

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.1 - phase: 0 /pseudo
(790 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
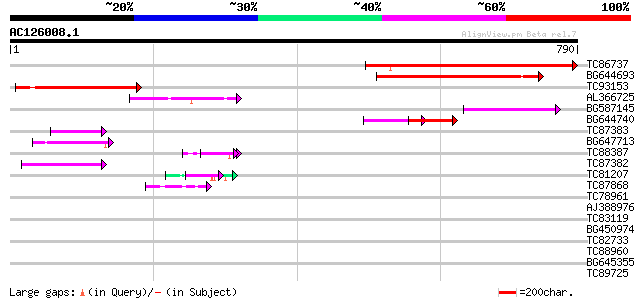
Score E
Sequences producing significant alignments: (bits) Value
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 244 7e-65
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 228 9e-60
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 176 4e-44
AL366725 111 1e-24
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 97 2e-20
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 82 6e-16
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 57 3e-08
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 56 6e-08
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 55 1e-07
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 53 5e-07
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 45 1e-04
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 43 5e-04
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 42 0.001
AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing fact... 39 0.008
TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 38 0.017
BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR p... 38 0.017
TC82733 similar to GP|10177404|dbj|BAB10535. gene_id:K24M7.12~pi... 37 0.038
TC88960 similar to GP|21537297|gb|AAM61638.1 DNA-damage inducibl... 28 0.057
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 36 0.065
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 35 0.15
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 244 bits (624), Expect = 7e-65
Identities = 132/303 (43%), Positives = 196/303 (64%), Gaps = 9/303 (2%)
Frame = +1
Query: 497 VVQEFPEVF-PEDITELPSEREV-EFAIDLVPGTS----PISIAP-YRMSASELGELKKQ 549
V++EFP++F PE ++P+ R + + AI L+P P+ P Y MS EL LKK
Sbjct: 343 VLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKKT 522
Query: 550 LEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQ 609
LE+LL+K FI+ S S GAPVL V+K G +R CVDYR LN +T K+RYPLP I + + +
Sbjct: 523 LEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRR 702
Query: 610 LVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNR 669
+ GA F+K+D+ + +H++R+K ED KTAFRTRYG +E+ V PFG++ AP F Y+N+
Sbjct: 703 VAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYINK 882
Query: 670 IFHPYLDRFVVVFIDDILVYSK-SEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSF 728
H +LD FV +IDD+L+Y+ S+++H +R VL+ L + L KCEF + V +
Sbjct: 883 TLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVTTVKY 1062
Query: 729 LGHVISKG-GISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLALPLTQL 787
+G +++ G G+S DP K+ A+ W P SV ++ G YY+ FI G+S++ PLT+L
Sbjct: 1063VGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEPLTRL 1242
Query: 788 TQK 790
T+K
Sbjct: 1243TRK 1251
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 228 bits (580), Expect = 9e-60
Identities = 121/233 (51%), Positives = 159/233 (67%)
Frame = +2
Query: 512 LPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVL 571
+P E +++F IDL+P +PI I YR++ +L LK QL++LLEK FI+PS+ P G VL
Sbjct: 17 VPPEWKIDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVVVL 196
Query: 572 LVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVK 631
+KKKDG +R+ +DY QLN + IK +YPLP ID+ D L G++ F KIDLR G HQ RV
Sbjct: 197 FLKKKDGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVI 376
Query: 632 AEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSK 691
ED+ KTAFR RYGHYE VM FG +N P FME MNR+F YLD V+VF +DIL+YSK
Sbjct: 377 GEDVPKTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSK 556
Query: 692 SEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSK 744
+E EH HLR+ L+VLK+ LC ++S ++ F HVIS G+ VD +
Sbjct: 557 NENEHENHLRLALKVLKDIGLC-QISYV*ILVEVGFFSLHVISGEGLKVDSKR 712
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 176 bits (445), Expect = 4e-44
Identities = 92/175 (52%), Positives = 116/175 (65%)
Frame = +2
Query: 9 GRDDAAIAQALATMAQVLAQGNENTAANRRNEGEAEERRLDRFLRNNPPTFKGRFDPEGA 68
G DAA+ AL +AQ + Q + + G R L+ FLRN+PPTFKGR+ P+GA
Sbjct: 8 GSSDAALVAALEAVAQAVQQ------LPKVDTGSDGTRMLETFLRNHPPTFKGRYAPDGA 169
Query: 69 QTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWWINAKGRLELGGDVVTWAKFKVEF 128
W++ +ERIFR M + QKV+ THM AEEA+ WWI+ LE VVTWA F+ EF
Sbjct: 170 *KWLKEIERIFRVMQCFETQKVQFGTHMLAEEADDWWISLLPVLEQDDAVVTWAMFRKEF 349
Query: 129 LRKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEELSRFCPYINAEDAMVSKCV 183
L +YFPED+R KKE+EFL LKQG MSV EYAAKF EL+ F P+ +AE A SKC+
Sbjct: 350 LGRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKCI 514
>AL366725
Length = 485
Score = 111 bits (277), Expect = 1e-24
Identities = 64/163 (39%), Positives = 97/163 (59%), Gaps = 7/163 (4%)
Frame = +2
Query: 167 RFCPYINAEDAMVSKCVKFESGLRPVIYQYMCVQEIRDFDTLVHKCRMFDDAGKAKSNYY 226
+F P+ AE A SKC+KFE+GLRP I + + Q++R F LV+ CR++++ KA +
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAIGYQQLRVFPDLVNTCRIYEEDTKA---HD 172
Query: 227 KVQNEKKGKGH-GVGKPYS--KDKGKKR---ESGGGSRPSLADVKCYRCGTLGHYANECM 280
KV NE+K KG KPYS DKGK+R + + + A++ C+ G GH +N C
Sbjct: 173 KVVNERKTKGQ*SRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIVCFNYGEKGHKSNVCP 352
Query: 281 NDL-NCHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKK 322
++ C +C K GH DC+ +I C+NC E+GHI ++ K+
Sbjct: 353 KEIKKCVRCDKKGHIVADCK--RNDIVCFNCNEEGHIGSQCKQ 475
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 97.1 bits (240), Expect = 2e-20
Identities = 50/135 (37%), Positives = 78/135 (57%)
Frame = +2
Query: 633 EDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKS 692
+D+ KTAF T G Y Y VMPFG+ NA + +NR+F L + V+IDD+LV S
Sbjct: 11 DDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVKSLR 190
Query: 693 EEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWE 752
+H HL+ + L E + +KC F + FLG+++++ GI V+P ++ A+L
Sbjct: 191 ATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAILDLP 370
Query: 753 SPKSVFEIKAIWGLD 767
SPK+ E++ + G D
Sbjct: 371 SPKNSREVQRLTGED 415
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 82.4 bits (202), Expect = 6e-16
Identities = 39/69 (56%), Positives = 51/69 (73%)
Frame = -1
Query: 556 KQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEV 615
K+F +PS+SP GA +L V+KKDG R+C+DYRQ NK+T KN+YPLPRID+ D++
Sbjct: 274 KRFQQPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCY 95
Query: 616 FSKIDLRSG 624
F IDLR G
Sbjct: 94 F*NIDLRLG 68
Score = 47.4 bits (111), Expect = 2e-05
Identities = 32/87 (36%), Positives = 48/87 (54%)
Frame = -2
Query: 494 VLPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEEL 553
V+ V++ F VFP++ +P ERE+ F IDL+ T IS P M +EL +LK L++
Sbjct: 459 VVLVLKGFS*VFPDNFPVIPLEREIFFCIDLLLDTQLISNPP*LMDRTELKKLKI*LKDS 280
Query: 554 LEKQFIRPSVSPRGAPVLLVKKKDGSM 580
LEK F R L +K+ G++
Sbjct: 279 LEKGFNNQVFLHRVLRCCLCEKRMGTL 199
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 57.0 bits (136), Expect = 3e-08
Identities = 27/80 (33%), Positives = 41/80 (50%), Gaps = 2/80 (2%)
Frame = -2
Query: 57 PTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWWINAKGRLELGG 116
P F+G P+ W+Q MER+F+ ++QKV++ + A WW N K R + G
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 117 --DVVTWAKFKVEFLRKYFP 134
+ TW K + + RKY P
Sbjct: 544 KSKIKTWEKMRQKLTRKYLP 485
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 55.8 bits (133), Expect = 6e-08
Identities = 28/120 (23%), Positives = 60/120 (49%), Gaps = 7/120 (5%)
Frame = +2
Query: 32 NTAANRRNEGEAEERRLDRFLRNNPPTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVR 91
+++++R + + R+ + ++ + P F G+ P+ W+Q +ER+F+ ++QKV+
Sbjct: 275 DSSSSRSSRSHRRQTRMSK-IKVDIPDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVK 451
Query: 92 LATHMFAEEAEYWWINAKGRLELGG--DVVTWAKFKVEFLRK-----YFPEDLRTKKEVE 144
+ + A WW N K + G + TW K + + RK Y+ ++ KK ++
Sbjct: 452 IVAAKLKKHASIWWKNLKRKRNCEGKSKIKTWDKMRQKLTRKYLHPHYYQDNFTQKKNIQ 631
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 55.1 bits (131), Expect = 1e-07
Identities = 24/58 (41%), Positives = 34/58 (58%), Gaps = 4/58 (6%)
Frame = +1
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVVARE----ITCYNCGEKGHISTK 319
C+ C GH A+ C N+ CH CGKAGH+A +C V + C NC ++GHI+ +
Sbjct: 538 CWNCKEPGHMASSCPNEGICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVE 711
Score = 50.8 bits (120), Expect = 2e-06
Identities = 27/77 (35%), Positives = 38/77 (49%)
Frame = +1
Query: 242 PYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVV 301
PY +D + G S+ +L C C GHY EC N CH C GH A +C
Sbjct: 379 PYRRDSRR-----GFSQDNL----CKNCKRPGHYVRECPNVAVCHNCSLPGHIASEC--- 522
Query: 302 AREITCYNCGEKGHIST 318
+ + C+NC E GH+++
Sbjct: 523 STKSLCWNCKEPGHMAS 573
Score = 44.7 bits (104), Expect = 1e-04
Identities = 21/58 (36%), Positives = 28/58 (48%)
Frame = +1
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKA 323
C C GH A EC N+ C+ C K GH A DC + C C GH++ + K+
Sbjct: 673 CNNCYKQGHIAVECTNEKACNNCRKTGHLARDC---PNDPICNLCNISGHVARQCPKS 837
Score = 39.3 bits (90), Expect = 0.006
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 3/56 (5%)
Frame = +1
Query: 246 DKGKKRESGGGSRPS-LADVKCYRCGTLGHYANECMND--LNCHKCGKAGHKAVDC 298
D+G GG R DV C C GH + +CM + C CG GH+A +C
Sbjct: 850 DRGGGGSLRGGYRDGGFRDVVCRSCQQFGHMSRDCMGGPLMICQNCGGRGHQAYEC 1017
Score = 35.8 bits (81), Expect = 0.065
Identities = 23/88 (26%), Positives = 33/88 (37%), Gaps = 23/88 (26%)
Frame = +1
Query: 251 RESGGGSRPSLADVKCYRCGTLGHYANEC-----------------------MNDLNCHK 287
R++G +R D C C GH A +C D+ C
Sbjct: 742 RKTGHLARDCPNDPICNLCNISGHVARQCPKSNVIGDRGGGGSLRGGYRDGGFRDVVCRS 921
Query: 288 CGKAGHKAVDCRVVAREITCYNCGEKGH 315
C + GH + DC + + C NCG +GH
Sbjct: 922 CQQFGHMSRDC-MGGPLMICQNCGGRGH 1002
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 52.8 bits (125), Expect = 5e-07
Identities = 32/122 (26%), Positives = 57/122 (46%), Gaps = 4/122 (3%)
Frame = +2
Query: 17 QALATMAQVLAQGNENTAANRRNEGEAEERRLDRF--LRNNPPTFKGRFDPEGAQTWVQG 74
QAL Q +G+ + + + + +RR + ++ + P F+G + W+Q
Sbjct: 503 QALLEAEQRRFEGDVSYSDSSSSRSSHSQRRQLQMNDIKVDIPDFEGNLQLDDFLDWLQT 682
Query: 75 MERIFRAMVTGDDQKVRLATHMFAEEAEYWWINAKGRLELGG--DVVTWAKFKVEFLRKY 132
+ER+F ++QKV++ + A WW N K R + G + TW K + + RKY
Sbjct: 683 IERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREGKSKIKTWDKMRQKLTRKY 862
Query: 133 FP 134
P
Sbjct: 863 LP 868
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 45.1 bits (105), Expect = 1e-04
Identities = 37/131 (28%), Positives = 49/131 (37%), Gaps = 30/131 (22%)
Frame = +3
Query: 217 DAGKAKSNYYKVQNEKKGKGHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYA 276
D K +V+ + G G G G+ + +G +R +GGG CY CG GH A
Sbjct: 240 DVTGPKGEPLQVRQDNHGGGGG-GRGF---RGGERRNGGGG--------CYTCGDTGHIA 383
Query: 277 NEC----MNDLN--------------CHKCGKAGHKAVDCR------------VVAREIT 306
+C ND N C+ CG H A DC +
Sbjct: 384 RDCDRSDRNDRNDRSGGGGGGDRDRACYTCGSFEHFARDCMRGGGNNNNGGGGYGGGGTS 563
Query: 307 CYNCGEKGHIS 317
CY CG GHI+
Sbjct: 564 CYRCGGVGHIA 596
Score = 43.5 bits (101), Expect = 3e-04
Identities = 20/68 (29%), Positives = 29/68 (42%), Gaps = 15/68 (22%)
Frame = +3
Query: 246 DKGKKRESGGGSRPSLADVKCYRCGTLGHYANECM---------------NDLNCHKCGK 290
D+ + + GG D CY CG+ H+A +CM +C++CG
Sbjct: 402 DRNDRNDRSGGGGGGDRDRACYTCGSFEHFARDCMRGGGNNNNGGGGYGGGGTSCYRCGG 581
Query: 291 AGHKAVDC 298
GH A DC
Sbjct: 582 VGHIARDC 605
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 42.7 bits (99), Expect = 5e-04
Identities = 30/92 (32%), Positives = 39/92 (41%)
Frame = +3
Query: 190 RPVIYQYMCVQEIRDFDTLVHKCRMFDDAGKAKSNYYKVQNEKKGKGHGVGKPYSKDKGK 249
RP Y ++ + RD + D GK N + G G G G G+
Sbjct: 147 RPPGYAFIDFDDRRDAQDAIR-----DLDGKNGWRVELSHNSRSGGGGGGG-----GGGR 296
Query: 250 KRESGGGSRPSLADVKCYRCGTLGHYANECMN 281
R GGG +D+KCY CG GH+A EC N
Sbjct: 297 GRGGGGGGG---SDLKCYECGEPGHFARECRN 383
Score = 30.0 bits (66), Expect = 3.6
Identities = 10/19 (52%), Positives = 15/19 (78%)
Frame = +3
Query: 281 NDLNCHKCGKAGHKAVDCR 299
+DL C++CG+ GH A +CR
Sbjct: 324 SDLKCYECGEPGHFARECR 380
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 42.0 bits (97), Expect = 0.001
Identities = 18/43 (41%), Positives = 23/43 (52%)
Frame = +2
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKAAGKV 327
C CG GH A DC+ + CY CGE+GHI K + K+
Sbjct: 359 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKL 487
Score = 29.3 bits (64), Expect = 6.1
Identities = 10/17 (58%), Positives = 10/17 (58%)
Frame = +2
Query: 265 KCYRCGTLGHYANECMN 281
KCYRCG GH C N
Sbjct: 422 KCYRCGERGHIEKNCKN 472
>AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing factor
[imported] - Arabidopsis thaliana, partial (62%)
Length = 508
Score = 38.9 bits (89), Expect = 0.008
Identities = 20/46 (43%), Positives = 23/46 (49%)
Frame = +2
Query: 234 GKGHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANEC 279
G G G G G+ R GGGS D+KCY CG GH+A C
Sbjct: 263 GGGGGRGGGGGGGGGRGRSGGGGS-----DLKCYXCGEPGHFARXC 385
>TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 421
Score = 37.7 bits (86), Expect = 0.017
Identities = 20/48 (41%), Positives = 23/48 (47%)
Frame = +3
Query: 242 PYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLNCHKCG 289
PY +D + G SR +L C C GHYA EC N CH CG
Sbjct: 303 PYRRDSSR-----GFSRDNL----CKNCKRPGHYARECPNVAVCHNCG 419
>BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (54%)
Length = 364
Score = 37.7 bits (86), Expect = 0.017
Identities = 31/93 (33%), Positives = 41/93 (43%), Gaps = 3/93 (3%)
Frame = +1
Query: 190 RPVIYQYMCVQEIRDFDTLVHKCRMFDDAGKAKSNYYKVQ---NEKKGKGHGVGKPYSKD 246
RP Y ++ + RD +H+ GK N ++VQ N K G G G
Sbjct: 145 RPPGYAFIEFDDPRDALDAIHELD-----GK---NGWRVQLSHNSKSGGGGG-------- 276
Query: 247 KGKKRESGGGSRPSLADVKCYRCGTLGHYANEC 279
+G R GG D+KCY CG GH+A EC
Sbjct: 277 RGGGRGGRGGD-----DLKCYECGEPGHFAREC 360
Score = 30.0 bits (66), Expect = 3.6
Identities = 10/19 (52%), Positives = 15/19 (78%)
Frame = +1
Query: 281 NDLNCHKCGKAGHKAVDCR 299
+DL C++CG+ GH A +CR
Sbjct: 307 DDLKCYECGEPGHFARECR 363
>TC82733 similar to GP|10177404|dbj|BAB10535.
gene_id:K24M7.12~pir||S42136~similar to unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 710
Score = 36.6 bits (83), Expect = 0.038
Identities = 35/129 (27%), Positives = 50/129 (38%), Gaps = 15/129 (11%)
Frame = +3
Query: 216 DDAGKAKSNYYKVQNEKKGKGHGVGKPYSKDK---GKKRESGGGSRPSLADVKCYRCGTL 272
+D K K N +K + KP S K GK+ G +P + C+ C L
Sbjct: 165 NDPSKKKKNKFKRK-----------KPDSNSKPRTGKRPLRVPGMKPGDS---CFICKGL 302
Query: 273 GHYANECMNDLN------CHKCGKAGHKAVDCRVVAREITC---YNCGEKGHISTK---P 320
H A C C +C + GH+A +C + YNCG+ GH P
Sbjct: 303 DHIAKFCTQKAEWEKNKICLRCRRRGHRAQNCPDGGSKEDFKY*YNCGDNGHSLANCPHP 482
Query: 321 KKAAGKVFA 329
+ G +FA
Sbjct: 483 LQEGGTMFA 509
>TC88960 similar to GP|21537297|gb|AAM61638.1 DNA-damage inducible protein
DDI1-like {Arabidopsis thaliana}, partial (91%)
Length = 1523
Score = 28.5 bits (62), Expect(2) = 0.057
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = +2
Query: 348 INSTPLIAIIDTGATHSFISASCVE 372
+N PL A +D+GA + IS +C E
Sbjct: 740 VNGVPLKAFVDSGAQSTIISKTCAE 814
Score = 26.2 bits (56), Expect(2) = 0.057
Identities = 9/30 (30%), Positives = 17/30 (56%)
Frame = +1
Query: 405 PVNFGDVDFKLDLVCLPLKNMDVIFGMDWL 434
P+ G++ + + L NM+ +FG+D L
Sbjct: 910 PIKIGNIFYPCSFLVLDSSNMEFLFGLDML 999
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 35.8 bits (81), Expect = 0.065
Identities = 20/60 (33%), Positives = 27/60 (44%), Gaps = 14/60 (23%)
Frame = -1
Query: 253 SGGGSRPSLADVKCYRCGTLGHYANECMN--------------DLNCHKCGKAGHKAVDC 298
SG G A KCY+C GH+A+ C + NC+KC + GH A +C
Sbjct: 528 SGSGG----ASGKCYKCQQPGHWASNCPSMSAANRVSGGSGGASGNCYKCNQPGHWANNC 361
Score = 33.1 bits (74), Expect = 0.42
Identities = 20/64 (31%), Positives = 24/64 (37%)
Frame = -1
Query: 218 AGKAKSNYYKVQNEKKGKGHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYAN 277
+G A YK Q GH S + G G A CY+C GH+AN
Sbjct: 522 SGGASGKCYKCQQP----GHWASNCPSMSAANRVSGGSGG----ASGNCYKCNQPGHWAN 367
Query: 278 ECMN 281
C N
Sbjct: 366 NCPN 355
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 34.7 bits (78), Expect = 0.15
Identities = 17/54 (31%), Positives = 22/54 (40%), Gaps = 21/54 (38%)
Frame = +1
Query: 266 CYRCGTLGHYANECMN---------------------DLNCHKCGKAGHKAVDC 298
CY CG GH A EC + +C+ CG++GH A DC
Sbjct: 16 CYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDC 177
Score = 30.8 bits (68), Expect = 2.1
Identities = 16/44 (36%), Positives = 18/44 (40%)
Frame = +1
Query: 236 GHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANEC 279
G G G Y G GGG CY CG GH+A +C
Sbjct: 67 GGGGGGRYGGGGGGGGGGGGGG-------SCYSCGESGHFARDC 177
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,338,491
Number of Sequences: 36976
Number of extensions: 325350
Number of successful extensions: 1507
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 1421
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1478
length of query: 790
length of database: 9,014,727
effective HSP length: 104
effective length of query: 686
effective length of database: 5,169,223
effective search space: 3546086978
effective search space used: 3546086978
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126008.1


