

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.23 + phase: 0
(150 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
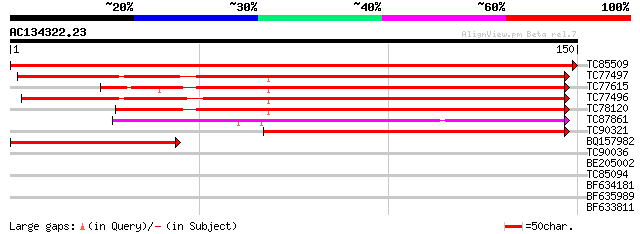
Score E
Sequences producing significant alignments: (bits) Value
TC85509 similar to SP|P09911|FER1_PEA Ferredoxin I chloroplast ... 305 6e-84
TC77497 similar to PIR|S62722|S62722 ferredoxin [2Fe-2S] fd1 pre... 159 4e-40
TC77615 similar to SP|P27788|FER3_MAIZE Ferredoxin III chloropl... 158 9e-40
TC77496 similar to PIR|S62722|S62722 ferredoxin [2Fe-2S] fd1 pre... 153 2e-38
TC78120 similar to PIR|G84673|G84673 probable ferredoxin [import... 149 3e-37
TC87861 similar to PIR|B71412|B71412 ferredoxin [2Fe-2S] - Arabi... 77 2e-15
TC90321 similar to GP|13174248|gb|AAK14422.1 putative ferredoxin... 73 5e-14
BQ157982 homologue to GP|2440148|emb| ABC-type transporter puta... 57 4e-09
TC90036 29 0.82
BE205002 similar to GP|14334760|gb putative receptor-protein kin... 26 5.3
TC85094 similar to PIR|T01449|T01449 cytoskeletal protein homolo... 26 7.0
BF634181 homologue to SP|Q08436|PMA3 Plasma membrane ATPase 3 (E... 26 7.0
BF635989 weakly similar to GP|9294446|dbj| sulfate transporter {... 26 7.0
BF633811 25 9.1
>TC85509 similar to SP|P09911|FER1_PEA Ferredoxin I chloroplast precursor.
[Garden pea] {Pisum sativum}, partial (90%)
Length = 831
Score = 305 bits (780), Expect = 6e-84
Identities = 150/150 (100%), Positives = 150/150 (100%)
Frame = +2
Query: 1 MATTPALYGTAVSTSFLRRQPMPVSISTTTKAFPSGFGLKSKTGKRGDLAVAMATYKVKL 60
MATTPALYGTAVSTSFLRRQPMPVSISTTTKAFPSGFGLKSKTGKRGDLAVAMATYKVKL
Sbjct: 119 MATTPALYGTAVSTSFLRRQPMPVSISTTTKAFPSGFGLKSKTGKRGDLAVAMATYKVKL 298
Query: 61 VTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDD 120
VTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDD
Sbjct: 299 VTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDD 478
Query: 121 QIEEGWVLTCVAYPTSDVTIETHKEEELTA 150
QIEEGWVLTCVAYPTSDVTIETHKEEELTA
Sbjct: 479 QIEEGWVLTCVAYPTSDVTIETHKEEELTA 568
>TC77497 similar to PIR|S62722|S62722 ferredoxin [2Fe-2S] fd1 precursor
non-photosynthetic - sweet orange, partial (84%)
Length = 880
Score = 159 bits (402), Expect = 4e-40
Identities = 83/147 (56%), Positives = 105/147 (70%), Gaps = 1/147 (0%)
Frame = +2
Query: 3 TTPALYGTAVSTSFLRRQPMPVSISTTTKAFPSGFGLKSKTGKRGDLAVAMATYKVKLVT 62
TT +LYGTA S + + P S+ + K FGLKS + + AMA YKVKL+
Sbjct: 152 TTASLYGTAPSRTSCALRKSPSSLRSV-KNVSKTFGLKSSSFR----VSAMAVYKVKLIG 316
Query: 63 PEGTQ-EFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQ 121
P+GT+ EFD P D YILD AE+ G++LPYSCRAG+CS+CAG+V G+VDQSD SFLD+ Q
Sbjct: 317 PDGTENEFDAPDDSYILDSAEDAGVELPYSCRAGACSTCAGQVVTGSVDQSDQSFLDEQQ 496
Query: 122 IEEGWVLTCVAYPTSDVTIETHKEEEL 148
IE+G++LTCV+YP SD I THKEEEL
Sbjct: 497 IEKGYLLTCVSYPKSDTVIYTHKEEEL 577
>TC77615 similar to SP|P27788|FER3_MAIZE Ferredoxin III chloroplast
precursor (Fd III). [Maize] {Zea mays}, partial (65%)
Length = 906
Score = 158 bits (399), Expect = 9e-40
Identities = 81/135 (60%), Positives = 102/135 (75%), Gaps = 11/135 (8%)
Frame = +3
Query: 25 SISTTTKAFPSGFG----------LKSKTGKRGDLAVAMATYKVKLVTPEGTQ-EFDCPS 73
+I+TT K FPS FG LKS + R AMA YKVKL+ P+G + EFD P
Sbjct: 168 TIATTAK-FPSSFGSTKTYSKTCGLKSSSSYR---TTAMAAYKVKLIGPDGKENEFDAPD 335
Query: 74 DVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAY 133
DVYILD AE+ G++LPYSCRAG+CS+CAGK+ +G+VDQSDGSFLDD+Q+++G+VLTCV+Y
Sbjct: 336 DVYILDAAEDAGVELPYSCRAGACSTCAGKIVSGSVDQSDGSFLDDNQLKDGFVLTCVSY 515
Query: 134 PTSDVTIETHKEEEL 148
PT+D IETHKE EL
Sbjct: 516 PTADCIIETHKEGEL 560
>TC77496 similar to PIR|S62722|S62722 ferredoxin [2Fe-2S] fd1 precursor
non-photosynthetic - sweet orange, partial (84%)
Length = 827
Score = 153 bits (387), Expect = 2e-38
Identities = 81/146 (55%), Positives = 103/146 (70%), Gaps = 1/146 (0%)
Frame = +2
Query: 4 TPALYGTAVSTSFLRRQPMPVSISTTTKAFPSGFGLKSKTGKRGDLAVAMATYKVKLVTP 63
T +L+GTA S + P ++ + K FGLKS + + AMA YKVKL+ P
Sbjct: 98 TASLHGTASSRTSCALLKNPSTLRSV-KNVSKRFGLKSSSFRIS----AMAVYKVKLIQP 262
Query: 64 EGTQ-EFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQI 122
+GT+ EFD P D YILD AEE G++LPYSCRAG+CS+CAG+V +G+VDQSD SFLD QI
Sbjct: 263 DGTENEFDAPDDYYILDSAEEAGVELPYSCRAGACSTCAGQVVSGSVDQSDQSFLDKQQI 442
Query: 123 EEGWVLTCVAYPTSDVTIETHKEEEL 148
E+G++LTCV+YP SD I THKEEEL
Sbjct: 443 EKGYLLTCVSYPKSDTVIYTHKEEEL 520
>TC78120 similar to PIR|G84673|G84673 probable ferredoxin [imported] -
Arabidopsis thaliana, partial (75%)
Length = 853
Score = 149 bits (377), Expect = 3e-37
Identities = 73/121 (60%), Positives = 89/121 (73%), Gaps = 1/121 (0%)
Frame = +1
Query: 29 TTKAFPSGFGLKSKTGKRGDLAVAMATYKVKLVTPEGTQ-EFDCPSDVYILDHAEEVGID 87
+ K FGLKS R A+A YKVKL+ P+G + EF+ D YILD AE G++
Sbjct: 199 SAKNVSRSFGLKSSASSR---VTAVAAYKVKLIGPDGKENEFEATDDTYILDAAENAGVE 369
Query: 88 LPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVTIETHKEEE 147
LPYSCRAG+CS+CAGKV +G+VDQSDGSFLDD+Q+ EG+VLTCVAYPTSD I THKE +
Sbjct: 370 LPYSCRAGACSTCAGKVVSGSVDQSDGSFLDDNQLNEGYVLTCVAYPTSDCVIHTHKEGD 549
Query: 148 L 148
L
Sbjct: 550 L 552
>TC87861 similar to PIR|B71412|B71412 ferredoxin [2Fe-2S] - Arabidopsis
thaliana, partial (66%)
Length = 695
Score = 77.4 bits (189), Expect = 2e-15
Identities = 43/127 (33%), Positives = 66/127 (51%), Gaps = 6/127 (4%)
Frame = +2
Query: 28 TTTKAFPSGFGLKSKTGKRGDLAVAMATYKVK-----LVTPEG-TQEFDCPSDVYILDHA 81
TT +PS + + G + T+ V+ ++ EG T + + D IL A
Sbjct: 95 TTKLPYPSQLNTRPRLGSGSHPSSPSLTFTVRSSYKVVIEHEGKTTQLEVEPDETILSKA 274
Query: 82 EEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVTIE 141
+ G+D+P+ C+ G C +C ++ +G VDQSDG L DD +E G+ L C +YP SD I
Sbjct: 275 LDSGLDVPHDCKLGVCMTCPARLISGTVDQSDG-MLSDDVVERGYALLCASYPRSDCHIR 451
Query: 142 THKEEEL 148
E+EL
Sbjct: 452 VIPEDEL 472
>TC90321 similar to GP|13174248|gb|AAK14422.1 putative ferredoxin {Oryza
sativa}, partial (65%)
Length = 649
Score = 72.8 bits (177), Expect = 5e-14
Identities = 33/81 (40%), Positives = 50/81 (60%)
Frame = +3
Query: 68 EFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWV 127
EF P D YIL AE I LP++CR G C+SCA ++ G + Q + + + E+G+
Sbjct: 261 EFLVPEDQYILHTAESQNITLPFACRHGCCTSCAVRIKNGKIKQPEALGISAELREQGYA 440
Query: 128 LTCVAYPTSDVTIETHKEEEL 148
L CV++P SD+ +ET E+E+
Sbjct: 441 LLCVSFPYSDLEVETQDEDEV 503
>BQ157982 homologue to GP|2440148|emb| ABC-type transporter putative
membrane subunit {Pseudomonas aeruginosa}, partial (2%)
Length = 926
Score = 56.6 bits (135), Expect = 4e-09
Identities = 30/45 (66%), Positives = 34/45 (74%)
Frame = +1
Query: 1 MATTPALYGTAVSTSFLRRQPMPVSISTTTKAFPSGFGLKSKTGK 45
+ATTPA GTA STSFL RQPMP +I+ TT A PSG GL+S T K
Sbjct: 76 VATTPA*NGTAKSTSFLMRQPMPKTITPTTNALPSGSGLES*TRK 210
>TC90036
Length = 627
Score = 28.9 bits (63), Expect = 0.82
Identities = 14/64 (21%), Positives = 30/64 (46%), Gaps = 5/64 (7%)
Frame = +3
Query: 67 QEFDCPSDVYILDHAEEVGIDLPYSCRAGSC-----SSCAGKVAAGAVDQSDGSFLDDDQ 121
+E CP +YI ++ ++G + Y C +C + C G++ + ++ +
Sbjct: 75 KESGCP--IYICNYCNQLGFESSYQCLNSNCTYILHTQCVGRIVELVEHRPTTECVERQR 248
Query: 122 IEEG 125
I+EG
Sbjct: 249 IDEG 260
>BE205002 similar to GP|14334760|gb putative receptor-protein kinase
{Arabidopsis thaliana}, partial (16%)
Length = 555
Score = 26.2 bits (56), Expect = 5.3
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +3
Query: 57 KVKLVTPEGTQEFDCPSDVYI 77
+VKL P GT + PSDVY+
Sbjct: 369 EVKLEYPPGTPXYIAPSDVYV 431
>TC85094 similar to PIR|T01449|T01449 cytoskeletal protein homolog
F24O1.11 - Arabidopsis thaliana, partial (11%)
Length = 577
Score = 25.8 bits (55), Expect = 7.0
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = +1
Query: 4 TPALYGTAVSTSFLRRQPMPVSISTTTKA 32
TP ++ + VST L P+P S S+T ++
Sbjct: 61 TPCIHFSLVSTIILNPNPLPTSSSSTHRS 147
>BF634181 homologue to SP|Q08436|PMA3 Plasma membrane ATPase 3 (EC 3.6.3.6)
(Proton pump 3). [Leadwort-leaved tobacco], partial
(23%)
Length = 663
Score = 25.8 bits (55), Expect = 7.0
Identities = 12/20 (60%), Positives = 13/20 (65%)
Frame = -3
Query: 27 STTTKAFPSGFGLKSKTGKR 46
S T A PS F +KS TGKR
Sbjct: 196 SGKTPANPSAFSIKSSTGKR 137
>BF635989 weakly similar to GP|9294446|dbj| sulfate transporter {Arabidopsis
thaliana}, partial (9%)
Length = 655
Score = 25.8 bits (55), Expect = 7.0
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = +1
Query: 76 YILDHAEEVGIDLPYSCRAGSCSSCAGKVA 105
YI+ +E G + +C SC C GKVA
Sbjct: 16 YIMRT*KETGTKITSACIGKSCWKCDGKVA 105
>BF633811
Length = 313
Score = 25.4 bits (54), Expect = 9.1
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -2
Query: 84 VGIDLPYSCRAGSCSSCAGKV 104
VG+ LP++ G C C+G V
Sbjct: 159 VGLILPFAVATGVCEDCSGAV 97
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,402,713
Number of Sequences: 36976
Number of extensions: 57293
Number of successful extensions: 212
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 208
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 208
length of query: 150
length of database: 9,014,727
effective HSP length: 87
effective length of query: 63
effective length of database: 5,797,815
effective search space: 365262345
effective search space used: 365262345
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC134322.23


