

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133139.1 - phase: 0 /pseudo
(365 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
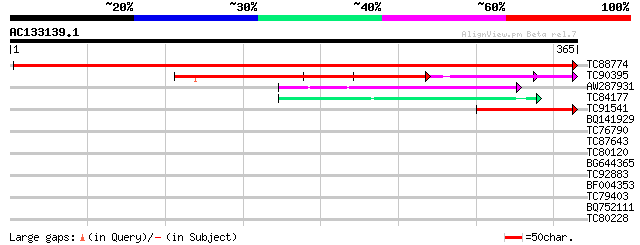
Score E
Sequences producing significant alignments: (bits) Value
TC88774 similar to PIR|T01394|T01394 hypothetical protein T4I9.1... 702 0.0
TC90395 similar to GP|19310761|gb|AAL85111.1 unknown protein {Ar... 211 4e-55
AW287931 weakly similar to PIR|T01062|T01 hypothetical protein Y... 69 4e-12
TC84177 similar to PIR|T22172|T22172 hypothetical protein F44E5.... 43 2e-04
TC91541 weakly similar to GP|15810253|gb|AAL07014.1 unknown prot... 41 6e-04
BQ141929 similar to GP|19310761|gb| unknown protein {Arabidopsis... 34 0.10
TC76790 similar to PIR|T06817|T06817 RNA-binding protein - garde... 31 0.66
TC87643 similar to PIR|C71405|C71405 probable casein kinase I - ... 31 0.86
TC80120 homologue to GP|11994319|dbj|BAB02278. casein kinase {Ar... 31 0.86
BG644365 31 0.86
TC92883 29 2.5
BF004353 similar to SP|P22335|HSF2_ Heat shock factor protein HS... 29 2.5
TC79403 homologue to GP|10190525|emb|CAC09322. unnamed protein p... 29 3.3
BQ752111 similar to PIR|JQ2282|JQ22 negatively phytochrome regul... 28 5.6
TC80228 similar to GP|21592725|gb|AAM64674.1 unknown {Arabidopsi... 28 5.6
>TC88774 similar to PIR|T01394|T01394 hypothetical protein T4I9.13 -
Arabidopsis thaliana, partial (47%)
Length = 1375
Score = 702 bits (1813), Expect = 0.0
Identities = 357/363 (98%), Positives = 358/363 (98%)
Frame = +2
Query: 3 PFCFIMQIQWYPILKNMNFSGTITKPNFPFPPTEMPILTFGHSKLTAATLALQLPCNYIR 62
P F IQWYPILKNMNFSGTITKPNFPFPPTEMPILTFGHSKLTAATLALQLPCNYIR
Sbjct: 47 PSHFPFYIQWYPILKNMNFSGTITKPNFPFPPTEMPILTFGHSKLTAATLALQLPCNYIR 226
Query: 63 KIRFITKLQCSVAGRTDSYRTSAADSQGSKRDSGEIQRKRRGASSLYAGYARPSLSEMKK 122
KIRFITKLQCSVAGRTDSYRTSAADSQGSKRDSGEIQRKRRGASSLYAGYARPSLSEMKK
Sbjct: 227 KIRFITKLQCSVAGRTDSYRTSAADSQGSKRDSGEIQRKRRGASSLYAGYARPSLSEMKK 406
Query: 123 DKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIEDINNYPLVLG 182
DKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIEDINNYPLVLG
Sbjct: 407 DKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIEDINNYPLVLG 586
Query: 183 CSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDI 242
CSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDI
Sbjct: 587 CSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDI 766
Query: 243 PRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVE 302
PRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILT+FPEILGMRVGRVIKPFVE
Sbjct: 767 PRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTKFPEILGMRVGRVIKPFVE 946
Query: 303 YLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIG 362
YLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIG
Sbjct: 947 YLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIG 1126
Query: 363 TDL 365
TDL
Sbjct: 1127TDL 1135
>TC90395 similar to GP|19310761|gb|AAL85111.1 unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 785
Score = 211 bits (537), Expect = 4e-55
Identities = 95/169 (56%), Positives = 138/169 (81%), Gaps = 4/169 (2%)
Frame = +1
Query: 107 SLYAGYARPSLS----EMKKDKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVD 162
S + Y P+++ + +K+K R +++++L+G+GI+PDEL LELP TV+VM+ERV+
Sbjct: 232 SKFPEYEMPTVTWGVIQGRKEKLVSRVIIFDYLKGLGIIPDELQDLELPSTVEVMRERVE 411
Query: 163 FLHSLGLTIEDINNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVV 222
F+ LGLTI+DIN YPL+LGCSV+KNM+P+L YL K+G+ +S + +F++ YPQVLHASV+
Sbjct: 412 FIQKLGLTIDDINQYPLILGCSVRKNMIPILGYLEKIGISRSKLGEFIKNYPQVLHASVI 591
Query: 223 VDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGI 271
V+L PV+K+L+G+D++ DDI VL++YPE+LGFKLEGTMSTSVAYL+ I
Sbjct: 592 VELAPVIKFLRGLDVEKDDIGFVLQKYPELLGFKLEGTMSTSVAYLVSI 738
Score = 56.6 bits (135), Expect = 1e-08
Identities = 40/152 (26%), Positives = 74/152 (48%), Gaps = 1/152 (0%)
Frame = +1
Query: 190 VPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPV-VKYLQGMDIKPDDIPRVLER 248
V + DYL LG+ + Q L V+++ V+++Q + + DDI +
Sbjct: 310 VIIFDYLKGLGIIPDEL--------QDLELPSTVEVMRERVEFIQKLGLTIDDI----NQ 453
Query: 249 YPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLG 308
YP +LG + M + YL IG+ R +LG + +P++L V + P +++L L
Sbjct: 454 YPLILGCSVRKNMIPILGYLEKIGISRSKLGEFIKNYPQVLHASVIVELAPVIKFLRGLD 633
Query: 309 IPRLAIARLIETQPYILGFDLDEKVKPNVKSL 340
+ + I +++ P +LGF L+ + +V L
Sbjct: 634 VEKDDIGFVLQKYPELLGFKLEGTMSTSVAYL 729
Score = 53.5 bits (127), Expect = 1e-07
Identities = 35/144 (24%), Positives = 71/144 (49%)
Frame = +1
Query: 222 VVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGI 281
+V V + YL+G+ I PD++ + P + M V ++ +G+ ++
Sbjct: 298 LVSRVIIFDYLKGLGIIPDELQDL--ELPSTVE-----VMRERVEFIQKLGLTIDDIN-- 450
Query: 282 LTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLE 341
++P ILG V + + P + YLE +GI R + I+ P +L + ++ P +K L
Sbjct: 451 --QYPLILGCSVRKNMIPILGYLEKIGISRSKLGEFIKNYPQVLHASVIVELAPVIKFLR 624
Query: 342 EFNVRETSLASIIAQYPDIIGTDL 365
+V + + ++ +YP+++G L
Sbjct: 625 GLDVEKDDIGFVLQKYPELLGFKL 696
>AW287931 weakly similar to PIR|T01062|T01 hypothetical protein YUP8H12R.46 -
Arabidopsis thaliana, partial (20%)
Length = 503
Score = 68.6 bits (166), Expect = 4e-12
Identities = 39/156 (25%), Positives = 77/156 (49%)
Frame = +2
Query: 174 INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQ 233
I P +LGCS K + V D +LGV++ + Q + PQ+L D + VV + +
Sbjct: 41 IKRQPQLLGCSTSKLKLMV-DQFAELGVQRKKLYQVITRSPQLL-LQKPEDFLQVVMFFE 214
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRV 293
M ++I R+L R PE+ + T+ + + +L I V + + ++ ++PE+L +
Sbjct: 215 NMGFDKENIGRILARCPEIFATSISKTLQSKIEFLSRIEVSKAYIPLVIRKYPELLVSDI 394
Query: 294 GRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDL 329
+ + + Y +G+ IA ++ +LG+ +
Sbjct: 395 NKTLPQRIVYFMKVGLSEKEIALMVRKFSPLLGYSI 502
>TC84177 similar to PIR|T22172|T22172 hypothetical protein F44E5.2 -
Caenorhabditis elegans, partial (2%)
Length = 824
Score = 42.7 bits (99), Expect = 2e-04
Identities = 38/169 (22%), Positives = 65/169 (37%)
Frame = +2
Query: 174 INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQ 233
I P +L C+ K ++P +L G S I +R P+ L S+ + +L
Sbjct: 116 IKRVPWILSCNTHKTILPKFQFLLSKGASTSDIVHMVRGNPRFLELSLKNHQIKFEFFLS 295
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRV 293
I +L YP++L E + L ++ L + + +
Sbjct: 296 -KGASSSHIVSLLTTYPQILQTSFENRIIPLFKLLTRFFKTNKDTIVCLIQHSKWVTSHP 472
Query: 294 GRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEE 342
+I + + G+ IARL++ +P I G K +KSLEE
Sbjct: 473 HHLIVASINLISDFGVSDSVIARLLQNKPSIFG------SKDLIKSLEE 601
>TC91541 weakly similar to GP|15810253|gb|AAL07014.1 unknown protein
{Arabidopsis thaliana}, partial (66%)
Length = 820
Score = 41.2 bits (95), Expect = 6e-04
Identities = 19/66 (28%), Positives = 44/66 (65%), Gaps = 1/66 (1%)
Frame = +3
Query: 301 VEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSL-EEFNVRETSLASIIAQYPD 359
++YL SL + +++L++ P +LG +L E++K N+K L E++++ SL +++ + P
Sbjct: 582 LDYLRSLNLSDDDLSKLLKKFPEVLGCNLKEELKANIKILKEQWSIEGKSLKNLLLRNPK 761
Query: 360 IIGTDL 365
++G ++
Sbjct: 762 VLGYNI 779
Score = 38.9 bits (89), Expect = 0.003
Identities = 18/64 (28%), Positives = 38/64 (59%), Gaps = 1/64 (1%)
Frame = +3
Query: 231 YLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYL-IGIGVGRRELGGILTRFPEIL 289
YL+ +++ DD+ ++L+++PEVLG L+ + ++ L + + L +L R P++L
Sbjct: 588 YLRSLNLSDDDLSKLLKKFPEVLGCNLKEELKANIKILKEQWSIEGKSLKNLLLRNPKVL 767
Query: 290 GMRV 293
G +
Sbjct: 768 GYNI 779
Score = 36.6 bits (83), Expect = 0.016
Identities = 23/87 (26%), Positives = 47/87 (53%), Gaps = 1/87 (1%)
Frame = +3
Query: 247 ERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYL-E 305
+R EV F+ +++ ++ YL + + +L +L +FPE+LG + +K ++ L E
Sbjct: 537 DREKEVPKFE---SVNGTLDYLRSLNLSDDDLSKLLKKFPEVLGCNLKEELKANIKILKE 707
Query: 306 SLGIPRLAIARLIETQPYILGFDLDEK 332
I ++ L+ P +LG+++D K
Sbjct: 708 QWSIEGKSLKNLLLRNPKVLGYNIDWK 788
Score = 35.8 bits (81), Expect = 0.027
Identities = 19/67 (28%), Positives = 38/67 (56%), Gaps = 1/67 (1%)
Frame = +3
Query: 193 LDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYL-QGMDIKPDDIPRVLERYPE 251
LDYL L + +++ L+ +P+VL ++ +L +K L + I+ + +L R P+
Sbjct: 582 LDYLRSLNLSDDDLSKLLKKFPEVLGCNLKEELKANIKILKEQWSIEGKSLKNLLLRNPK 761
Query: 252 VLGFKLE 258
VLG+ ++
Sbjct: 762 VLGYNID 782
>BQ141929 similar to GP|19310761|gb| unknown protein {Arabidopsis thaliana},
partial (10%)
Length = 866
Score = 33.9 bits (76), Expect = 0.10
Identities = 13/30 (43%), Positives = 22/30 (73%)
Frame = +3
Query: 121 KKDKATLRKVVYEFLRGIGIVPDELDGLEL 150
+K+K R +++ +L+G GI+PDE + LEL
Sbjct: 129 RKEKLVTRVIIFNYLKG*GIIPDESNELEL 218
>TC76790 similar to PIR|T06817|T06817 RNA-binding protein - garden pea,
partial (79%)
Length = 1141
Score = 31.2 bits (69), Expect = 0.66
Identities = 15/36 (41%), Positives = 23/36 (63%)
Frame = +2
Query: 28 PNFPFPPTEMPILTFGHSKLTAATLALQLPCNYIRK 63
P+FPFPP P ++ SK TA + +L LP + ++K
Sbjct: 125 PSFPFPPN--PSISISPSKPTAQSQSLPLPYSLLKK 226
>TC87643 similar to PIR|C71405|C71405 probable casein kinase I - Arabidopsis
thaliana, partial (71%)
Length = 2349
Score = 30.8 bits (68), Expect = 0.86
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +3
Query: 155 DVMKERVDFLHSLGLTIEDINNYPLVLGCSVKKNMVPVLDY 195
D + RV+++HS G DI ++G K N V ++DY
Sbjct: 900 DQLINRVEYMHSRGFLHRDIKPDNFLMGLGRKANQVYIIDY 1022
>TC80120 homologue to GP|11994319|dbj|BAB02278. casein kinase {Arabidopsis
thaliana}, partial (71%)
Length = 1584
Score = 30.8 bits (68), Expect = 0.86
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +1
Query: 155 DVMKERVDFLHSLGLTIEDINNYPLVLGCSVKKNMVPVLDY 195
D + RV+++HS G DI ++G K N V ++DY
Sbjct: 529 DQLINRVEYMHSRGFLHRDIKPDNFLMGLGRKANQVYIIDY 651
>BG644365
Length = 555
Score = 30.8 bits (68), Expect = 0.86
Identities = 15/58 (25%), Positives = 27/58 (45%)
Frame = +3
Query: 297 IKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASII 354
+ F+E+ ++ G +LI +P +L +D+ +KP K L+ E II
Sbjct: 162 VSAFIEFFQNHGFSATQTKKLIRLRPKLLVSKVDKTLKPKFKFLQSIGFTEDERNKII 335
>TC92883
Length = 924
Score = 29.3 bits (64), Expect = 2.5
Identities = 17/49 (34%), Positives = 25/49 (50%), Gaps = 3/49 (6%)
Frame = +3
Query: 187 KNMVPVLDYLGKLGVRKSTITQFLRTYPQV---LHASVVVDLVPVVKYL 232
K+++P GKL V K + R V LH + ++D V VVK+L
Sbjct: 585 KSLIPFAIRTGKLAVLKLLVANGCRINDSVDFVLHEAAIIDRVDVVKFL 731
>BF004353 similar to SP|P22335|HSF2_ Heat shock factor protein HSF24 (Heat
shock transcription factor 24) (HSTF 24), partial (17%)
Length = 406
Score = 29.3 bits (64), Expect = 2.5
Identities = 13/50 (26%), Positives = 23/50 (46%)
Frame = -3
Query: 180 VLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVV 229
VL C +K + L YLGK+ S + +P H ++++ P +
Sbjct: 230 VLSCRIKLEKLLCLKYLGKVSSENSAGDHTTKPFPSSFHDTILMPFSPSI 81
>TC79403 homologue to GP|10190525|emb|CAC09322. unnamed protein product
{Pisum sativum}, partial (42%)
Length = 1027
Score = 28.9 bits (63), Expect = 3.3
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -3
Query: 13 YPILKNMNFSGTITKPNFPF 32
YP L N NFS TI+ NF +
Sbjct: 608 YPFLSNQNFSNTISSSNFDY 549
>BQ752111 similar to PIR|JQ2282|JQ22 negatively phytochrome regulated protein
I - swollen duckweed, partial (17%)
Length = 737
Score = 28.1 bits (61), Expect = 5.6
Identities = 16/43 (37%), Positives = 27/43 (62%)
Frame = +1
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRR 276
G++ + DD+ +VLE +V+G ++ G VAY+ G G+ RR
Sbjct: 619 GLEARFDDVAKVLEEKNDVVGSRVRG----QVAYVAG-GLPRR 732
>TC80228 similar to GP|21592725|gb|AAM64674.1 unknown {Arabidopsis
thaliana}, partial (42%)
Length = 798
Score = 28.1 bits (61), Expect = 5.6
Identities = 23/99 (23%), Positives = 41/99 (41%)
Frame = +1
Query: 112 YARPSLSEMKKDKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTI 171
+ R S+ + K+ +KVV +F+ GI + E VDV ++ ++ + +G+ +
Sbjct: 307 HRRDSVDDEPKEHKETKKVVSDFVDGIAKEQQQQKQKENGEEVDVNEDEIEMMKMMGIPV 486
Query: 172 EDINNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFL 210
S K VP D G V K Q++
Sbjct: 487 G---------FDSTKGKPVPGADVSGVRAVTKRQPRQYM 576
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.142 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,821,862
Number of Sequences: 36976
Number of extensions: 125036
Number of successful extensions: 672
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 651
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 662
length of query: 365
length of database: 9,014,727
effective HSP length: 97
effective length of query: 268
effective length of database: 5,428,055
effective search space: 1454718740
effective search space used: 1454718740
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC133139.1


