

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130803.17 + phase: 0
(532 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
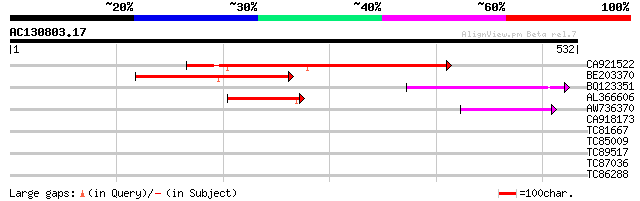
Score E
Sequences producing significant alignments: (bits) Value
CA921522 251 5e-67
BE203370 151 5e-37
BQ123351 109 2e-24
AL366606 107 1e-23
AW736370 47 1e-05
CA918173 weakly similar to GP|11034537|dbj putative glucosyl tra... 30 3.0
TC81667 similar to GP|8777488|dbj|BAA97068.1 contains similarity... 30 3.0
TC85009 similar to GP|14531294|gb|AAK66160.1 pectate lyase B {Fr... 29 3.9
TC89517 weakly similar to PIR|A85475|A85475 hypothetical protein... 29 5.1
TC87036 similar to GP|11138330|gb|AAG31326.1 putative serine/thr... 28 6.7
TC86288 similar to GP|2407790|gb|AAB70660.1| grr1 {Glycine max},... 28 8.8
>CA921522
Length = 767
Score = 251 bits (641), Expect = 5e-67
Identities = 128/256 (50%), Positives = 167/256 (65%), Gaps = 8/256 (3%)
Frame = +3
Query: 167 KTSDVSNHLITKSHIRGFASKYLLEQANLSTTRQDTL------EAILALLIYGLILFPNL 220
+T +V HLIT+ GF++ +L E+ TT D + +ILALL+YGL+LFP+L
Sbjct: 3 ETDEVKTHLITRGKFLGFSTDFLYER----TTFFDKMGVAYAFNSILALLVYGLVLFPSL 170
Query: 221 DNFVDMNAIEVFHSKNPVPTLLADTYHAIHDRTLKGRGYILCCISLLYRWFISHLPSS-- 278
DNFVD+ AI++F S+NPVPTLL DTY +IH RT GRG ILCC LLYRW SHLP +
Sbjct: 171 DNFVDIKAIQIFLSRNPVPTLLGDTYLSIHRRTQAGRGTILCCAQLLYRWITSHLPRTPR 350
Query: 279 FHDNSENWSYSQRIMALTPNEVVWITPAAQAKEIIMGCGDFLNVPLLGTRGGINYNPELA 338
F N EN +S+R+M+LTP EVVW II CG F NVPLLG GGI+YNP LA
Sbjct: 351 FTTNPENLLWSKRLMSLTPAEVVWYDRVYDKGTIIDSCGKFANVPLLGMEGGISYNPTLA 530
Query: 339 MRQFGFPMKSKPINLATSPEFFFYTNAPTGQRKAFMDAWSKVRRKSVKHLGVRSGVAHEA 398
QFG+PM+ KP+++ ++ + TG R+ + AW +RR+ LG ++G +
Sbjct: 531 RHQFGYPMERKPLSIYLENVYYLNADDSTGMREHVVRAWHTIRRRDKDQLGKKTGAISSS 710
Query: 399 YTQWVIDRAEEIGMPY 414
YTQWVIDRA +IGMPY
Sbjct: 711 YTQWVIDRAVQIGMPY 758
>BE203370
Length = 454
Score = 151 bits (382), Expect = 5e-37
Identities = 79/150 (52%), Positives = 101/150 (66%), Gaps = 2/150 (1%)
Frame = +1
Query: 119 CFTFPDFQLVPTLEAYSNLVGLPIAEKTPFTGPGTSLTPLVIAKDLYLKTSDVSNHLITK 178
CFTFPD+QL PTLE YS LVG P+ +K PFTG IA L+L+TS + L K
Sbjct: 4 CFTFPDYQLAPTLEEYSYLVG*PVLDKVPFTGFEPIPKAATIADALHLETSLIKAKLTIK 183
Query: 179 SHIRGFASKYLLEQAN--LSTTRQDTLEAILALLIYGLILFPNLDNFVDMNAIEVFHSKN 236
++ +K+L +QA+ D +ILALLIYGL+LFPN+DN+VD++AI++F +KN
Sbjct: 184 GNLPSLPTKFLYQQASDFAKVDNVDAFYSILALLIYGLVLFPNVDNYVDIHAIQMFLTKN 363
Query: 237 PVPTLLADTYHAIHDRTLKGRGYILCCISL 266
PVPTLLAD YH+IHDR GRG IL C L
Sbjct: 364 PVPTLLADMYHSIHDRNQVGRGAILGCAPL 453
>BQ123351
Length = 725
Score = 109 bits (273), Expect = 2e-24
Identities = 61/153 (39%), Positives = 93/153 (59%)
Frame = +2
Query: 373 FMDAWSKVRRKSVKHLGVRSGVAHEAYTQWVIDRAEEIGMPYPAMRYVSSSTPSIPLPLL 432
F+ AW V ++ LG RS + +E+YT+WVIDRA + G+P+P R SS+T S LP+
Sbjct: 8 FIQAWRAVHKREKGQLGRRS*LVYESYTKWVIDRAAKNGIPFPLQRLPSSTTLSPSLPMT 187
Query: 433 PATQDMYQEHLAMESHEKQVWKARYNQAENLIMTLDGRDEQKTHENLMLKKELAKARKEL 492
T++ Q LA + EK W+ RY +AE+ I TL G+ EQK HE L +++++ + L
Sbjct: 188 SKTKEEAQYLLAEMTREKDTWRIRYMEAESEIGTLRGQVEQKDHELLKMRQQMIERDDLL 367
Query: 493 EEKDELLLRDSKRAKGRRDFFARYCGSDSESDD 525
+EKD+LL + + K R D + G DS+ +D
Sbjct: 368 QEKDKLLKKHITK-KQRMDSMDLFDGPDSDFED 463
>AL366606
Length = 393
Score = 107 bits (267), Expect = 1e-23
Identities = 58/78 (74%), Positives = 59/78 (75%), Gaps = 6/78 (7%)
Frame = -1
Query: 205 AILALLIYGLILFPNLDNFVDMNAIEVFHSKNPVPTLLADTYHAIHDRTLKGRGYILCCI 264
AI L GLILFPNLDNFVDMNAIEVFHSKNPVPTLLADTYHAIHDRTLKGRGYIL
Sbjct: 381 AIXLCLSTGLILFPNLDNFVDMNAIEVFHSKNPVPTLLADTYHAIHDRTLKGRGYILWLH 202
Query: 265 SLL------YRWFISHLP 276
L +R F S LP
Sbjct: 201 ILFSNRVVYFRTFRSSLP 148
>AW736370
Length = 488
Score = 47.4 bits (111), Expect = 1e-05
Identities = 29/90 (32%), Positives = 43/90 (47%)
Frame = -2
Query: 424 TPSIPLPLLPATQDMYQEHLAMESHEKQVWKARYNQAENLIMTLDGRDEQKTHENLMLKK 483
TP+ PLP+ T+ YQE L E WK Y + + T+ G EQ+ +++
Sbjct: 481 TPAPPLPMTFDTEKEYQERLTEAEREIHRWKREYQKKDQDYETVMGLLEQEAYDSR*KDV 302
Query: 484 ELAKARKELEEKDELLLRDSKRAKGRRDFF 513
+AK + ++E D L R R K R D F
Sbjct: 301 IIAKLNERIKENDAALDRIPGRKKRRMDLF 212
>CA918173 weakly similar to GP|11034537|dbj putative glucosyl transferase
{Oryza sativa (japonica cultivar-group)}, partial (19%)
Length = 769
Score = 29.6 bits (65), Expect = 3.0
Identities = 13/22 (59%), Positives = 18/22 (81%)
Frame = -3
Query: 105 LVHTLVQFYDPSFRCFTFPDFQ 126
L+HTLVQ + PS RCF FP+++
Sbjct: 113 LIHTLVQKW-PS*RCFPFPNYE 51
>TC81667 similar to GP|8777488|dbj|BAA97068.1 contains similarity to SF16
protein~gene_id:K15M2.19 {Arabidopsis thaliana}, partial
(34%)
Length = 1216
Score = 29.6 bits (65), Expect = 3.0
Identities = 17/59 (28%), Positives = 25/59 (41%)
Frame = -2
Query: 329 GGINYNPELAMRQFGFPMKSKPINLATSPEFFFYTNAPTGQRKAFMDAWSKVRRKSVKH 387
GG+N NP FP L +P F+T A K + W + R+S++H
Sbjct: 780 GGLNLNPN-------FPRTRCNPTLHPAPTPAFFTKAIRSLTKILVTVWPPLMRESIRH 625
>TC85009 similar to GP|14531294|gb|AAK66160.1 pectate lyase B {Fragaria x
ananassa}, partial (55%)
Length = 914
Score = 29.3 bits (64), Expect = 3.9
Identities = 12/51 (23%), Positives = 26/51 (50%)
Frame = +3
Query: 360 FFYTNAPTGQRKAFMDAWSKVRRKSVKHLGVRSGVAHEAYTQWVIDRAEEI 410
F +TNA ++ K++ +S+ + + ++H+A + +D EEI
Sbjct: 138 FAFTNAIDSNESRVVNGEEKLKMQSLNNSSMAESLSHDAINEHAVDNPEEI 290
>TC89517 weakly similar to PIR|A85475|A85475 hypothetical protein AT4g40080
[imported] - Arabidopsis thaliana, partial (46%)
Length = 1177
Score = 28.9 bits (63), Expect = 5.1
Identities = 23/87 (26%), Positives = 34/87 (38%), Gaps = 1/87 (1%)
Frame = -3
Query: 217 FPNLDNFVDMNAIEVFHSKNPVPTLLADTYHAIHDRTLKGRGYILCCISLLYRWFISHLP 276
F L F++ N H NP+ +H IH Y+ + L W I P
Sbjct: 764 FLPLPKFINSNQ-NFIHHHNPI------IFHQIH--------YLNNHLVLFTHWSIRFFP 630
Query: 277 SSFHDNSENWSYS-QRIMALTPNEVVW 302
+SFH + S+S Q + N +W
Sbjct: 629 NSFHQGQQRISFS*QITIGKIRNSFLW 549
>TC87036 similar to GP|11138330|gb|AAG31326.1 putative serine/threonine
kinase GDBrPK {Vitis vinifera}, partial (66%)
Length = 1212
Score = 28.5 bits (62), Expect = 6.7
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +2
Query: 270 WFISHLPSSFHDNSENWS 287
WF+ +LP F ++ ENWS
Sbjct: 692 WFLKNLPLEFMEDGENWS 745
>TC86288 similar to GP|2407790|gb|AAB70660.1| grr1 {Glycine max}, partial
(84%)
Length = 2900
Score = 28.1 bits (61), Expect = 8.8
Identities = 15/40 (37%), Positives = 23/40 (57%)
Frame = -2
Query: 470 RDEQKTHENLMLKKELAKARKELEEKDELLLRDSKRAKGR 509
+ + K +L KE +K R EK+E LLR+ K+ +GR
Sbjct: 178 KKQHKVKSEELLTKETSKKR----EKEEELLREKKKKRGR 71
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,233,197
Number of Sequences: 36976
Number of extensions: 211249
Number of successful extensions: 1141
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1135
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1138
length of query: 532
length of database: 9,014,727
effective HSP length: 101
effective length of query: 431
effective length of database: 5,280,151
effective search space: 2275745081
effective search space used: 2275745081
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130803.17


