

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC131249.9 - phase: 0 /pseudo
(421 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
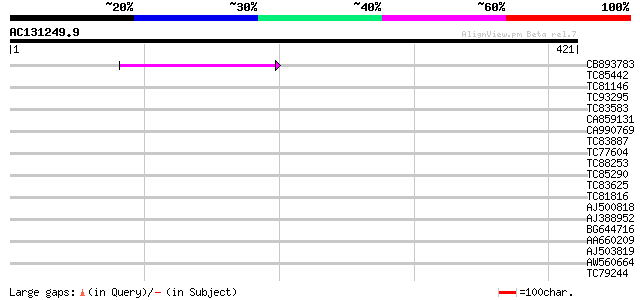
Score E
Sequences producing significant alignments: (bits) Value
CB893783 weakly similar to GP|22830935|dbj hypothetical protein~... 42 6e-04
TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein fo... 40 0.002
TC81146 similar to GP|13543783|gb|AAH06040.1 Unknown (protein fo... 38 0.006
TC93295 weakly similar to GP|5821157|dbj|BAA83720.1 GAMP {Procam... 35 0.054
TC83583 similar to PIR|T06379|T06379 SAR DNA-binding protein 2 -... 35 0.054
CA859131 weakly similar to GP|23496809|gb| 101 kd malaria antige... 35 0.054
CA990769 weakly similar to GP|21594005|gb| unknown {Arabidopsis ... 35 0.054
TC83887 weakly similar to GP|13310891|gb|AAK13153.2 hypothetical... 35 0.071
TC77604 homologue to PIR|T06377|T06377 SAR DNA-binding protein-1... 34 0.12
TC88253 weakly similar to GP|15810597|gb|AAL07186.1 unknown prot... 33 0.16
TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsi... 33 0.21
TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 33 0.27
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 33 0.27
AJ500818 weakly similar to GP|10435304|dbj unnamed protein produ... 33 0.27
AJ388952 similar to PIR|T04526|T045 hypothetical protein F16A16.... 33 0.27
BG644716 homologue to GP|7298552|gb| brat gene product {Drosophi... 32 0.35
AA660209 similar to GP|6642637|gb|A unknown protein {Arabidopsis... 32 0.35
AJ503819 32 0.35
AW560664 similar to GP|23495077|gb hypothetical protein {Plasmod... 32 0.46
TC79244 weakly similar to PIR|B86193|B86193 hypothetical protein... 32 0.60
>CB893783 weakly similar to GP|22830935|dbj hypothetical protein~similar to
gag-pol polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 853
Score = 41.6 bits (96), Expect = 6e-04
Identities = 37/121 (30%), Positives = 63/121 (51%), Gaps = 1/121 (0%)
Frame = +1
Query: 82 W*ISHD*EDVGKPS*GGRIKSKGKSFPYKMLSARKGMFSNY*WR*LY*CC*CTHGG*IRI 141
W*I++ *+ + + R +K + P+ + AR + ++* L CC +GG I
Sbjct: 151 W*ITNS*KKLTHHNSQRRTMAKTQYLPHSLNLARHSL*CHH*QWKL*ECCIQLYGGKIGA 330
Query: 142 RNKTSP*ALQTSMA**KCRNAC*QTSRGLF*NW-EV*RCCCV*CSTNGGLSFVARKTMAI 200
+K SP ++Q +MA + R +F +W ++ R C V*C +G LS + +T+AI
Sbjct: 331 ADKRSPTSIQVTMAKKR*RGQSL*VLPCIFFHWAKIQRQCMV*CHLHGCLSHASWETLAI 510
Query: 201 * 201
*
Sbjct: 511 * 513
>TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein for MGC:7642)
{Mus musculus}, partial (62%)
Length = 585
Score = 40.0 bits (92), Expect = 0.002
Identities = 20/43 (46%), Positives = 29/43 (66%), Gaps = 2/43 (4%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRKK--RKRNRKRKERKFHGKKRRNKK 273
K+R K+ RK +K KR+K RK+ RKRK RK GK+++ +K
Sbjct: 109 KKRKKKQRRKKRRKKGKRRKNQRKKKRKRKRRKKKGKRKKRRK 237
Score = 39.7 bits (91), Expect = 0.002
Identities = 17/40 (42%), Positives = 32/40 (79%)
Frame = +1
Query: 234 RRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
+R K+++RK +RK+KKRK+ ++RK+R+ GK+R+N++
Sbjct: 64 QRRKKSQRK--KNQRKKKKRKKKQRRKKRRKKGKRRKNQR 177
Score = 38.5 bits (88), Expect = 0.005
Identities = 16/41 (39%), Positives = 29/41 (70%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
K++ ++ RK R++ ++K+KR RKR+++K KKRR +K
Sbjct: 121 KKQRRKKRRKKGKRRKNQRKKKRKRKRRKKKGKRKKRRKRK 243
Score = 31.2 bits (69), Expect = 0.78
Identities = 13/35 (37%), Positives = 23/35 (65%)
Frame = +1
Query: 238 ENERKICARKRKRKKRKRNRKRKERKFHGKKRRNK 272
+ +R +K +RKK +R +K++++K KKRR K
Sbjct: 49 DQQRSQRRKKSQRKKNQRKKKKRKKKQRRKKRRKK 153
Score = 29.6 bits (65), Expect = 2.3
Identities = 14/31 (45%), Positives = 22/31 (70%), Gaps = 1/31 (3%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRK-KRKRNRKRKER 262
KRR + ++K ++RK+K KRK+ RKRK +
Sbjct: 157 KRRKNQRKKKRKRKRRKKKGKRKKRRKRKSQ 249
>TC81146 similar to GP|13543783|gb|AAH06040.1 Unknown (protein for MGC:7642)
{Mus musculus}, partial (77%)
Length = 1023
Score = 38.1 bits (87), Expect = 0.006
Identities = 17/41 (41%), Positives = 26/41 (62%)
Frame = +2
Query: 237 KENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKKCHCH 277
K +RK RK +++K+K RKR+ +K KK+R +K H H
Sbjct: 575 KSLKRKKKPRKMEKRKKKERRKRERKKMERKKKRRRKKHPH 697
Score = 36.2 bits (82), Expect = 0.024
Identities = 22/44 (50%), Positives = 29/44 (65%)
Frame = +2
Query: 235 RSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKKCHCHQ 278
+ K+ RK+ RK+K ++RKR RK+ ERK KKRR KK HQ
Sbjct: 584 KRKKKPRKMEKRKKK-ERRKRERKKMERK---KKRRRKKHPHHQ 703
Score = 30.8 bits (68), Expect = 1.0
Identities = 14/31 (45%), Positives = 20/31 (64%)
Frame = +2
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRKERK 263
K R E +K RKR+RKK +R +KR+ +K
Sbjct: 596 KPRKMEKRKKKERRKRERKKMERKKKRRRKK 688
>TC93295 weakly similar to GP|5821157|dbj|BAA83720.1 GAMP {Procambarus
clarkii}, partial (11%)
Length = 504
Score = 35.0 bits (79), Expect = 0.054
Identities = 15/35 (42%), Positives = 22/35 (62%)
Frame = +1
Query: 234 RRSKENERKICARKRKRKKRKRNRKRKERKFHGKK 268
+ K E+K +KRK +KRK+ +K K+ K GKK
Sbjct: 334 KEKKRKEKKRKGKKRKERKRKKRKKNKKNKKKGKK 438
Score = 32.7 bits (73), Expect = 0.27
Identities = 16/37 (43%), Positives = 24/37 (64%)
Frame = +1
Query: 237 KENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
KE +RK +KRK KKRK +++K +K K++ KK
Sbjct: 334 KEKKRK--EKKRKGKKRKERKRKKRKKNKKNKKKGKK 438
Score = 30.8 bits (68), Expect = 1.0
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 11/52 (21%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRKKRKR-----------NRKRKERKFHGKKRRNKK 273
+ R K+N K K+ R K+ R +KRKE+K GKKR+ +K
Sbjct: 235 QNRRKQNRMKQTRTKQTRTKQTRMK*NWM*KWMKEKKRKEKKRKGKKRKERK 390
>TC83583 similar to PIR|T06379|T06379 SAR DNA-binding protein 2 - garden
pea, partial (33%)
Length = 852
Score = 35.0 bits (79), Expect = 0.054
Identities = 16/40 (40%), Positives = 26/40 (65%)
Frame = +3
Query: 234 RRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
+R K +RK ++RKRK+R + + ++RK KKRR K+
Sbjct: 402 KRKKRRKRKKRRKRRKRKRRLKMLRNQKRKLLRKKRRRKR 521
Score = 33.9 bits (76), Expect = 0.12
Identities = 14/31 (45%), Positives = 23/31 (74%)
Frame = +3
Query: 243 ICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
+ +++KR+KRK+ RKR++RK K RN+K
Sbjct: 393 LLTKRKKRRKRKKRRKRRKRKRRLKMLRNQK 485
>CA859131 weakly similar to GP|23496809|gb| 101 kd malaria antigen
{Plasmodium falciparum 3D7}, partial (4%)
Length = 395
Score = 35.0 bits (79), Expect = 0.054
Identities = 15/35 (42%), Positives = 22/35 (62%)
Frame = +2
Query: 234 RRSKENERKICARKRKRKKRKRNRKRKERKFHGKK 268
+ K E+K +KRK +KRK+ +K K+ K GKK
Sbjct: 116 KEKKRKEKKRKGKKRKERKRKKRKKNKKNKKKGKK 220
Score = 32.7 bits (73), Expect = 0.27
Identities = 16/37 (43%), Positives = 24/37 (64%)
Frame = +2
Query: 237 KENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
KE +RK +KRK KKRK +++K +K K++ KK
Sbjct: 116 KEKKRK--EKKRKGKKRKERKRKKRKKNKKNKKKGKK 220
Score = 30.8 bits (68), Expect = 1.0
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 11/52 (21%)
Frame = +2
Query: 233 KRRSKENERKICARKRKRKKRKR-----------NRKRKERKFHGKKRRNKK 273
+ R K+N K K+ R K+ R +KRKE+K GKKR+ +K
Sbjct: 17 QNRRKQNRMKQTRTKQTRTKQTRMK*NWM*KWMKEKKRKEKKRKGKKRKERK 172
Score = 29.6 bits (65), Expect = 2.3
Identities = 13/39 (33%), Positives = 23/39 (58%)
Frame = +2
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRN 271
++ K +K RKRK++K+ + K+K +K KR+N
Sbjct: 128 RKEKKRKGKKRKERKRKKRKKNKKNKKKGKKT*MTKRKN 244
>CA990769 weakly similar to GP|21594005|gb| unknown {Arabidopsis thaliana},
partial (20%)
Length = 488
Score = 35.0 bits (79), Expect = 0.054
Identities = 16/44 (36%), Positives = 26/44 (58%)
Frame = +2
Query: 234 RRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKKCHCH 277
RRSK +K ++ K++KR+R KRK +K +K N + H +
Sbjct: 212 RRSKRKRKKKRQKEEKKRKRRRKAKRKSQKKRKRKSLNHQNHVY 343
Score = 32.7 bits (73), Expect = 0.27
Identities = 10/41 (24%), Positives = 30/41 (72%)
Frame = +2
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
+++++ N+R ++++++KKR++ K+++R+ K++ KK
Sbjct: 182 RKQNRNNQRLRRSKRKRKKKRQKEEKKRKRRRKAKRKSQKK 304
Score = 28.5 bits (62), Expect = 5.1
Identities = 13/34 (38%), Positives = 23/34 (67%), Gaps = 1/34 (2%)
Frame = +2
Query: 233 KRRSKENERKICARKRKRK-KRKRNRKRKERKFH 265
++R K+ +++ RKR+RK KRK +KRK + +
Sbjct: 224 RKRKKKRQKEEKKRKRRRKAKRKSQKKRKRKSLN 325
>TC83887 weakly similar to GP|13310891|gb|AAK13153.2 hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (8%)
Length = 778
Score = 34.7 bits (78), Expect = 0.071
Identities = 18/41 (43%), Positives = 29/41 (69%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
K +KENERK RK +R K++++R+ +ER+ K+RR K+
Sbjct: 250 KEEAKENERK---RKEERAKKEKDREERERR-KSKQRREKE 360
>TC77604 homologue to PIR|T06377|T06377 SAR DNA-binding protein-1 - garden
pea, partial (85%)
Length = 1625
Score = 33.9 bits (76), Expect = 0.12
Identities = 14/31 (45%), Positives = 23/31 (74%)
Frame = +1
Query: 243 ICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
+ +++KR+KRK+ RKR++RK K RN+K
Sbjct: 1519 LLTKRKKRRKRKKRRKRRKRKRRLKMLRNQK 1611
Score = 28.9 bits (63), Expect = 3.9
Identities = 10/25 (40%), Positives = 19/25 (76%)
Frame = +1
Query: 246 RKRKRKKRKRNRKRKERKFHGKKRR 270
RKRK+++++R RKR+ + +KR+
Sbjct: 1543 RKRKKRRKRRKRKRRLKMLRNQKRK 1617
>TC88253 weakly similar to GP|15810597|gb|AAL07186.1 unknown protein
{Arabidopsis thaliana}, partial (51%)
Length = 1122
Score = 33.5 bits (75), Expect = 0.16
Identities = 17/42 (40%), Positives = 24/42 (56%)
Frame = +1
Query: 232 CKRRSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
C R K + +KR++K+RK+ RKRK K KK+ KK
Sbjct: 661 CSRGLKSKLQL*RRKKRRKKRRKKKRKRKRSKKKRKKKVKKK 786
Score = 30.4 bits (67), Expect = 1.3
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRKERK 263
K+R K+ +K RKR +KKRK+ K+K ++
Sbjct: 703 KKRRKKRRKKKRKRKRSKKKRKKKVKKKTKE 795
Score = 29.6 bits (65), Expect = 2.3
Identities = 14/27 (51%), Positives = 22/27 (80%)
Frame = +1
Query: 234 RRSKENERKICARKRKRKKRKRNRKRK 260
+R+++ ERK ++++RKKRKR RKRK
Sbjct: 112 KRTRKKERKKKPKRKRRKKRKR-RKRK 189
Score = 28.9 bits (63), Expect = 3.9
Identities = 11/27 (40%), Positives = 20/27 (73%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRKKRKRNRKR 259
KRR K+ +RK +KRK+K +K+ +++
Sbjct: 718 KRRKKKRKRKRSKKKRKKKVKKKTKEK 798
>TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsis
thaliana}, partial (32%)
Length = 1558
Score = 33.1 bits (74), Expect = 0.21
Identities = 14/38 (36%), Positives = 23/38 (59%)
Frame = +1
Query: 235 RSKENERKICARKRKRKKRKRNRKRKERKFHGKKRRNK 272
R K R + +K++ KKRK+ +++K K KKRR +
Sbjct: 721 RKKLKRRNVSVKKKRPKKRKKEKRKKNVKRRRKKRRKE 834
Score = 32.0 bits (71), Expect = 0.46
Identities = 15/42 (35%), Positives = 29/42 (68%), Gaps = 2/42 (4%)
Frame = +1
Query: 234 RRSKENERKICARKRKRKKRKRN--RKRKERKFHGKKRRNKK 273
+R + +K +KRK++KRK+N R+RK+R+ +++N+K
Sbjct: 733 KRRNVSVKKKRPKKRKKEKRKKNVKRRRKKRRKENVRKKNQK 858
>TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (9%)
Length = 908
Score = 32.7 bits (73), Expect = 0.27
Identities = 20/49 (40%), Positives = 26/49 (52%), Gaps = 10/49 (20%)
Frame = +3
Query: 233 KRRSKENERK----------ICARKRKRKKRKRNRKRKERKFHGKKRRN 271
KRR K+ RK + R R+R+KRKR R+ K+ K KRRN
Sbjct: 645 KRRRKKKRRKRMLRVKKKMVMKRRIRRRRKRKRRRREKKTKTRMVKRRN 791
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 32.7 bits (73), Expect = 0.27
Identities = 20/49 (40%), Positives = 26/49 (52%), Gaps = 10/49 (20%)
Frame = +1
Query: 233 KRRSKENERK----------ICARKRKRKKRKRNRKRKERKFHGKKRRN 271
KRR K+ RK + R R+R+KRKR R+ K+ K KRRN
Sbjct: 253 KRRRKKKRRKRMLRVKKKMVMKRRIRRRRKRKRRRREKKTKTRMVKRRN 399
>AJ500818 weakly similar to GP|10435304|dbj unnamed protein product {Homo
sapiens}, partial (12%)
Length = 457
Score = 32.7 bits (73), Expect = 0.27
Identities = 23/41 (56%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Frame = -1
Query: 237 KENERKICARKR---KRKKRKRN-RKRKERKFHGKKRRNKK 273
K N RK ARKR KR RKRN RKR RK + +KR KK
Sbjct: 175 KRNVRKRNARKRNARKRNARKRNVRKRNVRKRNVRKRNVKK 53
Score = 32.7 bits (73), Expect = 0.27
Identities = 22/41 (53%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Frame = -1
Query: 237 KENERKICARKR---KRKKRKRN-RKRKERKFHGKKRRNKK 273
K N RK ARKR KR RKRN RKR RK + +KR +K
Sbjct: 220 KRNVRKRNARKRNARKRNVRKRNARKRNARKRNARKRNVRK 98
Score = 32.3 bits (72), Expect = 0.35
Identities = 22/41 (53%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Frame = -1
Query: 237 KENERKICARKR---KRKKRKRN-RKRKERKFHGKKRRNKK 273
K N RK ARKR KR RKRN RKR RK + +KR +K
Sbjct: 205 KRNARKRNARKRNVRKRNARKRNARKRNARKRNVRKRNVRK 83
Score = 32.3 bits (72), Expect = 0.35
Identities = 21/41 (51%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Frame = -1
Query: 237 KENERKICARKR---KRKKRKRN-RKRKERKFHGKKRRNKK 273
+ N RK ARKR KR RKRN RKR RK + +KR +K
Sbjct: 250 RRNARKRNARKRNVRKRNARKRNARKRNVRKRNARKRNARK 128
Score = 31.2 bits (69), Expect = 0.78
Identities = 22/37 (59%), Positives = 23/37 (61%), Gaps = 4/37 (10%)
Frame = -1
Query: 237 KENERKICARKR---KRKKRKRN-RKRKERKFHGKKR 269
K N RK ARKR KR RKRN RKR RK + KKR
Sbjct: 160 KRNARKRNARKRNARKRNVRKRNVRKRNVRKRNVKKR 50
Score = 30.0 bits (66), Expect = 1.7
Identities = 19/41 (46%), Positives = 26/41 (63%), Gaps = 4/41 (9%)
Frame = -1
Query: 237 KENERKICARKR---KRKKRKRN-RKRKERKFHGKKRRNKK 273
+ N+++ ARKR KR RKRN RKR RK + +KR +K
Sbjct: 265 RRNDKRRNARKRNARKRNVRKRNARKRNARKRNVRKRNARK 143
Score = 28.1 bits (61), Expect = 6.6
Identities = 20/41 (48%), Positives = 24/41 (57%), Gaps = 4/41 (9%)
Frame = -1
Query: 237 KENERKICARKR---KRKKRKRN-RKRKERKFHGKKRRNKK 273
K N RK RKR KR RKRN +KR K + +KR +KK
Sbjct: 130 KRNARKRNVRKRNVRKRNVRKRNVKKRNV*KRNVRKRNDKK 8
>AJ388952 similar to PIR|T04526|T045 hypothetical protein F16A16.160 -
Arabidopsis thaliana, partial (24%)
Length = 591
Score = 32.7 bits (73), Expect = 0.27
Identities = 15/24 (62%), Positives = 18/24 (74%)
Frame = -1
Query: 245 ARKRKRKKRKRNRKRKERKFHGKK 268
+R+RKRKK+KR RKRK K KK
Sbjct: 282 SRRRKRKKKKRKRKRKREKEMKKK 211
>BG644716 homologue to GP|7298552|gb| brat gene product {Drosophila
melanogaster}, partial (2%)
Length = 339
Score = 32.3 bits (72), Expect = 0.35
Identities = 15/19 (78%), Positives = 15/19 (78%)
Frame = +1
Query: 245 ARKRKRKKRKRNRKRKERK 263
ARKRKRKKRK RKRK K
Sbjct: 268 ARKRKRKKRKTKRKRKNMK 324
>AA660209 similar to GP|6642637|gb|A unknown protein {Arabidopsis thaliana},
partial (4%)
Length = 544
Score = 32.3 bits (72), Expect = 0.35
Identities = 20/42 (47%), Positives = 28/42 (66%), Gaps = 4/42 (9%)
Frame = +2
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRK----ERKFHGKKRR 270
KR++ E ++ RKRKRK+RKR+RK K E+K K+RR
Sbjct: 2 KRKTAEKKK---LRKRKRKERKRSRKIKMINEEKKXKIKRRR 118
>AJ503819
Length = 294
Score = 32.3 bits (72), Expect = 0.35
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +1
Query: 233 KRRSKENERKICARKRKRKKRKRNRKRKERK 263
KR +KE+E + C R+ K+K + + K KER+
Sbjct: 172 KRATKEDEEEDCERRMKQKLKSKGSKEKERR 264
>AW560664 similar to GP|23495077|gb hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 545
Score = 32.0 bits (71), Expect = 0.46
Identities = 13/30 (43%), Positives = 22/30 (73%)
Frame = -1
Query: 234 RRSKENERKICARKRKRKKRKRNRKRKERK 263
+R+K +K RK+KRKK+++ +KRK+ K
Sbjct: 383 QRTKGKRKKNQRRKKKRKKKRKEKKRKQAK 294
Score = 31.2 bits (69), Expect = 0.78
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = -1
Query: 240 ERKICARKRKRKKRKRNRKRKERKFHGKKRRNKK 273
ERK K KRKK +R +K++++K KKR+ K
Sbjct: 395 ERK*QRTKGKRKKNQRRKKKRKKKRKEKKRKQAK 294
Score = 27.7 bits (60), Expect = 8.7
Identities = 17/57 (29%), Positives = 27/57 (46%), Gaps = 9/57 (15%)
Frame = -2
Query: 234 RRSKENERKICARKRKRKKRKRNRKRKERK---------FHGKKRRNKKCHCHQAAP 281
+ + ERKI KRK KK + +KRKE K G++ + + +C + P
Sbjct: 382 KEQRGKERKIKEEKRKEKK--KGKKRKENKQSCSPSWHHIEGERPKTQSFYCIKIIP 218
Score = 27.7 bits (60), Expect = 8.7
Identities = 11/27 (40%), Positives = 20/27 (73%)
Frame = -1
Query: 233 KRRSKENERKICARKRKRKKRKRNRKR 259
K + K+N+R+ RK+KRK++KR + +
Sbjct: 374 KGKRKKNQRRKKKRKKKRKEKKRKQAK 294
>TC79244 weakly similar to PIR|B86193|B86193 hypothetical protein [imported]
- Arabidopsis thaliana, partial (57%)
Length = 1114
Score = 31.6 bits (70), Expect = 0.60
Identities = 19/39 (48%), Positives = 24/39 (60%), Gaps = 1/39 (2%)
Frame = -2
Query: 233 KRRSKENERKICARKRKRKKRKRN-RKRKERKFHGKKRR 270
K R KE E KRK KKRK++ RK +ER G++RR
Sbjct: 129 KNRGKEEEEG----KRKEKKRKKDKRKNRERD*RGRERR 25
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.385 0.173 0.749
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,553,674
Number of Sequences: 36976
Number of extensions: 191358
Number of successful extensions: 4967
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 2222
Number of HSP's successfully gapped in prelim test: 182
Number of HSP's that attempted gapping in prelim test: 2351
Number of HSP's gapped (non-prelim): 2777
length of query: 421
length of database: 9,014,727
effective HSP length: 99
effective length of query: 322
effective length of database: 5,354,103
effective search space: 1724021166
effective search space used: 1724021166
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC131249.9


