: 1063 DLSGYGGPGESYGAYGGAGAGAAAGGGGSSGAGA------YQSGYDASLGGGYGGAASGA 1224
Query: 527 GSQAVNGSWSGSHGVNHQQG 546
G + G+ G H G
Sbjct: 1225 SFYGSRGGYGGA-GRYHPYG 1281
>BQ143930
Length = 502
Score = 42.7 bits (99), Expect = 0.001
Identities = 36/127 (28%), Positives = 51/127 (39%)
Frame = +2
Query: 435 SQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYG 494
S +H Q S + +G G+ Q GN + GV SQ N G
Sbjct: 17 SSEEHHSKQQSSSSEEHHKKTGGSGLGQ-GNVGNAQGVASQG-----------NYGGQNT 160
Query: 495 GSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNHQQGFDMYATEA 554
GSQG+ +Q+ G G G GN GG+Q G+ S +G G Q + T
Sbjct: 161 GSQGIPNQSSTGGQGGLTGT--TGNVGGNQNYGNPGGATSTTGVTGGAGQPITEKTTTHE 334
Query: 555 STKIGNN 561
+T++ N
Sbjct: 335 TTRLPGN 355
>TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (94%)
Length = 1034
Score = 42.4 bits (98), Expect = 0.001
Identities = 31/107 (28%), Positives = 38/107 (34%)
Frame = +1
Query: 434 YSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNY 493
Y+ G H G NG G HG G G HG + ++ VN A
Sbjct: 322 YNGGGGHGGHG----GYNGG--GGHGGYNGGGGHGGHGAAESVAVQTEEKTNEVNDAKYG 483
Query: 494 GGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG 540
GGS G +G S +HG + GG G G G HG
Sbjct: 484 GGSYNHGGSYNHGGGSYNHGGGSYHHGGGGYNHGGGGHGGHGGGGHG 624
Score = 40.8 bits (94), Expect = 0.004
Identities = 33/122 (27%), Positives = 44/122 (36%), Gaps = 5/122 (4%)
Frame = +1
Query: 459 GVSQAGNYDG---SHGVGSQAVNGSW-SGSHGVNQAGN-YGGSQGVGSQAVNGSWSGSHG 513
G + G Y+G +HG G G + +G G N G Y G G G G G
Sbjct: 205 GYNHGGGYNGGGYNHGGGYHNGGGGYHNGGGGYNHGGGGYNGGGGHGGHGGYNGGGGHGG 384
Query: 514 VNQAGNYGGSQGVGSQAVNGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHS 573
N G +GG S AV + + + G Y S G + + G HH
Sbjct: 385 YNGGGGHGGHGAAESVAVQTEEKTNEVNDAKYGGGSYNHGGSYNHGGGSYNHGGGSYHHG 564
Query: 574 YG 575
G
Sbjct: 565 GG 570
Score = 40.4 bits (93), Expect = 0.005
Identities = 35/130 (26%), Positives = 48/130 (36%), Gaps = 10/130 (7%)
Frame = +1
Query: 421 TSSFNDDNSLYSEYSQAGNHVSQGVGSQAVN--GSWSGSHGVSQAGNYDGSHGVGSQAVN 478
T+ ND + Y Y+ G + G N G + G G +HG G
Sbjct: 160 TNEVND--AKYGGYNHGGYNHGGGYNGGGYNHGGGYHNGGGGYHNGGGGYNHGGGGYNGG 333
Query: 479 GSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSG--------SHGVNQAGNYGGSQGVGSQA 530
G G G N G +GG G G +G+ ++ VN A GGS G
Sbjct: 334 GGHGGHGGYNGGGGHGGYNGGGGHGGHGAAESVAVQTEEKTNEVNDAKYGGGSYNHGGSY 513
Query: 531 VNGSWSGSHG 540
+G S +HG
Sbjct: 514 NHGGGSYNHG 543
Score = 39.3 bits (90), Expect = 0.012
Identities = 36/123 (29%), Positives = 46/123 (37%), Gaps = 10/123 (8%)
Frame = +1
Query: 434 YSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNY 493
Y+ G + + G G NG +HG G Y+G G G G G N G +
Sbjct: 241 YNHGGGYHNGGGGYH--NGGGGYNHG---GGGYNGGGGHGGHGGYNGGGGHGGYNGGGGH 405
Query: 494 GGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG----------VNH 543
GG S AV ++ VN A GGS G +G S +HG NH
Sbjct: 406 GGHGAAESVAVQTE-EKTNEVNDAKYGGGSYNHGGSYNHGGGSYNHGGGSYHHGGGGYNH 582
Query: 544 QQG 546
G
Sbjct: 583 GGG 591
Score = 29.6 bits (65), Expect = 9.5
Identities = 18/59 (30%), Positives = 21/59 (35%)
Frame = +1
Query: 439 NHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQ 497
N G GS GS++ G G HG G G G HG G +G Q
Sbjct: 466 NDAKYGGGSYNHGGSYNHGGGSYNHGGGSYHHGGGGYNHGGGGHGGHGGGGHGGHGAEQ 642
>TC77327 homologue to GP|16224254|gb|AAL15651.1 dehydrin-like protein
{Medicago sativa}, partial (71%)
Length = 1330
Score = 41.6 bits (96), Expect = 0.002
Identities = 41/175 (23%), Positives = 58/175 (32%), Gaps = 20/175 (11%)
Frame = +2
Query: 415 GHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGS 474
G + + + + + Q GN +S G G G G H D + G GS
Sbjct: 122 GDQTRRVDEYGNPLTSQGQVDQYGNPISGG-GMTGATGHGHGHHQQHHGVGVDQTTGFGS 298
Query: 475 QAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAG----NYGGSQGVGSQA 530
G+ G+H + + GG G G+ GS + G YG G G+
Sbjct: 299 NTGTGTGYGTHTGSGGTHTGGVGGYGTTTEYGSTNTGSGYGNTDIGGTGYGTGTGTGTTG 478
Query: 531 V----NGSWSGSHGVNHQQ------------GFDMYATEASTKIGNNTASSGNQQ 569
G+ G G H G D A+ T G T G+QQ
Sbjct: 479 YGATGGGTGVGYGGTGHDNRGVMDKIKEKIPGTDQNASTYGTGTGYGTTGIGHQQ 643
Score = 29.6 bits (65), Expect = 9.5
Identities = 29/107 (27%), Positives = 43/107 (40%), Gaps = 12/107 (11%)
Frame = +2
Query: 501 SQAVNGSWSGSHGVNQAGNYGGSQG----VGSQAVNGSWSGS--HGVNHQQGFDMYATEA 554
SQ G + V++ GN SQG G+ G +G+ HG H Q +
Sbjct: 101 SQYQQGYGDQTRRVDEYGNPLTSQGQVDQYGNPISGGGMTGATGHGHGHHQQHHGVGVDQ 280
Query: 555 STKIGNNTASSGNQQVH------HSYGNQQVNTSSSFGSVALNNKGS 595
+T G+NT + H H+ G T++ +GS N GS
Sbjct: 281 TTGFGSNTGTGTGYGTHTGSGGTHTGGVGGYGTTTEYGS---TNTGS 412
>BG648786 weakly similar to SP|P10495|GRP1_ Glycine-rich cell wall structural
protein 1.0 precursor (GRP 1.0). [Kidney bean French
bean], partial (17%)
Length = 668
Score = 40.4 bits (93), Expect = 0.005
Identities = 45/189 (23%), Positives = 73/189 (37%), Gaps = 4/189 (2%)
Frame = +1
Query: 403 SSVQSSVHGLQNGHTSTSTSSFNDDNSL-YSEYSQAGNHVSQGVGSQAVNGSWSGSHGVS 461
++V G N ++ + NS Y G S GS V+G + G++
Sbjct: 109 NNVSYGAAGTANYGSAPDVGGYGGGNSANYGADVGGGESNSVNYGSAPVDGGYGGANNA- 285
Query: 462 QAGNYDGSHGVG-SQAVN-GSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGN 519
NY + G S +VN GS G +G + NY + + G +G N
Sbjct: 286 ---NYGAAVGARESNSVNYGSAPGGYGGGNSANYSDRESNSVNYGSAPGVGGYGSGNNAN 456
Query: 520 YGGSQGVG-SQAVNGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQ 578
YG + G G S +VN + G G+ A ++ +T G + GNQ
Sbjct: 457 YGAAVGGGESNSVNYGTAPVDG-----GYGAAAGFGNSDNHYSTGGVGGGPSNGFSGNQY 621
Query: 579 VNTSSSFGS 587
+++ +GS
Sbjct: 622 PGSATGYGS 648
>TC92156 similar to GP|15724296|gb|AAL06541.1 AT5g61660/k11j9_180
{Arabidopsis thaliana}, partial (67%)
Length = 650
Score = 40.0 bits (92), Expect = 0.007
Identities = 36/97 (37%), Positives = 44/97 (45%), Gaps = 5/97 (5%)
Frame = +2
Query: 449 AVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGS---HGVNQAGN-YGGSQGVGSQAV 504
A GS +G+ G N+D + G GS +G GS H + G YG G GS
Sbjct: 164 ASGGSPTGASGSGHGPNWDYNWGWGSAPGSGWGYGSGSGHSPSGFGRGYGYGFGTGS--- 334
Query: 505 NGSWSGS-HGVNQAGNYGGSQGVGSQAVNGSWSGSHG 540
GS SGS +G G +GG G GS S G HG
Sbjct: 335 -GSGSGSGYGYGSGGAHGGGYGSGS---GNSGGGGHG 433
Score = 37.0 bits (84), Expect = 0.060
Identities = 27/85 (31%), Positives = 35/85 (40%), Gaps = 1/85 (1%)
Frame = +2
Query: 444 GVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGS-HGVNQAGNYGGSQGVGSQ 502
G GS +G GS + +G G +GS SGS +G G +GG G GS
Sbjct: 230 GWGSAPGSGWGYGSGSGHSPSGFGRGYGYGFGTGSGSGSGSGYGYGSGGAHGGGYGSGS- 406
Query: 503 AVNGSWSGSHGVNQAGNYGGSQGVG 527
S G HG + N S+ G
Sbjct: 407 --GNSGGGGHGGGRTNNRSPSESKG 475
Score = 32.3 bits (72), Expect = 1.5
Identities = 23/70 (32%), Positives = 31/70 (43%), Gaps = 1/70 (1%)
Frame = +2
Query: 432 SEYSQAGNHVSQGVGSQAVNGSWSGS-HGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQA 490
S +S +G G G +GS SGS +G G + G +G GS S G HG +
Sbjct: 275 SGHSPSGFGRGYGYGFGTGSGSGSGSGYGYGSGGAHGGGYGSGS---GNSGGGGHGGGRT 445
Query: 491 GNYGGSQGVG 500
N S+ G
Sbjct: 446 NNRSPSESKG 475
Score = 31.2 bits (69), Expect = 3.3
Identities = 26/79 (32%), Positives = 31/79 (38%), Gaps = 5/79 (6%)
Frame = +2
Query: 473 GSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVN 532
G S SG HG N N+G GS GS SG YG G GS + +
Sbjct: 170 GGSPTGASGSG-HGPNWDYNWGWGSAPGSGWGYGSGSGHSPSGFGRGYGYGFGTGSGSGS 346
Query: 533 GSW-----SGSHGVNHQQG 546
GS G+HG + G
Sbjct: 347 GSGYGYGSGGAHGGGYGSG 403
>BE248994 similar to GP|16604364|gb| AT5g39570/MIJ24_40 {Arabidopsis
thaliana}, partial (20%)
Length = 579
Score = 39.7 bits (91), Expect = 0.009
Identities = 39/150 (26%), Positives = 60/150 (40%), Gaps = 9/150 (6%)
Frame = -2
Query: 438 GNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHG-VNQAGNYGGS 496
G G G Q + SG G Q G Y +G G A G + +G N+ YG
Sbjct: 575 GRKQGSGYGGQE-SEIGSGYGGRKQEGEYGSGYG-GGGAQEGEYGSGYGRKNEESEYGSG 402
Query: 497 QGVGSQAVNGSWSGSHGV--------NQAGNYGGSQGVGSQAVNGSWSGSHGVNHQQGFD 548
G G + +G SG G N+ YG G G + +G SG G +++ +
Sbjct: 401 YG-GRKQESGYGSGYGGAQESEYGRKNEESGYGSGYG-GRKQESGYGSGYGGRKNEEESE 228
Query: 549 MYATEASTKIGNNTASSGNQQVHHSYGNQQ 578
Y + + G + SG ++ YG ++
Sbjct: 227 GYGRKNEYESGESEYGSGRKK--SGYGEEE 144
Score = 33.5 bits (75), Expect = 0.66
Identities = 37/137 (27%), Positives = 53/137 (38%), Gaps = 13/137 (9%)
Frame = -2
Query: 464 GNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG-VNQAGNYGG 522
G GS G ++ GS G G Q G YG G G A G + +G N+ YG
Sbjct: 575 GRKQGSGYGGQESEIGSGYG--GRKQEGEYGSGYG-GGGAQEGEYGSGYGRKNEESEYGS 405
Query: 523 SQGVGSQAVNGSWSGSHGV--------NHQQGFDMYATEASTKIGNNTASSG--NQQVHH 572
G G + +G SG G N + G+ + G + G N++
Sbjct: 404 GYG-GRKQESGYGSGYGGAQESEYGRKNEESGYGSGYGGRKQESGYGSGYGGRKNEEESE 228
Query: 573 SYG--NQQVNTSSSFGS 587
YG N+ + S +GS
Sbjct: 227 GYGRKNEYESGESEYGS 177
>TC77900 similar to GP|9759182|dbj|BAB09797.1 ATP-dependent Clp protease
regulatory subunit CLPX {Arabidopsis thaliana}, partial
(61%)
Length = 2144
Score = 38.5 bits (88), Expect = 0.021
Identities = 33/102 (32%), Positives = 48/102 (46%), Gaps = 9/102 (8%)
Frame = +3
Query: 409 VHGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGS--------WSGSHGV 460
+H N +TST+++S +L + +Q NHV V + V S WSG V
Sbjct: 378 IHKSSNNNTSTASTS----TTLIEDSNQNPNHVPYDVTAYRVLVSSFVDPPEVWSGGGIV 545
Query: 461 SQAGNYDGSHGVGSQAVNG-SWSGSHGVNQAGNYGGSQGVGS 501
+ GNYD S G G G + SG G ++ +GGS G+
Sbjct: 546 VRPGNYDVSGGGGGGGGGGAASSGDSGNSKDWCWGGSNMGGT 671
>BF636409
Length = 648
Score = 37.4 bits (85), Expect = 0.046
Identities = 46/144 (31%), Positives = 65/144 (44%), Gaps = 25/144 (17%)
Frame = +3
Query: 427 DNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGS------------HGV-- 472
DN+L+ HV+ G+ + A GS GVS++ N + H +
Sbjct: 96 DNALF--------HVNSGINNTARGGSIGKFSGVSESNNLVDAMKFASSPTTFHPHSLPE 251
Query: 473 --GSQAVNGSWSGSHGV-NQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNY-----GGSQ 524
GS A ++ S + N+AGN G GV ++A NG HG++ GN GGS
Sbjct: 252 FHGSLANGSPYTFSSTISNKAGNIGA--GV-TEASNG--RHIHGISSVGNLAEFNGGGSS 416
Query: 525 GVGSQA---VNGSWSGSHGVNHQQ 545
G G A +N WSGS+ HQQ
Sbjct: 417 GNGINAHHGLNHIWSGSN--LHQQ 482
>TC80000 weakly similar to GP|15451136|gb|AAK96839.1 periaxin-like protein
{Arabidopsis thaliana}, partial (55%)
Length = 1033
Score = 37.0 bits (84), Expect = 0.060
Identities = 34/122 (27%), Positives = 48/122 (38%), Gaps = 8/122 (6%)
Frame = -2
Query: 482 SGSHGVNQAGNYG--GSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSH 539
SG+ G + GN G G G G + G+ G+ +G G S V S + G+
Sbjct: 990 SGTLGCSSLGNSGTLGRLGFGISGI----LGNSGLGNSGTLGNSGFVNSGVLGNCGLGNS 823
Query: 540 GVNHQQGF------DMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSSFGSVALNNK 593
G+ GF D + S +GN+ S GN SS FG+ + N
Sbjct: 822 GILGNSGFGNSETLDSLGSCNSGALGNSCF*SSGTLGTSDLGNSGALGSSGFGNSGILNL 643
Query: 594 GS 595
GS
Sbjct: 642 GS 637
>TC93894
Length = 521
Score = 37.0 bits (84), Expect = 0.060
Identities = 26/104 (25%), Positives = 48/104 (46%), Gaps = 2/104 (1%)
Frame = +1
Query: 91 DGGGDDDGKEGNLLMGSSSVECDSKTELGKEEIGIGSEFTAVAPVGKSNEVASSGLMDIA 150
+GGGDD +G + +S+ S E GK+ + GS +GK + +++
Sbjct: 43 NGGGDDIVVDGEAVDATSAPAMVSNVEAGKKSVVGGSRRKGAGNLGKLD-------LNVE 201
Query: 151 AVGESNEVALSGIKEKDWNCF--SADDANGVGGFGSYSDFFSEL 192
E E + SG+++ + N + D+ G+G F +F + L
Sbjct: 202 FSNEVEEASASGVRQGNVNGTGNAEDNIEGIGFFEGLDEFLNSL 333
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.128 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,091,684
Number of Sequences: 36976
Number of extensions: 622192
Number of successful extensions: 3649
Number of sequences better than 10.0: 107
Number of HSP's better than 10.0 without gapping: 3230
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3556
length of query: 1555
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1446
effective length of database: 4,984,343
effective search space: 7207359978
effective search space used: 7207359978
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC137077.1


