

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.7 - phase: 0
(339 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
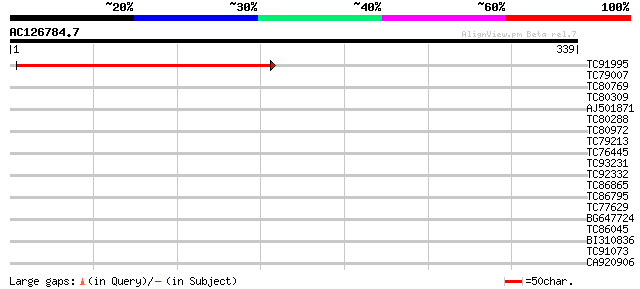
Sequences producing significant alignments: (bits) Value
TC91995 similar to GP|21554881|gb|AAM63718.1 unknown {Arabidopsi... 307 4e-84
TC79007 similar to GP|20260516|gb|AAM13156.1 unknown protein {Ar... 32 0.46
TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 29 2.3
TC80309 similar to GP|20804963|dbj|BAB92640. hypothetical protei... 29 2.3
AJ501871 homologue to GP|16197625|gb|A anaphase promoting comple... 29 3.0
TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein... 29 3.0
TC80972 weakly similar to GP|23496972|gb|AAN36523.1 hypothetical... 28 5.1
TC79213 similar to GP|15706274|emb|CAC69997. HMG I/Y like protei... 28 5.1
TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - so... 28 5.1
TC93231 similar to PIR|T49293|T49293 hypothetical protein T16L24... 28 5.1
TC92332 similar to GP|18377648|gb|AAL66974.1 unknown protein {Ar... 28 5.1
TC86865 similar to PIR|E96497|E96497 probable bZIP transcription... 28 5.1
TC86795 similar to GP|4105180|gb|AAD02288.1| plastoglobule assoc... 28 5.1
TC77629 similar to GP|6648962|gb|AAF21309.1| seed maturation pro... 28 6.6
BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}... 28 6.6
TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Ar... 28 6.6
BI310836 homologue to PIR|T12621|T126 Dc3 promoter-binding facto... 28 6.6
TC91073 similar to PIR|T52301|T52301 GYMNOS/PICKLE protein [impo... 28 6.6
CA920906 similar to GP|4894170|emb cytochrome P450 {Cicer arieti... 27 8.6
>TC91995 similar to GP|21554881|gb|AAM63718.1 unknown {Arabidopsis
thaliana}, partial (10%)
Length = 540
Score = 307 bits (786), Expect = 4e-84
Identities = 153/155 (98%), Positives = 153/155 (98%)
Frame = +1
Query: 5 ANSEIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQS 64
ANSEIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQS
Sbjct: 76 ANSEIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQS 255
Query: 65 SWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHLEGKC 124
SWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHLEGKC
Sbjct: 256 SWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHLEGKC 435
Query: 125 CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRL 159
CIDLMDGGVECVTVGDDGRINLVT GDSN NYRRL
Sbjct: 436 CIDLMDGGVECVTVGDDGRINLVTXGDSNXNYRRL 540
>TC79007 similar to GP|20260516|gb|AAM13156.1 unknown protein {Arabidopsis
thaliana}, partial (96%)
Length = 1561
Score = 31.6 bits (70), Expect = 0.46
Identities = 32/146 (21%), Positives = 54/146 (36%), Gaps = 11/146 (7%)
Frame = +2
Query: 116 NELHLEGKCCIDLMDGGVECVTVGDDGRIN-LVTVGDSNLNYRRLFDSGGLVSYTSVKWA 174
N LE ++ G ++C+ GR + L+++ D NL L S S A
Sbjct: 413 NSPQLEKITSLETQSGKIKCILWWPTGRHDKLISIDDENLCLWSLDVSKKTAQVQSQDSA 592
Query: 175 SPVEFATGGYGFGLHWWDQRKPGGPVSQFKGN---WDK-------NLNSGIVHSIDIHPS 224
+ +GG WD + + + WD ++ V S+D HP
Sbjct: 593 GMMHKLSGGA------WDPHDMNSVAATSESSLQFWDVRTMKKTLSIECSHVRSVDYHPR 754
Query: 225 RKHTCLAAGSLGTVLAWDLRMQQQPI 250
+ + + A + WD R + PI
Sbjct: 755 KNNILVTAEHESGIRIWDQRKPKVPI 832
>TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (40%)
Length = 1362
Score = 29.3 bits (64), Expect = 2.3
Identities = 18/70 (25%), Positives = 28/70 (39%)
Frame = +3
Query: 174 ASPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAG 233
AS AT + + WD PG + F G+ S V S+D HP++ +
Sbjct: 522 ASMPRLATSSFDKTVRVWDVDNPGYSLRTFTGH------STSVMSLDFHPNKDDLICSCD 683
Query: 234 SLGTVLAWDL 243
G + W +
Sbjct: 684 GDGEIRYWSI 713
>TC80309 similar to GP|20804963|dbj|BAB92640. hypothetical protein~similar
to Arabidopsis thaliana chromosome 3 At3g51270, partial
(3%)
Length = 828
Score = 29.3 bits (64), Expect = 2.3
Identities = 32/92 (34%), Positives = 42/92 (44%), Gaps = 10/92 (10%)
Frame = -2
Query: 10 HRFPQYK-----YIDAVRWLPVLSAFNRFAV---LATSDFDSNLSSIEIHSFKP-NPLSF 60
H FP Y+ I L + F+ F V T+ + +L+ + F P +P S
Sbjct: 395 HCFPPYRSPLPFQIRCFHLLQISPLFHDFHVPHPAPTNVKNQHLNHLLHSGFSPRSPSSL 216
Query: 61 -EFQSSWTSPSPISSIKSSQFLQKSIIASSTS 91
EF S TSPS I SS FL K + ASS S
Sbjct: 215 SEFVSLSTSPSINL*ISSSSFLPKPLAASSLS 120
>AJ501871 homologue to GP|16197625|gb|A anaphase promoting complex subunit
11 {Arabidopsis thaliana}, partial (84%)
Length = 669
Score = 28.9 bits (63), Expect = 3.0
Identities = 15/37 (40%), Positives = 22/37 (58%)
Frame = +2
Query: 45 LSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFL 81
L+ I IH+F+ P F+F+SS + SS SQ+L
Sbjct: 17 LTKIYIHTFEFGPSPFQFKSSSSQTLTSSSENESQYL 127
>TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein T28I19.100
- Arabidopsis thaliana, partial (14%)
Length = 1460
Score = 28.9 bits (63), Expect = 3.0
Identities = 26/94 (27%), Positives = 41/94 (42%)
Frame = -2
Query: 43 SNLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADS 102
S+LSS + S + LSF ++ +S S +S S F S S+S+ SL F+
Sbjct: 760 SSLSSFSVSSSPSSSLSFSSSTTSSSRSLSASDFS*SFSSISTSPPSSSTSSLLLSFSSC 581
Query: 103 TDARLVSEVSVPENELHLEGKCCIDLMDGGVECV 136
T A + S + L C + + V C+
Sbjct: 580 TSAPCLFSSSSCSSLL----SCTLSSLSSSVSCL 491
>TC80972 weakly similar to GP|23496972|gb|AAN36523.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (9%)
Length = 662
Score = 28.1 bits (61), Expect = 5.1
Identities = 19/43 (44%), Positives = 25/43 (57%)
Frame = +3
Query: 59 SFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFAD 101
++ + SS +S S SS SS F S ASS+SS S+H F D
Sbjct: 525 TYPYNSSCSSSSTSSS-SSSSFSSLSSSASSSSSSSIHQDFFD 650
>TC79213 similar to GP|15706274|emb|CAC69997. HMG I/Y like protein {Glycine
max}, partial (16%)
Length = 811
Score = 28.1 bits (61), Expect = 5.1
Identities = 20/73 (27%), Positives = 31/73 (42%), Gaps = 3/73 (4%)
Frame = +1
Query: 27 LSAFNRFAVLATSDFDSNL--SSIEIHSF-KPNPLSFEFQSSWTSPSPISSIKSSQFLQK 83
++A +LAT D + L +++ F +P PL E S P P+ SQ L
Sbjct: 220 VAAIQELEMLATLDLKAPLRDETLQPQQFPEPQPLQQELPSQQLHPQPVPQALPSQQLHP 399
Query: 84 SIIASSTSSGSLH 96
+ + S LH
Sbjct: 400 QPVPQAPPSQQLH 438
>TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - soybean
(fragment), partial (76%)
Length = 821
Score = 28.1 bits (61), Expect = 5.1
Identities = 23/81 (28%), Positives = 34/81 (41%), Gaps = 17/81 (20%)
Frame = +1
Query: 68 SPSPISSIKSSQFLQ-----------------KSIIASSTSSGSLHFLFADSTDARLVSE 110
S SPIS SSQ LQ S + TS+ SLH F+ +T+ +L S
Sbjct: 226 SSSPISHFSSSQLLQIPSTTIAISSTSIPLRESSPTSKVTSTTSLHLCFSTTTNLQLAS- 402
Query: 111 VSVPENELHLEGKCCIDLMDG 131
++++ + K C G
Sbjct: 403 ----QHQITIGDKLCSSFHQG 453
>TC93231 similar to PIR|T49293|T49293 hypothetical protein T16L24.70 -
Arabidopsis thaliana, partial (30%)
Length = 722
Score = 28.1 bits (61), Expect = 5.1
Identities = 29/82 (35%), Positives = 37/82 (44%), Gaps = 6/82 (7%)
Frame = +1
Query: 52 SFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEV 111
SF P P+ SS TSPS S K S FL + + FL+ + VS V
Sbjct: 94 SFSPYPIQLFHSSSSTSPS-FSEDKDSTFLVFFRAIHFLFANNFLFLWLVNAAQIFVSWV 270
Query: 112 SV----PENELHLEGKC--CID 127
SV PE L+L+ + CID
Sbjct: 271 SVIYEFPEKPLNLKLRVLDCID 336
>TC92332 similar to GP|18377648|gb|AAL66974.1 unknown protein {Arabidopsis
thaliana}, partial (34%)
Length = 1164
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = +2
Query: 216 VHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQ 248
V S+D HPS L S G + W+L ++++
Sbjct: 458 VTSLDFHPSHHTLLLVGSSNGEITLWELSLRER 556
>TC86865 similar to PIR|E96497|E96497 probable bZIP transcription factor
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1171
Score = 28.1 bits (61), Expect = 5.1
Identities = 19/62 (30%), Positives = 27/62 (42%)
Frame = +1
Query: 40 DFDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLF 99
DF+S + E+ F S +PSP+S I++ IASS+S H L
Sbjct: 160 DFESLFAEYEVADFLQEDDINPSVSVADTPSPLSEIENLLMSDAEGIASSSSDSDYHKLL 339
Query: 100 AD 101
D
Sbjct: 340 QD 345
>TC86795 similar to GP|4105180|gb|AAD02288.1| plastoglobule associated
protein PG1 precursor {Pisum sativum}, partial (95%)
Length = 1400
Score = 28.1 bits (61), Expect = 5.1
Identities = 29/95 (30%), Positives = 38/95 (39%)
Frame = +2
Query: 29 AFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIAS 88
+FNRFA+ S S L I S P P S S SPSP +S +F I++
Sbjct: 95 SFNRFAMSLLS---STLRVPLIFSQNPKPSSLSSLPSRISPSP----RSHRFPSLRFISA 253
Query: 89 STSSGSLHFLFADSTDARLVSEVSVPENELHLEGK 123
+ G D +D VPE + L K
Sbjct: 254 TGDIGDADIPSTDISDEWGDKSEPVPEAKPSLSSK 358
>TC77629 similar to GP|6648962|gb|AAF21309.1| seed maturation protein PM23
{Glycine max}, partial (76%)
Length = 1741
Score = 27.7 bits (60), Expect = 6.6
Identities = 21/69 (30%), Positives = 35/69 (50%)
Frame = +3
Query: 43 SNLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADS 102
S+ SS+ +H F+P+P SF F ++P+ S FL + +S+S S F S
Sbjct: 126 SSSSSLPLHRFRPSP-SFSFSPLRSNPTK----SPSPFL----VLASSSHDSNDFTSKKS 278
Query: 103 TDARLVSEV 111
+ L+ E+
Sbjct: 279 ALSELIQEI 305
>BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}, partial
(25%)
Length = 757
Score = 27.7 bits (60), Expect = 6.6
Identities = 18/58 (31%), Positives = 27/58 (46%), Gaps = 1/58 (1%)
Frame = +2
Query: 169 TSVKWA-SPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSR 225
T V+++ S AT Y + WD PG + F G +S V S+D HP++
Sbjct: 569 TDVRFSPSMPRLATSSYDKTVRVWDVDNPGYSLRTFTG------HSAAVMSLDFHPNK 724
>TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana, partial (69%)
Length = 1166
Score = 27.7 bits (60), Expect = 6.6
Identities = 36/106 (33%), Positives = 47/106 (43%), Gaps = 1/106 (0%)
Frame = -1
Query: 21 VRWLPVLSAFNRFAVLATSDFDSNLSSIEIHS-FKPNPLSFEFQSSWTSPSPISSIKSSQ 79
V L ++ FN +A F S L E + + + LS SS +S S SS+ SS
Sbjct: 407 VTLLTPVTGFNPSLSIAFRLFFSLLLCFEPSAPARASSLSSSSSSSSSSSSSSSSLSSS- 231
Query: 80 FLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHLEGKCC 125
L SII +S+SSG FL R+VS EG CC
Sbjct: 230 -LTSSIIGASSSSGWGFFL-------RVVSST---------EGICC 144
>BI310836 homologue to PIR|T12621|T126 Dc3 promoter-binding factor-1 - common
sunflower, partial (6%)
Length = 656
Score = 27.7 bits (60), Expect = 6.6
Identities = 21/52 (40%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Frame = +2
Query: 177 VEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIH--PSRK 226
V ATGGYG + GGPVS + N NSG ID++ P RK
Sbjct: 479 VNGATGGYGVAVA--QTMGMGGPVSPVSSDGIGNENSGGQFGIDMNGLPRRK 628
>TC91073 similar to PIR|T52301|T52301 GYMNOS/PICKLE protein [imported] -
Arabidopsis thaliana, partial (14%)
Length = 627
Score = 27.7 bits (60), Expect = 6.6
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = -2
Query: 93 GSLHFLFADSTDARLVSEVSVPENELHLEGKCC 125
G + F+F+D+T + S + N +HL G CC
Sbjct: 233 GKVKFIFSDNTTSLGRSNI---HNNIHLRGPCC 144
>CA920906 similar to GP|4894170|emb cytochrome P450 {Cicer arietinum},
partial (46%)
Length = 792
Score = 27.3 bits (59), Expect = 8.6
Identities = 21/46 (45%), Positives = 27/46 (58%), Gaps = 2/46 (4%)
Frame = +2
Query: 73 SSIKSS--QFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPEN 116
SSIK S L SI + ST+SG + L D + +V+EVSVP N
Sbjct: 536 SSIKCSCPM*LSNSIFSFSTTSGWFNKL--DIAHSNVVAEVSVPAN 667
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,066,301
Number of Sequences: 36976
Number of extensions: 155953
Number of successful extensions: 949
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 937
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 945
length of query: 339
length of database: 9,014,727
effective HSP length: 97
effective length of query: 242
effective length of database: 5,428,055
effective search space: 1313589310
effective search space used: 1313589310
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126784.7


