

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.7 + phase: 0
(123 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
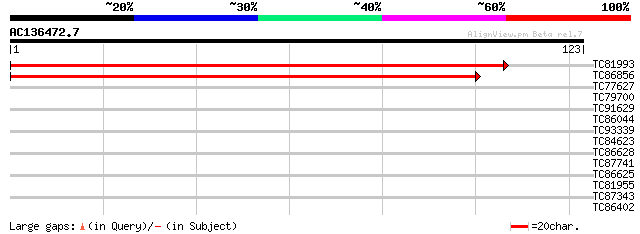
Sequences producing significant alignments: (bits) Value
TC81993 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase... 209 2e-55
TC86856 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase... 184 1e-47
TC77627 similar to GP|15028089|gb|AAK76575.1 unknown protein {Ar... 33 0.021
TC79700 similar to GP|16226834|gb|AAL16275.1 AT3g54190/F24B22_15... 27 2.0
TC91629 similar to PIR|A96608|A96608 hypothetical protein F25P12... 25 5.7
TC86044 similar to GP|4235430|gb|AAD13216.1| latex-abundant prot... 25 7.5
TC93339 similar to PIR|T08950|T08950 GTP binding protein Arac7 [... 25 7.5
TC84623 similar to GP|9759369|dbj|BAB09828.1 gene_id:MYJ24.10~re... 25 9.8
TC86628 similar to PIR|T08971|T08971 hypothetical protein F19B15... 25 9.8
TC87741 similar to GP|7269844|emb|CAB79703.1 serine/threonine-sp... 25 9.8
TC86625 similar to GP|16226834|gb|AAL16275.1 AT3g54190/F24B22_15... 25 9.8
TC81955 similar to SP|P93841|ISPE_LYCES 4-diphosphocytidyl-2-C-m... 25 9.8
TC87343 homologue to GP|16226834|gb|AAL16275.1 AT3g54190/F24B22_... 25 9.8
TC86402 homologue to GP|15148926|gb|AAK84890.1 TGA-type basic le... 25 9.8
>TC81993 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase
{Arabidopsis thaliana}, partial (51%)
Length = 1096
Score = 209 bits (533), Expect = 2e-55
Identities = 102/107 (95%), Positives = 103/107 (95%)
Frame = +3
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA
Sbjct: 96 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 275
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLR 107
LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG +R
Sbjct: 276 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGIEERMR 416
>TC86856 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase {Arabidopsis
thaliana}, partial (90%)
Length = 1911
Score = 184 bits (466), Expect = 1e-47
Identities = 86/101 (85%), Positives = 96/101 (94%)
Frame = +3
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MSIDAMEE+AKARKSGQRVIGEP++SGLALDDSWLWHPDF TAAKY+MSPPIR +GHDKA
Sbjct: 900 MSIDAMEEIAKARKSGQRVIGEPIVSGLALDDSWLWHPDFTTAAKYLMSPPIRSKGHDKA 1079
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
LQ+AL+TGVLQLVGTDHC +NSTQKA G+DDFR+I NGVNG
Sbjct: 1080 LQSALATGVLQLVGTDHCAWNSTQKARGVDDFRQIINGVNG 1202
>TC77627 similar to GP|15028089|gb|AAK76575.1 unknown protein {Arabidopsis
thaliana}, partial (88%)
Length = 1964
Score = 33.5 bits (75), Expect = 0.021
Identities = 22/63 (34%), Positives = 30/63 (46%)
Frame = +1
Query: 38 PDFDTAAKYVMSPPIRKQGHDKALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPN 97
PD DT +Y SPPIR + + L AL G + L+ +DH K DF K
Sbjct: 1012 PDKDT--RYKCSPPIRDASNREKLWEALLDGHIDLLSSDHSPTVPKLKLLKEGDFLKAWG 1185
Query: 98 GVN 100
G++
Sbjct: 1186 GIS 1194
>TC79700 similar to GP|16226834|gb|AAL16275.1 AT3g54190/F24B22_150
{Arabidopsis thaliana}, partial (79%)
Length = 1644
Score = 26.9 bits (58), Expect = 2.0
Identities = 21/87 (24%), Positives = 37/87 (42%), Gaps = 8/87 (9%)
Frame = +1
Query: 29 ALDDSWLWHPDFDTAAKYVMSPP------IRKQGHDKALQAALSTGVLQLVGTDHCV--F 80
+L+D LWHP +T Y+ S + + D+ ++A + + + T CV
Sbjct: 1099 SLEDHLLWHPHCNTNNIYITSDQDLIISYCKAESEDQWMEANAGSITVSNILTGKCVAKI 1278
Query: 81 NSTQKAFGIDDFRKIPNGVNGDHSNLR 107
N+ +DD + + D S LR
Sbjct: 1279 NAANCISKVDDCSSTCSCKHTDSSQLR 1359
>TC91629 similar to PIR|A96608|A96608 hypothetical protein F25P12.93
[imported] - Arabidopsis thaliana, partial (67%)
Length = 1291
Score = 25.4 bits (54), Expect = 5.7
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 1/57 (1%)
Frame = -1
Query: 35 LWHPDFDTAAKYVMSPPIRKQGHDKALQAALSTGVLQLVGTDHCV-FNSTQKAFGID 90
L HP F + K M+PP ++ Q +ST V ++ V N+T FG D
Sbjct: 460 LGHPPFSNSPKAFMNPPCIFPTFGRSYQ-EISTSVFHCTASNGVVPLNATTIKFGSD 293
>TC86044 similar to GP|4235430|gb|AAD13216.1| latex-abundant protein {Hevea
brasiliensis}, partial (92%)
Length = 1551
Score = 25.0 bits (53), Expect = 7.5
Identities = 15/48 (31%), Positives = 20/48 (41%)
Frame = +1
Query: 52 IRKQGHDKALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGV 99
+ GH L A TG G D C+ S DDF++ +GV
Sbjct: 310 VHYSGHGTRLPA--ETGEDDDTGFDECIVPSDMNLITDDDFKEFVDGV 447
>TC93339 similar to PIR|T08950|T08950 GTP binding protein Arac7 [imported]
- Arabidopsis thaliana, partial (32%)
Length = 467
Score = 25.0 bits (53), Expect = 7.5
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +1
Query: 40 FDTAAKYVMSPPIRKQ 55
FDTA K V+ PP RK+
Sbjct: 82 FDTAIKVVLQPPRRKE 129
>TC84623 similar to GP|9759369|dbj|BAB09828.1
gene_id:MYJ24.10~ref|NP_055178.1~similar to unknown
protein {Arabidopsis thaliana}, partial (3%)
Length = 634
Score = 24.6 bits (52), Expect = 9.8
Identities = 12/35 (34%), Positives = 17/35 (48%)
Frame = -2
Query: 43 AAKYVMSPPIRKQGHDKALQAALSTGVLQLVGTDH 77
++KY +P RKQ H K L+T + T H
Sbjct: 396 SSKY*QNPLQRKQHHYKPKAQVLATAITNRTATTH 292
>TC86628 similar to PIR|T08971|T08971 hypothetical protein F19B15.190 -
Arabidopsis thaliana, partial (88%)
Length = 743
Score = 24.6 bits (52), Expect = 9.8
Identities = 14/57 (24%), Positives = 26/57 (45%), Gaps = 3/57 (5%)
Frame = +3
Query: 57 HDKALQ---AALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLR 110
HD+ + A +T + + T + QKA IDD K + +N N++ ++
Sbjct: 390 HDQMIMLEGAKATTETVDALRTGAATMKAMQKATNIDDVDKTMDEINEQTENMKQIQ 560
>TC87741 similar to GP|7269844|emb|CAB79703.1 serine/threonine-specific
receptor protein kinase-like protein {Arabidopsis
thaliana}, partial (14%)
Length = 796
Score = 24.6 bits (52), Expect = 9.8
Identities = 17/90 (18%), Positives = 36/90 (39%), Gaps = 7/90 (7%)
Frame = +2
Query: 37 HPDFDTAAKYVMSPPIRKQGHDKALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIP 96
+ D DT KYV + + TGV + + +D+ + + + D R P
Sbjct: 164 YTDEDTKIKYVTDG------------SYIQTGVNKNISSDYAYPKNPNLPYPLSDLRSFP 307
Query: 97 NG-------VNGDHSNLRYLRPKYILDDFE 119
+G + G +L +R ++ +++
Sbjct: 308 HGNRNCYRLIAGTKGSLHLIRASFLYGNYD 397
>TC86625 similar to GP|16226834|gb|AAL16275.1 AT3g54190/F24B22_150
{Arabidopsis thaliana}, partial (80%)
Length = 1447
Score = 24.6 bits (52), Expect = 9.8
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = +1
Query: 29 ALDDSWLWHPDFDTAAKYVMS 49
+ +D LWHPD +T Y+ S
Sbjct: 1273 SFEDHLLWHPDCNTNNIYITS 1335
>TC81955 similar to SP|P93841|ISPE_LYCES
4-diphosphocytidyl-2-C-methyl-D-erythritol kinase
chloroplast precursor (EC 2.7.1.148) (CMK), partial
(77%)
Length = 1140
Score = 24.6 bits (52), Expect = 9.8
Identities = 14/39 (35%), Positives = 21/39 (52%), Gaps = 2/39 (5%)
Frame = +1
Query: 78 CVFNS--TQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYI 114
C NS T + + FR PNG + HS L + RP+++
Sbjct: 154 CSNNSIPTFRTIHLPQFR--PNGSSNFHSKLHFHRPQFV 264
>TC87343 homologue to GP|16226834|gb|AAL16275.1 AT3g54190/F24B22_150
{Arabidopsis thaliana}, partial (64%)
Length = 1328
Score = 24.6 bits (52), Expect = 9.8
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = +3
Query: 29 ALDDSWLWHPDFDTAAKYVMS 49
+ +D LWHPD +T Y+ S
Sbjct: 621 SFEDHLLWHPDCNTNNIYITS 683
>TC86402 homologue to GP|15148926|gb|AAK84890.1 TGA-type basic leucine
zipper protein TGA2.2 {Phaseolus vulgaris}, partial
(96%)
Length = 1970
Score = 24.6 bits (52), Expect = 9.8
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +1
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQ 84
LQ + T +LQL+G HC + T+
Sbjct: 391 LQTRI*TSLLQLLGLRHCNYKKTR 462
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,464,256
Number of Sequences: 36976
Number of extensions: 39614
Number of successful extensions: 167
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 167
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 167
length of query: 123
length of database: 9,014,727
effective HSP length: 84
effective length of query: 39
effective length of database: 5,908,743
effective search space: 230440977
effective search space used: 230440977
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC136472.7


