

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
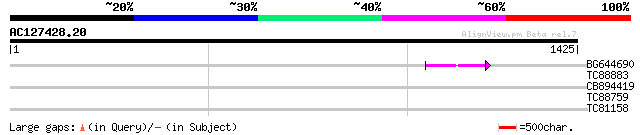
Query= AC127428.20 + phase: 0 /pseudo
(1425 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 47 4e-05
TC88883 similar to PIR|B84247|B84247 proline permease [imported]... 35 0.16
CB894419 weakly similar to GP|10177935|db copia-type polyprotein... 32 1.3
TC88759 similar to GP|16588826|gb|AAL26909.1 dehydration-respons... 30 6.7
TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidops... 30 6.7
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 47.4 bits (111), Expect = 4e-05
Identities = 47/163 (28%), Positives = 68/163 (40%)
Frame = -1
Query: 1045 YQ*LNLKQLMKHSQMMNGY*QCRKS*ISFKEPMCGI*YPNPSRRTLLEQNGYSETS*MRK 1104
Y L+ + L KH M G C+K+ IS KE G + + + LE G+ ETS
Sbjct: 599 YHLLSPRMLKKHCVMQTGSILCKKNSISLKEVRYGTWFLDLKAKQ*LELGGFLETSLSDY 420
Query: 1105 VK*PETKPDLLHKDIVNKKALIILKLLLQLQDWKQSGYFYLCNQSWNNNISNRC*ECFS* 1164
K* + + N+ + Q+ + C + N C EC *
Sbjct: 419 *K*IQVGGARIQSKRRNR---L**GFFTCCQNGSY*NFNSFCCIHGVQAVPNGCEECIY* 249
Query: 1165 WCH*RRSICQTTSWF*GS*IS*PCL*A*EITIWLETSSQSLL* 1207
W R +CQ TSW * + C+ TIW E SS+S++*
Sbjct: 248 WRSQRGGVCQATSWI*RCRGTKSCVQIE*DTIWSEASSKSMV* 120
>TC88883 similar to PIR|B84247|B84247 proline permease [imported] -
Halobacterium sp. NRC-1, partial (4%)
Length = 676
Score = 35.4 bits (80), Expect = 0.16
Identities = 14/29 (48%), Positives = 19/29 (65%)
Frame = -2
Query: 989 HLMKMLLKIFQKTLSKQFHQSLNTNHHIL 1017
HL +L K+ Q LS ++HQ L T HH+L
Sbjct: 312 HLQLLLTKLLQHQLSAKYHQQLQTKHHLL 226
>CB894419 weakly similar to GP|10177935|db copia-type polyprotein {Arabidopsis
thaliana}, partial (2%)
Length = 170
Score = 32.3 bits (72), Expect = 1.3
Identities = 14/47 (29%), Positives = 26/47 (54%)
Frame = +2
Query: 1303 EEVQTRRLQSDEHSNASYLHLKQRR*WNKSRPEAV*RYDWFFVIFHC 1349
EE+Q +++ H+ + +R+ W K R ++ ++DW F IF C
Sbjct: 26 EEIQDGAFKANFHAG*RKVEADKRKRW*KGRLNSLQKFDWKFEIFDC 166
>TC88759 similar to GP|16588826|gb|AAL26909.1 dehydration-responsive protein
RD22 {Prunus persica}, partial (14%)
Length = 1111
Score = 30.0 bits (66), Expect = 6.7
Identities = 21/66 (31%), Positives = 27/66 (40%), Gaps = 2/66 (3%)
Frame = +1
Query: 1269 NEHDGRTKVLSRNSNQSKQRWSLCSSNKIHKGASEEVQTRRLQSDEHSNASY--LHLKQR 1326
+ HD K R S KQ CS + S QT S H + Y LHLK+
Sbjct: 46 SNHDFLWKQSKRISWNRKQTLGYCSEQNCY**TSNRTQTSCT*SQSHGSFCYGVLHLKRL 225
Query: 1327 R*WNKS 1332
+ W K+
Sbjct: 226 KSWEKN 243
>TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidopsis
thaliana}, partial (6%)
Length = 890
Score = 30.0 bits (66), Expect = 6.7
Identities = 14/47 (29%), Positives = 32/47 (67%), Gaps = 1/47 (2%)
Frame = +2
Query: 931 LMIESLEVKLQSKVKVLQVN-KILKMHQNLI*HSTLKRVQKLNQIQK 976
+ I+ +EVK++ KVKVL +N ++L +H+ + + +K + + +++K
Sbjct: 401 MRIKMMEVKVKVKVKVLSLNLRVLLLHRRRLVLTVMKTTKTMQKMKK 541
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.377 0.169 0.647
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,705,826
Number of Sequences: 36976
Number of extensions: 588047
Number of successful extensions: 9419
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1664
Number of HSP's successfully gapped in prelim test: 351
Number of HSP's that attempted gapping in prelim test: 7632
Number of HSP's gapped (non-prelim): 2387
length of query: 1425
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1317
effective length of database: 5,021,319
effective search space: 6613077123
effective search space used: 6613077123
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC127428.20


