

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
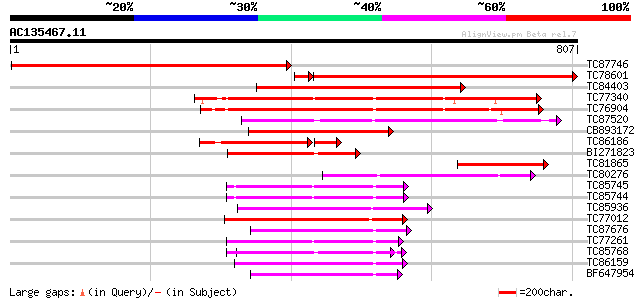
Query= AC135467.11 - phase: 0
(807 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87746 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease... 786 0.0
TC78601 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotea... 742 0.0
TC84403 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotea... 522 e-148
TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog ... 431 e-121
TC76904 homologue to GP|4325041|gb|AAD17230.1| FtsH-like protein... 399 e-111
TC87520 similar to GP|9757998|dbj|BAB08420.1 cell division prote... 330 1e-90
CB893172 homologue to PIR|T45642|T45 FtsH metalloproteinase-like... 234 1e-61
TC86186 similar to GP|4325041|gb|AAD17230.1| FtsH-like protein P... 179 5e-55
BI271823 similar to GP|21592745|gb cell division protein FtsH-li... 211 7e-55
TC81865 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease... 206 4e-53
TC80276 similar to GP|20197264|gb|AAC31223.2 FtsH protease puta... 205 6e-53
TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome re... 192 4e-49
TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15... 191 1e-48
TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidops... 187 1e-47
TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulator... 183 2e-46
TC87676 PIR|G96577|G96577 26S proteasome ATPase subunit [importe... 181 1e-45
TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AA... 177 1e-44
TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle pr... 176 3e-44
TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AA... 168 9e-42
BF647954 homologue to GP|11094192|db 26S proteasome regulatory p... 161 1e-39
>TC87746 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (49%)
Length = 1274
Score = 786 bits (2030), Expect = 0.0
Identities = 394/399 (98%), Positives = 395/399 (98%)
Frame = +2
Query: 3 FSRIGRSLSRSSRVKNLLHGETRLGTLYGVSRTNVFVDDVEKGLGFVRGYVSSAIARNNG 62
F +G SLSRSSRVKNLLHGETRLGTLYGVSRTNVFVDDVEKGLGFVRGYVSSAIARNNG
Sbjct: 77 FQELGGSLSRSSRVKNLLHGETRLGTLYGVSRTNVFVDDVEKGLGFVRGYVSSAIARNNG 256
Query: 63 FGSNLYDFKSIAANRMLHRMFSSESPKKKNYEKFYPKEKKEVPKGEEKKSESKDESKSNT 122
FGSNLYDFKSIAANRMLHRMFSSESPKKKNYEKFYPKEKKEVPKGEEKKSESKDESKSNT
Sbjct: 257 FGSNLYDFKSIAANRMLHRMFSSESPKKKNYEKFYPKEKKEVPKGEEKKSESKDESKSNT 436
Query: 123 EDGGSFHEAFIKQFQNYLTPLLVVGLFLSSLSLGPRDQQQISFQEFKNKLLEPGLVDHIV 182
EDGGSFHEAFIKQFQNYLTPLLVVGLFLSSLSLGPRDQQQISFQEFKNKLLEPGLVDHIV
Sbjct: 437 EDGGSFHEAFIKQFQNYLTPLLVVGLFLSSLSLGPRDQQQISFQEFKNKLLEPGLVDHIV 616
Query: 183 VSNKSVAKIYVRNSPLNQADSEVQGTLPAKGSGGQYKYIINIGSVESFEEKLEEAQEALG 242
VSNKSVAKIYVRNSPLNQADSEVQGTLPAKGSGGQYKYIINIGSVESFEEKLEEAQEALG
Sbjct: 617 VSNKSVAKIYVRNSPLNQADSEVQGTLPAKGSGGQYKYIINIGSVESFEEKLEEAQEALG 796
Query: 243 VDSHNFVPVTYSSEMVWYQELMRFAPTLLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIF 302
VDSHNFVPVTYSSEMVWYQELMRFAPTLLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIF
Sbjct: 797 VDSHNFVPVTYSSEMVWYQELMRFAPTLLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIF 976
Query: 303 NIGKAHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLV 362
NIGKAHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLV
Sbjct: 977 NIGKAHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLV 1156
Query: 363 GPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGP 401
GPPGTGKTLL KATAGESGVPFLSISGSDFMEMFVGVGP
Sbjct: 1157GPPGTGKTLLXKATAGESGVPFLSISGSDFMEMFVGVGP 1273
>TC78601 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (49%)
Length = 1338
Score = 742 bits (1916), Expect(2) = 0.0
Identities = 375/375 (100%), Positives = 375/375 (100%)
Frame = +1
Query: 433 RGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISID 492
RGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISID
Sbjct: 85 RGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISID 264
Query: 493 VPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDES 552
VPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDES
Sbjct: 265 VPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDES 444
Query: 553 QVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVP 612
QVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVP
Sbjct: 445 QVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVP 624
Query: 613 RGTAALGFAQYVPSENLLRTKEQLLDMTCMTLGGRAAEQVLIGAISTGAQNDLEKVTKMT 672
RGTAALGFAQYVPSENLLRTKEQLLDMTCMTLGGRAAEQVLIGAISTGAQNDLEKVTKMT
Sbjct: 625 RGTAALGFAQYVPSENLLRTKEQLLDMTCMTLGGRAAEQVLIGAISTGAQNDLEKVTKMT 804
Query: 673 YAQVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHK 732
YAQVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHK
Sbjct: 805 YAQVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHK 984
Query: 733 EKLAQIAELLLEKEVLHQEDLVRILGERPFKSAEPTNYDRFKLGFQDEEKAAETTVDEAE 792
EKLAQIAELLLEKEVLHQEDLVRILGERPFKSAEPTNYDRFKLGFQDEEKAAETTVDEAE
Sbjct: 985 EKLAQIAELLLEKEVLHQEDLVRILGERPFKSAEPTNYDRFKLGFQDEEKAAETTVDEAE 1164
Query: 793 EGSGSSPLEPEVVPT 807
EGSGSSPLEPEVVPT
Sbjct: 1165EGSGSSPLEPEVVPT 1209
Score = 57.8 bits (138), Expect(2) = 0.0
Identities = 27/27 (100%), Positives = 27/27 (100%)
Frame = +3
Query: 406 NLFQEARQCAPSIVFIDEIDAIGRKRG 432
NLFQEARQCAPSIVFIDEIDAIGRKRG
Sbjct: 3 NLFQEARQCAPSIVFIDEIDAIGRKRG 83
>TC84403 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (36%)
Length = 897
Score = 522 bits (1344), Expect = e-148
Identities = 263/298 (88%), Positives = 278/298 (93%)
Frame = +3
Query: 352 GAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEA 411
GAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDF+E+FVGVG SRVRNLF+EA
Sbjct: 3 GAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFLELFVGVGSSRVRNLFKEA 182
Query: 412 RQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNR 471
R+CAPSIVFIDEIDAIGR RG G +NDERESTLNQLLVEMDGFGTTAGVVVLAGTNR
Sbjct: 183 RKCAPSIVFIDEIDAIGRARGSRGGDRANDERESTLNQLLVEMDGFGTTAGVVVLAGTNR 362
Query: 472 ADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFA 531
D+LD ALLRPGRFDR I+ID PDIKGRDQIFQIYLK IKLDHEPSYYS +LA+LTPGFA
Sbjct: 363 PDVLDKALLRPGRFDRQITIDKPDIKGRDQIFQIYLKGIKLDHEPSYYSHKLASLTPGFA 542
Query: 532 GADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAG 591
GADIANVCNEAALIAART+E+ VT DHFEAAIDRIIGGLEKKNRVISK +RRTVAYHEAG
Sbjct: 543 GADIANVCNEAALIAARTEEAHVTHDHFEAAIDRIIGGLEKKNRVISKLQRRTVAYHEAG 722
Query: 592 HAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSENLLRTKEQLLDMTCMTLGGRAA 649
HAVAGWFLEH +PLLKVTIVPRGT+ALGFAQYVP+ENLL TKEQL D TCMTLGGRAA
Sbjct: 723 HAVAGWFLEHTDPLLKVTIVPRGTSALGFAQYVPNENLLMTKEQLFDRTCMTLGGRAA 896
>TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog chloroplast
precursor (EC 3.4.24.-). [Alfalfa] {Medicago sativa},
complete
Length = 2473
Score = 431 bits (1108), Expect = e-121
Identities = 238/512 (46%), Positives = 335/512 (64%), Gaps = 18/512 (3%)
Frame = +3
Query: 263 LMRFAPTLLL----LGTLWFMGRKMQGGFGVGGGSTGKGSRGIFNIGKAHVTKVDKNTKN 318
L F +LLL L+ + R+ QGG G GG G + G++ +K + +
Sbjct: 834 LFSFVGSLLLPFLAFAGLFLIFRRGQGGPGGPGGLGGP-----MDFGRSK-SKFQEVPET 995
Query: 319 KVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAG 378
V F DVAG ++AK E+ E V FLKNP KY LGAKIPKG LLVGPPGTGKTLLA+A AG
Sbjct: 996 GVTFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAG 1175
Query: 379 ESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSG 438
E+G PF S + S+F+E+FVGVG SRVR+LF++A+ AP IVFIDEIDA+GR+RG G G
Sbjct: 1176 EAGTPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRG-AGLGG 1352
Query: 439 SNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKG 498
NDERE T+NQLL EMDGF +GV+VLA TNR D+LD+ALLRPGRFDR +++D PD+ G
Sbjct: 1353 GNDEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAG 1532
Query: 499 RDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDH 558
R +I Q++ + L + + ++A TPGF GAD+ N+ NEAA++AAR D +++ D
Sbjct: 1533 RVKILQVHSRGKALAKDVDF--DKIARRTPGFTGADLQNLMNEAAILAARRDLKEISKDE 1706
Query: 559 FEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAAL 618
A++RII G EKKN V+S+ +++ VAYHEAGHA+ G + +P+ K++I+PRG A
Sbjct: 1707 IADALERIIAGPEKKNAVVSEEKKKLVAYHEAGHALVGALMPEYDPVAKISIIPRGQAG- 1883
Query: 619 GFAQYVPSENLLR----TKEQLLDMTCMTLGGRAAEQVLIGA--ISTGAQNDLEKVTKMT 672
G + PSE L ++ L + + LGGR AE+V+ G ++TGA ND +V+++
Sbjct: 1884 GLTFFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGQDNVTTGASNDFMQVSRVA 2063
Query: 673 YAQVAIYGFSEKVGLLS--------FPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERT 724
V +GFS+K+G ++ F + K YS T +I+D+EVR+ V+ AYER
Sbjct: 2064 RQMVERFGFSKKIGQVAIGGGGGNPFLGQQMSSQKDYSMATADIVDKEVRELVDKAYERA 2243
Query: 725 VQLIEEHKEKLAQIAELLLEKEVLHQEDLVRI 756
Q+I H + L ++A+LL+EKE + E+ + +
Sbjct: 2244 TQIINTHIDILHKLAQLLIEKETVDGEEFMSL 2339
>TC76904 homologue to GP|4325041|gb|AAD17230.1| FtsH-like protein Pftf
precursor {Nicotiana tabacum}, partial (91%)
Length = 2667
Score = 399 bits (1025), Expect = e-111
Identities = 229/500 (45%), Positives = 317/500 (62%), Gaps = 12/500 (2%)
Frame = +1
Query: 272 LLGTLWFMGRKMQGGFGVGGGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCEEA 331
++G L+ + R+ GG G G G G F KA K V F DVAG +EA
Sbjct: 697 VIGVLFLLSRR-SGGMG---GPGGPGFPMQFGQSKA---KFQMEPNTGVTFDDVAGVDEA 855
Query: 332 KQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSD 391
KQ+ ME V FLK P+++ +GA+IPKG LLVGPPGTGKTLLAKA AGE+GVPF SISGS+
Sbjct: 856 KQDFMEVVEFLKKPERFTSVGARIPKGVLLVGPPGTGKTLLAKAIAGEAGVPFFSISGSE 1035
Query: 392 FMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLL 451
F+EMFVG+G SRVR+LF++A++ AP IVF+DEIDA+GR+RG G G NDERE TLNQLL
Sbjct: 1036 FVEMFVGIGASRVRDLFKKAKENAPCIVFVDEIDAVGRQRGT-GIGGGNDEREQTLNQLL 1212
Query: 452 VEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIK 511
EMDGF GV+V+A TNRADILD+ALLRPGRFDR +S+DVPD++GR +I +++ K
Sbjct: 1213 TEMDGFEGNTGVIVVAATNRADILDSALLRPGRFDRQVSVDVPDVRGRTEILKVHANNKK 1392
Query: 512 LDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLE 571
D++ S + +A TPGF+GAD+AN+ NEAA++A R + ++ + +IDRI+ G+E
Sbjct: 1393 FDNDVSL--EVIAMRTPGFSGADLANLLNEAAILAGRRGRTGISPKEIDDSIDRIVAGME 1566
Query: 572 KKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSEN-LL 630
+ + + VAYHE GHA+ G + + KVT+VPRG A G ++PS++ L
Sbjct: 1567 -GTLMTDGKSKSLVAYHEVGHAICGTLTPGHDAVQKVTLVPRGQAR-GLTWFIPSDDPTL 1740
Query: 631 RTKEQLLDMTCMTLGGRAAEQVLIG--AISTGAQNDLEKVTKMTYAQVAIYGFSEKVGLL 688
+K+QL LGGRAAE+++ G ++TGA DL+++T + V +G S+ +G
Sbjct: 1741 ISKQQLFARIVGGLGGRAAEEIIFGEPEVTTGAGGDLQQITGIARQMVVTFGMSD-IGPW 1917
Query: 689 SFPQNEDQFG---------KPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQIA 739
S + Q G S ID V+ + AYE + I ++E + +I
Sbjct: 1918 SLMDSSAQSGDVIMRMMARNSMSEKLAEDIDTAVKRLSDEAYEIALTQIRNNREAIDKIV 2097
Query: 740 ELLLEKEVLHQEDLVRILGE 759
E+LLEKE L ++ +L E
Sbjct: 2098 EVLLEKETLSGDEFRALLSE 2157
>TC87520 similar to GP|9757998|dbj|BAB08420.1 cell division protein FtsH
protease-like {Arabidopsis thaliana}, partial (53%)
Length = 1667
Score = 330 bits (847), Expect = 1e-90
Identities = 197/460 (42%), Positives = 275/460 (58%), Gaps = 5/460 (1%)
Frame = +2
Query: 331 AKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGS 390
AKQE+ E V +LKNP K+ LG K+PKG LL G PGTGKTLLAKA AGE+GVPF +GS
Sbjct: 2 AKQELEEVVEYLKNPAKFTRLGGKLPKGILLTGAPGTGKTLLAKAIAGEAGVPFFYRAGS 181
Query: 391 DFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIG--RKRGRGGFSGSNDERESTLN 448
+F EMFVGVG RVR+LFQ A++ AP I+FIDEIDA+G RK+ G + TL+
Sbjct: 182 EFEEMFVGVGARRVRSLFQAAKKKAPCIIFIDEIDAVGSTRKQWEG-------HTKKTLH 340
Query: 449 QLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYL- 507
QLLVEMDGF G++++A TN DILD AL RPGRFDR I + PD++GR +I ++YL
Sbjct: 341 QLLVEMDGFEQNEGIILMAATNLPDILDPALTRPGRFDRHIVVPNPDVRGRQEILELYLH 520
Query: 508 KKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRII 567
K D+ + +A TPGF GAD+AN+ N AA+ AA ++T E A DRII
Sbjct: 521 DKPTADNVD---IKAIARGTPGFNGADLANLVNIAAIKAAVEGAEKLTASQLEFAKDRII 691
Query: 568 GGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSE 627
G E+K +S+ ++ AYHE+GHA+ + P+ K TI+PRG+A Q S+
Sbjct: 692 MGTERKTMFVSEESKKLTAYHESGHAIVALNTDGAHPIHKATIMPRGSALGMVTQLPSSD 871
Query: 628 NLLRTKEQLLDMTCMTLGGRAAEQVLIGA--ISTGAQNDLEKVTKMTYAQVAIYGFSEKV 685
+K+QLL + +GGR AE+++ G ++TGA +DL+ T++ V+ G S+ +
Sbjct: 872 ETSISKKQLLARLDVCMGGRVAEELIFGRDNVTTGASSDLQSATELAQYMVSSCGMSDTI 1051
Query: 686 GLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQIAELLLEK 745
G + + S + + ID EV + AY+R L+++H++ L +A LLE
Sbjct: 1052GPIHIKERP-------SSEMQSRIDAEVVKLLRDAYDRVKALLKKHEKALHTLANALLES 1210
Query: 746 EVLHQEDLVRILGERPFKSAEPTNYDRFKLGFQDEEKAAE 785
E L E++ R+L Y KL Q E++ AE
Sbjct: 1211ETLSAEEIRRLL----------LPYREGKLPEQQEQEEAE 1300
>CB893172 homologue to PIR|T45642|T45 FtsH metalloproteinase-like protein -
Arabidopsis thaliana, partial (25%)
Length = 661
Score = 234 bits (596), Expect = 1e-61
Identities = 117/206 (56%), Positives = 153/206 (73%)
Frame = +3
Query: 341 FLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVG 400
FL+NP +Y LGA+ P+G LLVG PGTGKTLLAKA AGE+ VPF+S S S+F+E++VG+G
Sbjct: 39 FLRNPDRYARLGARPPRGVLLVGLPGTGKTLLAKAVAGEADVPFISCSASEFVELYVGMG 218
Query: 401 PSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTT 460
SRVR+LF A++ APSI+FIDEIDA+ + R NDERE TLNQLL EMDGF +
Sbjct: 219 ASRVRDLFARAKKEAPSIIFIDEIDAVAKSRDGKFRIVGNDEREQTLNQLLTEMDGFDSN 398
Query: 461 AGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYS 520
+ V+VLA TNRAD+LD AL RPGRFDR + ++ PD GR+ I ++++ K +L Y
Sbjct: 399 SPVIVLAATNRADVLDPALRRPGRFDRIVMVETPDRIGRESILKVHVSKKELPLAKDVYI 578
Query: 521 QRLAALTPGFAGADIANVCNEAALIA 546
+A++T GF GAD+AN+ NEAAL+A
Sbjct: 579 GDIASMTTGFTGADLANLVNEAALLA 656
>TC86186 similar to GP|4325041|gb|AAD17230.1| FtsH-like protein Pftf
precursor {Nicotiana tabacum}, partial (47%)
Length = 1303
Score = 179 bits (454), Expect(2) = 5e-55
Identities = 90/161 (55%), Positives = 119/161 (73%)
Frame = +2
Query: 270 LLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCE 329
L+++G L+ + R+ GG G GGS G+F G++ K V F DVAG +
Sbjct: 716 LIVIGGLFLLSRRSSGGTGGPGGS------GLFGFGQSKA-KFQMEPNTGVTFDDVAGVD 874
Query: 330 EAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISG 389
EAKQ+ ME V FLK P+++ +GA+IPKG LL+GPPGTGKTLLAKA AGE+GVPF SISG
Sbjct: 875 EAKQDFMEVVEFLKKPERFTAIGARIPKGVLLIGPPGTGKTLLAKAIAGEAGVPFFSISG 1054
Query: 390 SDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRK 430
S+F+EMFVG+G SRVR+LF++A++ AP IVF+DEIDA+GR+
Sbjct: 1055SEFVEMFVGIGASRVRDLFKKAKENAPCIVFVDEIDAVGRQ 1177
Score = 54.7 bits (130), Expect(2) = 5e-55
Identities = 25/38 (65%), Positives = 28/38 (72%)
Frame = +3
Query: 435 GFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRA 472
G G NDERE TLNQLL EMDGF G++V+A TNRA
Sbjct: 1188 GIGGGNDEREHTLNQLLTEMDGFEGNTGIIVIAPTNRA 1301
>BI271823 similar to GP|21592745|gb cell division protein FtsH-like protein
{Arabidopsis thaliana}, partial (33%)
Length = 714
Score = 211 bits (538), Expect = 7e-55
Identities = 108/189 (57%), Positives = 137/189 (72%)
Frame = +3
Query: 311 KVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKT 370
K K V F+DV G + AK E+ME V L+ Y++LGAK+P+G LLVGPPGTGKT
Sbjct: 63 KKRKPKSQTVGFEDVQGVDSAKVELMEIVSCLQGDINYQKLGAKLPRGVLLVGPPGTGKT 242
Query: 371 LLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRK 430
LLA+A AGE+GVPF ++S S+F+EMFVG G +R+R+LF AR+ APSI+FIDE+DA+G K
Sbjct: 243 LLARAVAGEAGVPFFTVSASEFVEMFVGRGAARIRDLFSRARKFAPSIIFIDELDAVGGK 422
Query: 431 RGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTIS 490
RGR G N+ER+ TLNQLL EMDGF + VVV+A TNR + LD AL RPGRF +
Sbjct: 423 RGR----GFNEERDQTLNQLLTEMDGFESEIRVVVIAATNRPEALDPALCRPGRFSXKVF 590
Query: 491 IDVPDIKGR 499
+ PD +GR
Sbjct: 591 VGEPDEEGR 617
>TC81865 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (16%)
Length = 624
Score = 206 bits (523), Expect = 4e-53
Identities = 105/131 (80%), Positives = 115/131 (87%), Gaps = 2/131 (1%)
Frame = +1
Query: 638 DMTCMTLGGRAAEQVLIGAISTGAQNDLEKVTKMTYAQVAIYGFSEKVGLLSFPQNED-- 695
D TCMTLGGRAAEQVLIG ISTGAQ+DLEKVTKMTYAQVA+YGFSEKVGLLSFPQ ED
Sbjct: 1 DRTCMTLGGRAAEQVLIGTISTGAQDDLEKVTKMTYAQVAVYGFSEKVGLLSFPQKEDSL 180
Query: 696 QFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQIAELLLEKEVLHQEDLVR 755
+ KPYS TG IID EVR+WV+ AYERTVQLIE+ KEK+AQ+AELLLEKEVLHQ+DL+
Sbjct: 181 EMSKPYSNKTGAIIDNEVREWVDKAYERTVQLIEKQKEKVAQLAELLLEKEVLHQDDLLP 360
Query: 756 ILGERPFKSAE 766
ILG RPFKS E
Sbjct: 361 ILGVRPFKSTE 393
>TC80276 similar to GP|20197264|gb|AAC31223.2 FtsH protease putative
{Arabidopsis thaliana}, partial (44%)
Length = 1384
Score = 205 bits (521), Expect = 6e-53
Identities = 118/306 (38%), Positives = 181/306 (58%), Gaps = 3/306 (0%)
Frame = +2
Query: 446 TLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQI 505
TLNQ+LVE+DGF G++V+ TN + LD AL+RPGRFDR + + PD++GR QI +
Sbjct: 41 TLNQMLVELDGFKQNDGIIVIGATNFPESLDKALVRPGRFDRHVVVPNPDVEGRRQILES 220
Query: 506 YLKKI-KLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAID 564
++ KI K D + R TPGF+GAD+AN+ N AAL AA V+M E A D
Sbjct: 221 HMSKILKADDVDLMITARC---TPGFSGADLANLVNVAALKAAMDGSKAVSMHDLEFARD 391
Query: 565 RIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYV 624
+I+ G E+K+ VIS+ R+ A+HE GHA+ + P+ K TIVPRG A +Q
Sbjct: 392 KILMGSERKSAVISEETRKMTAFHEGGHALVAIHSDGALPVHKATIVPRGMALGMVSQLP 571
Query: 625 PSENLLRTKEQLLDMTCMTLGGRAAEQVLIG--AISTGAQNDLEKVTKMTYAQVAIYGFS 682
+ +++Q+L + +GGR AE+++ G +++GA +DL K TK+ V YG S
Sbjct: 572 DKDQTSHSRKQMLAELDVCMGGRVAEELIFGESEVTSGASSDLSKATKLARQMVTKYGMS 751
Query: 683 EKVGLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQIAELL 742
+VG ++ +D G+ S +T +I++EV++ + AY ++ H+++L +A L
Sbjct: 752 TEVGPVTHNYYDD--GRSMSSETRLLIEKEVKNLLERAYNNAKTILTTHEKELHALANAL 925
Query: 743 LEKEVL 748
LE E L
Sbjct: 926 LEHETL 943
>TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome regulatory
particle triple-A ATPase subunit2b {Oryza sativa
(japonica cultivar-group), partial (98%)
Length = 1696
Score = 192 bits (488), Expect = 4e-49
Identities = 111/260 (42%), Positives = 155/260 (58%), Gaps = 1/260 (0%)
Frame = +3
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
V KV+K + D+ G + QEI E V L +P+ YE++G K PKG +L G PGT
Sbjct: 615 VMKVEKAPLES--YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGT 788
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTLLAKA A + FL + GS+ ++ ++G GP VR LF+ A +P+IVFIDEIDA+
Sbjct: 789 GKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPAIVFIDEIDAV 968
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
G KR SG E + T+ +LL ++DGF + V V+ TNR + LD ALLRPGR DR
Sbjct: 969 GTKR-YDAHSGGEREIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDR 1145
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
I +PDIK R +IFQI+ ++ L + + + F+GADI +C EA L+A
Sbjct: 1146KIEFPLPDIKTRRRIFQIHTSRMTLADDVNL--EEFVMTKDEFSGADIKAICTEAGLLAL 1319
Query: 548 RTDESQVTMDHFEAAIDRII 567
R +VT F+ A D+++
Sbjct: 1320RERRMKVTHPDFKKAKDKVM 1379
>TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15.70 -
Arabidopsis thaliana, complete
Length = 1629
Score = 191 bits (484), Expect = 1e-48
Identities = 111/260 (42%), Positives = 154/260 (58%), Gaps = 1/260 (0%)
Frame = +2
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
V KV+K + D+ G + QEI E V L +P+ YE++G K PKG +L G PGT
Sbjct: 617 VMKVEKAPLES--YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGT 790
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTLLAKA A + FL + GS+ ++ ++G GP VR LF+ A +PSIVFIDEIDA+
Sbjct: 791 GKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAV 970
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
G KR SG E + T+ +LL ++DGF + V V+ TNR + LD ALLRPGR DR
Sbjct: 971 GTKR-YDAHSGGEREIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDR 1147
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
I +PDIK R +IF I+ ++ L + + + F+GADI +C EA L+A
Sbjct: 1148KIEFPLPDIKTRRRIFTIHTSRMTLADDVNL--EEFVMTKDEFSGADIKAICTEAGLLAL 1321
Query: 548 RTDESQVTMDHFEAAIDRII 567
R +VT F+ A D+++
Sbjct: 1322RERRMKVTHADFKKAKDKVM 1381
>TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidopsis
thaliana}, partial (98%)
Length = 1633
Score = 187 bits (475), Expect = 1e-47
Identities = 106/278 (38%), Positives = 158/278 (56%), Gaps = 1/278 (0%)
Frame = +1
Query: 325 VAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVP 383
+ G ++ +EI E + +K+P+ +E LG PKG LL GPPGTGKTLLA+A A +
Sbjct: 628 IGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLARAVAHHTDCT 807
Query: 384 FLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDER 443
F+ +SGS+ ++ ++G G VR LF AR+ APSI+F+DEID+IG R G + E
Sbjct: 808 FIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARMESGSGNGDSEV 987
Query: 444 ESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIF 503
+ T+ +LL ++DGF + + VL TNR DILD ALLRPGR DR I P+ + R I
Sbjct: 988 QRTMLELLNQLDGFEASNKIKVLMATNRIDILDQALLRPGRIDRKIEFPNPNEESRRDIL 1167
Query: 504 QIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAI 563
+I+ +++ L +++A G +GA++ VC EA + A R VT + FE A+
Sbjct: 1168KIHSRRMNLMR--GIDLKKIAEKMNGASGAELKAVCTEAGMFALRERRVHVTQEDFEMAV 1341
Query: 564 DRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEH 601
+++ +KN + K + V Y H EH
Sbjct: 1342AKVMKKETEKNMSLRKLWK*FVFYVSGAHCHVQIIREH 1455
>TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1)
(Mg(2+), complete
Length = 1793
Score = 183 bits (465), Expect = 2e-46
Identities = 102/261 (39%), Positives = 158/261 (60%), Gaps = 1/261 (0%)
Frame = +1
Query: 307 AHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPP 365
+ V ++ + K + D+ G E+ QE++E + + + +++++LG + PKG LL GPP
Sbjct: 592 SRVKAMEVDEKPTEDYNDIGGLEKQIQELVEAIVLPMTHKERFQKLGIRPPKGVLLYGPP 771
Query: 366 GTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEID 425
GTGKTL+A+A A ++ FL ++G ++MF+G G VR+ FQ A++ +P I+FIDEID
Sbjct: 772 GTGKTLMARACAAQTNATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEID 951
Query: 426 AIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRF 485
AIG KR SG E + T+ +LL ++DGF + + V+A TNRADILD AL+R GR
Sbjct: 952 AIGTKRFDSEVSGDR-EVQRTMLELLNQLDGFSSDDRIKVIAATNRADILDPALMRSGRL 1128
Query: 486 DRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALI 545
DR I P + R +I QI+ +K+ + P + LA T F GA + VC EA ++
Sbjct: 1129DRKIEFPHPTEEARARILQIHSRKMNV--HPDVNFEELARSTDDFNGAQLKAVCVEAGML 1302
Query: 546 AARTDESQVTMDHFEAAIDRI 566
A R D ++V + F I ++
Sbjct: 1303ALRRDATEVNHEDFNEGIIQV 1365
>TC87676 PIR|G96577|G96577 26S proteasome ATPase subunit [imported] -
Arabidopsis thaliana, partial (56%)
Length = 963
Score = 181 bits (458), Expect = 1e-45
Identities = 97/229 (42%), Positives = 137/229 (59%)
Frame = +2
Query: 344 NPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSR 403
+P+K+ +LG PKG L GPPGTGKTLLA+A A + F+ + GS+ ++ +VG G
Sbjct: 17 HPEKFVKLGIDPPKGVLCYGPPGTGKTLLARAVANRTDACFIRVIGSELVQKYVGEGARM 196
Query: 404 VRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGV 463
VR LFQ AR IVF DE+DAIG R G G N E + T+ +++ ++DGF +
Sbjct: 197 VRELFQMARSKKACIVFFDEVDAIGGARFDDGVGGDN-EVQRTMLEIVNQLDGFDARGNI 373
Query: 464 VVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRL 523
VL TNR D LD ALLRPGR DR + +PD++ R QIF+I+ + + + + + + L
Sbjct: 374 KVLMATNRPDTLDPALLRPGRLDRKVEFGLPDLESRTQIFKIHTRTMNCERDIRF--ELL 547
Query: 524 AALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLEK 572
A L P GADI +VC EA + A R VT F A++++I G +K
Sbjct: 548 ARLCPNSTGADIRSVCTEAGMYAIRARRKTVTEKDFLDAVNKVIKGYQK 694
>TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AAA-ATPase
subunit RPT3 {Arabidopsis thaliana}, partial (94%)
Length = 1575
Score = 177 bits (450), Expect = 1e-44
Identities = 98/253 (38%), Positives = 151/253 (58%), Gaps = 1/253 (0%)
Frame = +2
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
++ + ++ K V + D+ GC+ KQEI E V L + + Y+++G P+G LL GPPGT
Sbjct: 533 ISLLSQSEKPDVTYNDIGGCDIQKQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGT 712
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKT+LAKA A + F+ + GS+F++ ++G GP VR++F+ A++ AP+I+FIDE+DAI
Sbjct: 713 GKTMLAKAVANHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLAKENAPAIIFIDEVDAI 892
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
R +G++ E + L +LL +MDGF T V V+ TNRAD LD ALLRPGR DR
Sbjct: 893 ATAR-FDAQTGADREVQRILMELLNQMDGFDQTVNVKVIMATNRADTLDPALLRPGRLDR 1069
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
I +PD + + +FQ+ K+ L E + + + A+I+ +C EA + A
Sbjct: 1070KIEFPLPDRRQKRLVFQVCTAKMNLSDEVDL--EDYVSRPDKISAAEISAICQEAGMHAV 1243
Query: 548 RTDESQVTMDHFE 560
R + + FE
Sbjct: 1244RKNRYVILPKDFE 1282
>TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle protein 48
homolog (Valosin containing protein homolog) (VCP).
[Soybean], partial (95%)
Length = 2785
Score = 176 bits (446), Expect = 3e-44
Identities = 98/227 (43%), Positives = 134/227 (58%), Gaps = 2/227 (0%)
Frame = +2
Query: 324 DVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGV 382
D+ G E K+E+ E V + +++P+K+E+ G KG L GPPG GKTLLAKA A E
Sbjct: 1538 DIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIANECQA 1717
Query: 383 PFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRG-GFSGSND 441
F+SI G + + M+ G + VR +F +AR AP ++F DE+D+I +RG G +G
Sbjct: 1718 NFISIKGPELLTMWFGESEANVREIFDKARGSAPCVLFFDELDSIATQRGSSVGDAGGAA 1897
Query: 442 ERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQ 501
+R LNQLL EMDG V ++ TNR DI+D ALLRPGR D+ I I +PD R Q
Sbjct: 1898 DR--VLNQLLTEMDGMSAKKTVFIIGATNRPDIIDPALLRPGRLDQLIYIPLPDEDSRHQ 2071
Query: 502 IFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAAR 548
IF+ L+K + + + LA T GF+GADI +C A A R
Sbjct: 2072 IFKACLRKSPISKDVDI--RALAKYTQGFSGADITEICQRACKYAIR 2206
Score = 175 bits (444), Expect = 5e-44
Identities = 102/257 (39%), Positives = 151/257 (58%), Gaps = 1/257 (0%)
Frame = +2
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
+ + D+N ++V + DV G + +I E V L++P+ ++ +G K PKG LL GPPG+
Sbjct: 674 IKREDENRLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPPGS 853
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTL+A+A A E+G F I+G + M G S +R F+EA + APSI+FIDEID+I
Sbjct: 854 GKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEIDSI 1033
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
KR + + ++QLL MDG + A V+V+ TNR + +D AL R GRFDR
Sbjct: 1034APKREK----THGEVERRIVSQLLTLMDGLKSRAHVIVMGATNRPNSIDPALRRFGRFDR 1201
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
I I VPD GR ++ +I+ K +KL + ++++ T G+ GAD+A +C EAAL
Sbjct: 1202EIDIGVPDEVGRLEVLRIHTKNMKLAEDVDL--EKISKETHGYVGADLAALCTEAALQCI 1375
Query: 548 RTDESQVTMDHFEAAID 564
R E +D + ID
Sbjct: 1376R--EKMDVIDLEDETID 1420
>TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AAA-ATPase
subunit RPT4a-like protein {Arabidopsis thaliana},
complete
Length = 1640
Score = 168 bits (425), Expect = 9e-42
Identities = 97/248 (39%), Positives = 141/248 (56%), Gaps = 1/248 (0%)
Frame = +2
Query: 320 VYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAG 378
+ + V G + +E+ E + L NP+ + +G K PKG LL GPPGTGKTLLA+A A
Sbjct: 476 ISYSAVGGLSDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIAS 655
Query: 379 ESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSG 438
FL + S ++ ++G +R +F AR P I+F+DEIDAIG +R G S
Sbjct: 656 NIDANFLKVVSSAIIDKYIGESARLIREMFGYARDHQPCIIFMDEIDAIGGRRFSEGTS- 832
Query: 439 SNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKG 498
++ E + TL +LL ++DGF V ++ TNR D+LD ALLRPGR DR I I +P+ +
Sbjct: 833 ADREIQRTLMELLNQLDGFDQLGKVKMIMATNRPDVLDPALLRPGRLDRKIEIPLPNEQS 1012
Query: 499 RDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDH 558
R +I +I+ I E Y + + L GF GAD+ NVC EA + A R + V +
Sbjct: 1013RMEILKIHAAGIAKHGEIDY--EAVVKLAEGFNGADLRNVCTEAGMSAIRAERDYVIHED 1186
Query: 559 FEAAIDRI 566
F A+ ++
Sbjct: 1187FMKAVRKL 1210
>BF647954 homologue to GP|11094192|db 26S proteasome regulatory particle
triple-A ATPase subunit4 {Oryza sativa (japonica
cultivar-group)}, partial (53%)
Length = 649
Score = 161 bits (407), Expect = 1e-39
Identities = 90/216 (41%), Positives = 126/216 (57%)
Frame = +2
Query: 344 NPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSR 403
NP+ + +G K PKG LL GPPGTGKTLLA+A A FL + S ++ ++G
Sbjct: 5 NPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIASNIDANFLKVVSSAIIDKYIGESSRL 184
Query: 404 VRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGV 463
+R +F AR P I+F+DEIDAIG +R G S ++ E + TL +LL ++DGF V
Sbjct: 185 IREMFGYARDHQPCIIFMDEIDAIGGRRFSEGTS-ADREIQRTLMELLNQLDGFDQLGKV 361
Query: 464 VVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRL 523
++ TNR D+LD ALLRPGR DR I I +P+ + R +I +I+ I E Y + +
Sbjct: 362 KIIMATNRPDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHAAGIAKHGEIDY--EAV 535
Query: 524 AALTPGFAGADIANVCNEAALIAARTDESQVTMDHF 559
L GF GAD+ N+C EA + A R + V + F
Sbjct: 536 VKLAEGFNGADLRNICTEAGMSAIRAERDYVIHEDF 643
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,057,653
Number of Sequences: 36976
Number of extensions: 244515
Number of successful extensions: 1624
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 1500
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1546
length of query: 807
length of database: 9,014,727
effective HSP length: 104
effective length of query: 703
effective length of database: 5,169,223
effective search space: 3633963769
effective search space used: 3633963769
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135467.11


