

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124964.6 - phase: 0
(387 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
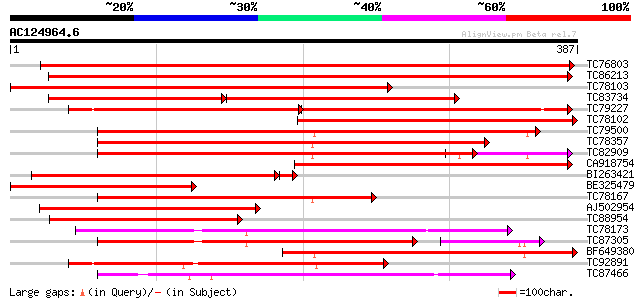
Score E
Sequences producing significant alignments: (bits) Value
TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold... 598 e-171
TC86213 homologue to PIR|S60121|S60121 mitogen-activated protein... 552 e-158
TC78103 SP|Q07176|MMK1_MEDSA Mitogen-activated protein kinase ho... 474 e-134
TC83734 homologue to PIR|S60121|S60121 mitogen-activated protein... 260 e-123
TC79227 homologue to GP|8132347|gb|AAF73257.1| MAP kinase PsMAPK... 242 e-114
TC78102 homologue to SP|Q07176|MMK1_MEDSA Mitogen-activated prot... 390 e-109
TC79500 similar to GP|9294059|dbj|BAB02016.1 MAP kinase {Arabido... 320 4e-88
TC78357 similar to GP|15289849|dbj|BAB63546. putative mitogen-ac... 301 4e-82
TC82909 homologue to GP|17064906|gb|AAL32607.1 Unknown protein {... 293 6e-80
CA918754 similar to PIR|S60121|S60 mitogen-activated protein kin... 275 3e-74
BI263421 similar to GP|21165527|db mitogen-activated protein kin... 259 3e-71
BE325479 SP|Q07176|MMK1 Mitogen-activated protein kinase homolog... 262 2e-70
TC78167 homologue to GP|9294393|dbj|BAB02403.1 mitogen-activated... 235 2e-62
AJ502954 homologue to GP|4456682|emb MAP kinase {Medicago sativa... 215 2e-56
TC88954 similar to PIR|S40470|S40470 mitogen-activated protein k... 207 7e-54
TC78173 SP|P24923|CC21_MEDSA Cell division control protein 2 hom... 206 2e-53
TC87305 homologue to SP|Q05006|CC22_MEDSA Cell division control ... 182 1e-51
BF649380 similar to GP|9294059|dbj MAP kinase {Arabidopsis thali... 192 2e-49
TC92891 similar to GP|4580577|gb|AAD24428.1| MAP protein kinase ... 188 3e-48
TC87466 homologue to GP|10185114|emb|CAC08564. wound-induced GSK... 188 3e-48
>TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold- and
drought-induced - alfalfa, complete
Length = 1709
Score = 598 bits (1541), Expect = e-171
Identities = 281/365 (76%), Positives = 323/365 (87%), Gaps = 1/365 (0%)
Frame = +2
Query: 22 QMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHV 81
Q G+ PA +HGG+F+QYN+FGN+FEVTAKY+PPIMPIG+GAYGIVCS N+ETNE V
Sbjct: 131 QNGVAEFPAVQTHGGQFVQYNVFGNLFEVTAKYRPPIMPIGRGAYGIVCSLLNTETNELV 310
Query: 82 AVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDT 141
AVKKIANAFDN +DAKRTLREIKLLRH+DHENV+ +RD++PPP R FNDVYI ELMDT
Sbjct: 311 AVKKIANAFDNHMDAKRTLREIKLLRHLDHENVIGLRDVIPPPLRREFNDVYITTELMDT 490
Query: 142 DLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFG 201
DLHQIIRSNQ LS+EHCQYFLYQILRGL+YIHSAN++HRDLKPSNLLLNANCDLKI DFG
Sbjct: 491 DLHQIIRSNQNLSDEHCQYFLYQILRGLRYIHSANIIHRDLKPSNLLLNANCDLKIIDFG 670
Query: 202 LARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDH 261
LAR T E+DFMTEYVVTRWYRAPELLLNSSDYT+AIDVWSVGCIFMELM++KPLFPG+DH
Sbjct: 671 LARPTMESDFMTEYVVTRWYRAPELLLNSSDYTSAIDVWSVGCIFMELMNKKPLFPGKDH 850
Query: 262 VHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLVEKM 320
VHQ+RLL EL+GTP++ D+G + N++A+RYIRQLP Y RQ FP VHP AIDLV+KM
Sbjct: 851 VHQMRLLTELLGTPTDADVGLVKNDDARRYIRQLPQYPRQPLNRVFPHVHPLAIDLVDKM 1030
Query: 321 LTFDPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYREALA 380
LT DP +RITVE+ALAHPYL LHD++DEP+CM PFSF+FEQ L EEQ+KE+IYREALA
Sbjct: 1031LTIDPTRRITVEEALAHPYLEKLHDVADEPICMEPFSFEFEQQHLDEEQIKEMIYREALA 1210
Query: 381 FNPEY 385
NPEY
Sbjct: 1211LNPEY 1225
>TC86213 homologue to PIR|S60121|S60121 mitogen-activated protein kinase
MMK2 (EC 2.7.1.-) - alfalfa, complete
Length = 1596
Score = 552 bits (1423), Expect = e-158
Identities = 256/359 (71%), Positives = 312/359 (86%), Gaps = 1/359 (0%)
Frame = +2
Query: 27 NIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKI 86
NI +HGGR++QYNI+GN+FEV+ KY PPI P+G+GAYGIVC+A N+ET E VA+KKI
Sbjct: 266 NIRGIPTHGGRYLQYNIYGNLFEVSRKYVPPIRPVGRGAYGIVCAAVNAETREEVAIKKI 445
Query: 87 ANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQI 146
NAFDN+IDAKRTLREIKLLRHMDHENV++I+DI+ PPQ+E FNDVYI ELMDTDLHQI
Sbjct: 446 GNAFDNRIDAKRTLREIKLLRHMDHENVMSIKDIIRPPQKENFNDVYIVSELMDTDLHQI 625
Query: 147 IRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVT 206
IRSNQ ++++HC+YF+YQ+LRGLKY+HSANVLHRDLKPSNLLLNANCDLKI DFGLAR T
Sbjct: 626 IRSNQPMTDDHCRYFVYQLLRGLKYVHSANVLHRDLKPSNLLLNANCDLKIGDFGLARTT 805
Query: 207 SETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLR 266
SETDFMTEYVVTRWYRAPELLLN S+YTAAID+WSVGCI E++ R+PLFPGRD+VHQLR
Sbjct: 806 SETDFMTEYVVTRWYRAPELLLNCSEYTAAIDIWSVGCILGEIVTRQPLFPGRDYVHQLR 985
Query: 267 LLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLVEKMLTFDP 325
L+ ELIG+P + LGFL +ENA+RY+RQLP Y +Q+F +FP + P A+DL+EKML FDP
Sbjct: 986 LVTELIGSPDDASLGFLRSENARRYVRQLPQYPQQNFSTRFPSMSPGAVDLLEKMLIFDP 1165
Query: 326 RKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYREALAFNPE 384
KRI V++AL HPY+ LHDI++EP+C PFSFDFE+ TEE +KELI++E++ FNP+
Sbjct: 1166SKRIRVDEALCHPYMAPLHDINEEPICARPFSFDFEEPMFTEEDIKELIWKESVRFNPD 1342
>TC78103 SP|Q07176|MMK1_MEDSA Mitogen-activated protein kinase homolog MMK1
(EC 2.7.1.-) (MAP kinase MSK7) (MAP kinase ERK1).,
partial (65%)
Length = 923
Score = 474 bits (1220), Expect = e-134
Identities = 236/261 (90%), Positives = 239/261 (91%)
Frame = +1
Query: 1 MEGGGAPPADTVMSDAAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMP 60
MEGGGAPPADTVMSDAAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMP
Sbjct: 136 MEGGGAPPADTVMSDAAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMP 315
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI
Sbjct: 316 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 495
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHR 180
VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHR
Sbjct: 496 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHR 675
Query: 181 DLKPSNLLLNANCDLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVW 240
DLKPSNLLLNANCDLKICDFGLARVTSETDFMTEYVVTRWYRAPELLL +
Sbjct: 676 DLKPSNLLLNANCDLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLTLLIIQQQLMYG 855
Query: 241 SVGCIFMELMDRKPLFPGRDH 261
+GCIF K L G +H
Sbjct: 856 LLGCIFHGTDGSKALXXGXNH 918
>TC83734 homologue to PIR|S60121|S60121 mitogen-activated protein kinase
MMK2 (EC 2.7.1.-) - alfalfa, partial (76%)
Length = 925
Score = 260 bits (664), Expect(2) = e-123
Identities = 119/160 (74%), Positives = 143/160 (89%), Gaps = 1/160 (0%)
Frame = +2
Query: 149 SNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVTSE 208
+NQ L+++HC+YFLYQ+LRGLKY+HSANVLHRDLKPSNLLLN+NCDLKI DFGLAR TSE
Sbjct: 437 TNQTLTDDHCRYFLYQLLRGLKYVHSANVLHRDLKPSNLLLNSNCDLKIGDFGLARTTSE 616
Query: 209 TDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLL 268
TDFMTEYVVTRWYRAPELLLN S+YTAAID+WSVGCI E+M RKPLFPG+D+VHQL+L+
Sbjct: 617 TDFMTEYVVTRWYRAPELLLNCSEYTAAIDIWSVGCILGEIMTRKPLFPGKDYVHQLKLI 796
Query: 269 MELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFP 307
ELIG+P + LGF+ ++NA+RY++QLP Y RQ F +FP
Sbjct: 797 TELIGSPDDASLGFIRSDNARRYVKQLPQYPRQQFAARFP 916
Score = 200 bits (509), Expect(2) = e-123
Identities = 91/122 (74%), Positives = 108/122 (87%)
Frame = +1
Query: 27 NIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKI 86
NI L+HGGR++QYNI+GN+FEV+ KY PPI PIG+GAYGIVC+A N+ET E VA+KK+
Sbjct: 70 NIKGVLTHGGRYVQYNIYGNLFEVSNKYVPPIRPIGRGAYGIVCAAVNAETREEVAIKKV 249
Query: 87 ANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQI 146
NAFDN+IDAKRTLREIKL RHMDHENV+A++DI+ PPQ+E FNDVYI YELMDTDLHQI
Sbjct: 250 GNAFDNRIDAKRTLREIKLQRHMDHENVIALKDIIRPPQKENFNDVYIVYELMDTDLHQI 429
Query: 147 IR 148
IR
Sbjct: 430 IR 435
>TC79227 homologue to GP|8132347|gb|AAF73257.1| MAP kinase PsMAPK2 {Pisum
sativum}, complete
Length = 1731
Score = 242 bits (618), Expect(2) = e-114
Identities = 111/160 (69%), Positives = 138/160 (85%)
Frame = +3
Query: 41 YNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTL 100
++++ +FE+ KY P I PIG+GAYGIVCS+ N ETNE VA+KKI NAF+N++DA RTL
Sbjct: 330 FSMWQTLFEIDTKYVP-IKPIGRGAYGIVCSSVNRETNEKVAIKKIQNAFENRVDALRTL 506
Query: 101 REIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQY 160
RE+KLLRH+ HENV+A++DI+ P R F DVY+ YELMDTDLHQII+S+QALS +HCQY
Sbjct: 507 RELKLLRHLHHENVIALKDIMMPNHRNNFKDVYLVYELMDTDLHQIIKSSQALSNDHCQY 686
Query: 161 FLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDF 200
FL+Q+LRGLKY+HSAN+LHRDLKP NLL+NANCDLKICDF
Sbjct: 687 FLFQLLRGLKYLHSANILHRDLKPGNLLINANCDLKICDF 806
Score = 186 bits (473), Expect(2) = e-114
Identities = 93/187 (49%), Positives = 130/187 (68%), Gaps = 2/187 (1%)
Frame = +1
Query: 200 FGLARVT-SETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPG 258
FGLAR S+ FMTEYVVTRWYRAPELLL +Y +IDVWSVGCIF EL+ RKP+ PG
Sbjct: 805 FGLARTNCSKNQFMTEYVVTRWYRAPELLLCWDNYGTSIDVWSVGCIFAELLGRKPISPG 984
Query: 259 RDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLV 317
+ ++QL+L++ ++G+ E+D+ F+ N AKRYI+ LP F +P HP AIDL+
Sbjct: 985 SECLNQLKLIINILGSQREEDIEFIDNPKAKRYIKSLPYSPGTPFSRLYPNAHPLAIDLL 1164
Query: 318 EKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYRE 377
KML FDP KRI+V +AL HP++ SL+D + +P + P D ++ L E+ ++EL++ E
Sbjct: 1165SKMLVFDPTKRISVTEALQHPFMASLYDPNSDPPAIIPIDLDIDED-LGEDVIRELMWSE 1341
Query: 378 ALAFNPE 384
L ++PE
Sbjct: 1342MLHYHPE 1362
>TC78102 homologue to SP|Q07176|MMK1_MEDSA Mitogen-activated protein kinase
homolog MMK1 (EC 2.7.1.-) (MAP kinase MSK7) (MAP kinase
ERK1)., partial (49%)
Length = 834
Score = 390 bits (1001), Expect = e-109
Identities = 189/191 (98%), Positives = 189/191 (98%)
Frame = +1
Query: 197 ICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLF 256
ICDF ARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLF
Sbjct: 1 ICDFDWARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLF 180
Query: 257 PGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDL 316
PGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDL
Sbjct: 181 PGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDL 360
Query: 317 VEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYR 376
VEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYR
Sbjct: 361 VEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYR 540
Query: 377 EALAFNPEYQQ 387
EALAFNPEYQQ
Sbjct: 541 EALAFNPEYQQ 573
>TC79500 similar to GP|9294059|dbj|BAB02016.1 MAP kinase {Arabidopsis
thaliana}, partial (55%)
Length = 1381
Score = 320 bits (821), Expect = 4e-88
Identities = 155/310 (50%), Positives = 221/310 (71%), Gaps = 8/310 (2%)
Frame = +2
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IGKG+YG+VCSA ++ T + VA+KKI + F++ DA R LREIKLLR + H ++V IR I
Sbjct: 449 IGKGSYGVVCSAIDTHTGQKVAIKKINHVFEHVSDATRILREIKLLRLLKHPDIVEIRHI 628
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHR 180
+ PP R F D+Y+ +ELM++DLHQ+I++N L+ +H Q+FLYQ+LRGLK+IH+ANV HR
Sbjct: 629 MLPPSRREFRDIYVVFELMESDLHQVIKANHDLTPQHHQFFLYQLLRGLKFIHTANVFHR 808
Query: 181 DLKPSNLLLNANCDLKICDFGLARVT----SETDFMTEYVVTRWYRAPELLLN-SSDYTA 235
DLKP N+L NA+C LKICDFGLARV+ T F T+YV TRWYRAPEL + S YT
Sbjct: 809 DLKPKNILANADCKLKICDFGLARVSFNDAPSTIFWTDYVATRWYRAPELCGSFFSKYTP 988
Query: 236 AIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQL 294
ID+WS+GCIF E++ +PLFPG++ VHQL ++ +L+GTP + + + NE A+RY+ +
Sbjct: 989 GIDIWSIGCIFAEMLTGRPLFPGKNVVHQLDIMTDLLGTPPPESIARIRNEKARRYLNSM 1168
Query: 295 PPYRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVC-- 352
+ F +KFP + P A+ ++E++L FDP+ R + E+AL+ PY L +I EP
Sbjct: 1169RKKQPVPFSQKFPNIDPLALRILERLLAFDPKNRPSAEEALSDPYFHGLSNIDREPSTHP 1348
Query: 353 MTPFSFDFEQ 362
++ F+FE+
Sbjct: 1349ISKLEFEFER 1378
>TC78357 similar to GP|15289849|dbj|BAB63546. putative mitogen-activated
protein kinase {Oryza sativa (japonica cultivar-group)},
partial (69%)
Length = 1751
Score = 301 bits (770), Expect = 4e-82
Identities = 149/273 (54%), Positives = 201/273 (73%), Gaps = 6/273 (2%)
Frame = +3
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IGKG+YG+VCSA ++ T E VA+KKI + F++ DA R LREIKLLR + H ++V I+ I
Sbjct: 549 IGKGSYGVVCSAIDTHTGEKVAIKKIHDIFEHISDAARILREIKLLRLLRHPDIVEIKHI 728
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHR 180
+ PP R+ F D+Y+ +ELM++DLHQ+I++N L++EH Q+FLYQ+LR LKYIH+ANV HR
Sbjct: 729 MLPPSRKDFKDIYVVFELMESDLHQVIKANDDLTKEHYQFFLYQLLRALKYIHTANVYHR 908
Query: 181 DLKPSNLLLNANCDLKICDFGLARV----TSETDFMTEYVVTRWYRAPELLLN-SSDYTA 235
DLKP N+L NANC LKICDFGLARV T T F T+YV TRWYRAPEL + S YT
Sbjct: 909 DLKPKNILANANCKLKICDFGLARVAFSDTPTTVFWTDYVATRWYRAPELCGSFYSKYTP 1088
Query: 236 AIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQL 294
AID+WS+GCIF E++ KPLFPG++ VHQL L+ +L+GTPS D + + NE A+RY+ +
Sbjct: 1089AIDIWSIGCIFAEVLTGKPLFPGKNVVHQLDLMTDLLGTPSLDTISRVRNEKARRYLTSM 1268
Query: 295 PPYRRQSFQEKFPQVHPEAIDLVEKMLTFDPRK 327
+ F +KF P A+ L+E++L F P++
Sbjct: 1269RKKQPVPFAQKFSNADPLALRLLERLLAF*PKR 1367
Score = 37.4 bits (85), Expect = 0.010
Identities = 24/82 (29%), Positives = 45/82 (54%), Gaps = 5/82 (6%)
Frame = +2
Query: 308 QVHPEAIDLVEKMLTFDPRKRITVEDAL-----AHPYLTSLHDISDEPVCMTPFSFDFEQ 362
Q P +I + K+ F +K + L + +L S + S +P+ T F+FE+
Sbjct: 1307 QCRPFSITTIGKITCFLTQKTGLLLKKLWLILTSRDWLKSRGNPSCQPI--TKMEFEFER 1480
Query: 363 HALTEEQMKELIYREALAFNPE 384
+T+E+++ELI+RE L ++P+
Sbjct: 1481 RRVTKEEIRELIFREILEYHPQ 1546
>TC82909 homologue to GP|17064906|gb|AAL32607.1 Unknown protein {Arabidopsis
thaliana}, partial (67%)
Length = 1274
Score = 293 bits (751), Expect = 6e-80
Identities = 144/265 (54%), Positives = 190/265 (71%), Gaps = 6/265 (2%)
Frame = +2
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IGKG+YG+VCSA+++ T E VA+KKI + F++ DA R LREIKLLR + H ++V I+ I
Sbjct: 203 IGKGSYGVVCSAYDTHTGEKVAIKKINDIFEHVSDATRILREIKLLRLLRHPDIVEIKHI 382
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHR 180
+ PP R F D+Y+ +ELM++DLHQ+I++N L+ EH Q+FLYQ+LRGLKYIH+ANV HR
Sbjct: 383 LLPPSRREFKDIYVVFELMESDLHQVIKANDDLTPEHYQFFLYQLLRGLKYIHTANVFHR 562
Query: 181 DLKPSNLLLNANCDLKICDFGLARV----TSETDFMTEYVVTRWYRAPELLLN-SSDYTA 235
DLKP N+L NA+C LKICDFGLARV T F T+YV TRWYRAPEL + S YT
Sbjct: 563 DLKPKNILANADCKLKICDFGLARVAFNDTPTAIFWTDYVATRWYRAPELCGSFFSKYTP 742
Query: 236 AIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQL 294
AID+WS+GCIF EL+ KPLFPG++ VHQL ++ + +GTPS D + + NE A+RY+ +
Sbjct: 743 AIDIWSIGCIFAELLTGKPLFPGKNVVHQLDIMTDFLGTPSPDAIARVRNEKARRYLSSM 922
Query: 295 PPYRRQSFQEKFPQVHPEAIDLVEK 319
+ F KFP P V+K
Sbjct: 923 RKKKPVPFSHKFPNADPPCSSAVKK 997
Score = 62.8 bits (151), Expect = 2e-10
Identities = 34/91 (37%), Positives = 53/91 (57%), Gaps = 4/91 (4%)
Frame = +3
Query: 298 RRQSFQEK--FPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVC--M 353
RR F + F P A+ L+++ML F+ + R T E+AL PY L + EP +
Sbjct: 927 RRSQFLSRTNFLMQIPLALRLLKRMLAFEAKDRPTAEEALTDPYFKGLAKVEREPSAQPV 1106
Query: 354 TPFSFDFEQHALTEEQMKELIYREALAFNPE 384
T F+FE+ +T+E ++ELIYRE L ++P+
Sbjct: 1107 TKMEFEFERRRITKEDVRELIYREILEYHPK 1199
>CA918754 similar to PIR|S60121|S60 mitogen-activated protein kinase MMK2 (EC
2.7.1.-) - alfalfa, partial (52%)
Length = 703
Score = 275 bits (702), Expect = 3e-74
Identities = 131/191 (68%), Positives = 161/191 (83%), Gaps = 1/191 (0%)
Frame = -1
Query: 195 LKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKP 254
LKI DFGLAR TSETDFMTEYVVTRWYRAPELLLN S+YT+AIDVWSVGCIF E+M R+P
Sbjct: 688 LKIGDFGLARTTSETDFMTEYVVTRWYRAPELLLNCSEYTSAIDVWSVGCIFAEIMTREP 509
Query: 255 LFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEA 313
LFPG+D+VHQLRL+ ELIG+P + L FL +ENA++Y+RQLP + +Q+ KFP + E
Sbjct: 508 LFPGKDYVHQLRLITELIGSPDDSSLRFLRSENARKYLRQLPQFGKQNLSVKFPSMSAEP 329
Query: 314 IDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKEL 373
++L+EKML FDP KRITV++AL HPYL+SLHDI+DEPV TPFSFDFEQ T+E +KEL
Sbjct: 328 LNLLEKMLVFDPVKRITVDEALCHPYLSSLHDINDEPVPPTPFSFDFEQPGCTQEHIKEL 149
Query: 374 IYREALAFNPE 384
I+RE++ FNP+
Sbjct: 148 IWRESVNFNPD 116
>BI263421 similar to GP|21165527|db mitogen-activated protein kinase {Solanum
tuberosum}, partial (46%)
Length = 682
Score = 259 bits (662), Expect(2) = 3e-71
Identities = 120/169 (71%), Positives = 142/169 (84%)
Frame = +3
Query: 16 AAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNS 75
AA A G I L+HGG++ YN++GN+FEV++KY PPI PIG+GAYGIVC+A NS
Sbjct: 141 AAAAASASGDTKIKRVLTHGGKYAHYNVYGNLFEVSSKYVPPIRPIGRGAYGIVCAAVNS 320
Query: 76 ETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIA 135
+T+E VA+KKI NAFDN IDAKRTLREIKLLRHMDH N++AI+DI+ PP++E FNDVYI
Sbjct: 321 DTHEQVAIKKIGNAFDNIIDAKRTLREIKLLRHMDHPNIIAIKDIIRPPKKEAFNDVYIV 500
Query: 136 YELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKP 184
YELMDTDLH II S+Q L EEHCQYFLYQ+LRGLKY+HSANVLHRDLKP
Sbjct: 501 YELMDTDLHHIIHSDQPLREEHCQYFLYQLLRGLKYVHSANVLHRDLKP 647
Score = 27.3 bits (59), Expect(2) = 3e-71
Identities = 11/12 (91%), Positives = 12/12 (99%)
Frame = +2
Query: 185 SNLLLNANCDLK 196
SNLL+NANCDLK
Sbjct: 647 SNLLVNANCDLK 682
>BE325479 SP|Q07176|MMK1 Mitogen-activated protein kinase homolog MMK1 (EC
2.7.1.-) (MAP kinase MSK7) (MAP kinase ERK1)., partial
(32%)
Length = 557
Score = 262 bits (669), Expect = 2e-70
Identities = 127/127 (100%), Positives = 127/127 (100%)
Frame = +2
Query: 1 MEGGGAPPADTVMSDAAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMP 60
MEGGGAPPADTVMSDAAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMP
Sbjct: 176 MEGGGAPPADTVMSDAAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMP 355
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI
Sbjct: 356 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 535
Query: 121 VPPPQRE 127
VPPPQRE
Sbjct: 536 VPPPQRE 556
>TC78167 homologue to GP|9294393|dbj|BAB02403.1 mitogen-activated protein
kinase {Arabidopsis thaliana}, partial (36%)
Length = 927
Score = 235 bits (600), Expect = 2e-62
Identities = 112/195 (57%), Positives = 150/195 (76%), Gaps = 5/195 (2%)
Frame = +1
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IGKG+YG+VC+A ++ T E VA+KKI +AF++ DA R LRE+KLLR + H ++V I+ I
Sbjct: 340 IGKGSYGVVCAAIDTHTGEKVAIKKIQDAFEHISDAVRILREVKLLRLLRHPDIVDIKRI 519
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHR 180
+ PP + F D+Y+ +ELM++DLHQ+I++N L+ EH Q+FLYQ+LR LKY+HSANV HR
Sbjct: 520 MLPPSKREFKDIYVVFELMESDLHQVIKANDDLTREHHQFFLYQMLRALKYMHSANVYHR 699
Query: 181 DLKPSNLLLNANCDLKICDFGLARV----TSETDFMTEYVVTRWYRAPELLLN-SSDYTA 235
DLKP N+L NANC LK+CDFGLARV + F T+YV TRWYRAPEL + +S YT
Sbjct: 700 DLKPKNILANANCKLKVCDFGLARVAFNDAPTSIFWTDYVATRWYRAPELCGSFASKYTP 879
Query: 236 AIDVWSVGCIFMELM 250
AID+WS+GCIF E++
Sbjct: 880 AIDLWSIGCIFAEVL 924
>AJ502954 homologue to GP|4456682|emb MAP kinase {Medicago sativa}, partial
(41%)
Length = 636
Score = 215 bits (548), Expect = 2e-56
Identities = 102/151 (67%), Positives = 123/151 (80%)
Frame = +3
Query: 21 PQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEH 80
P+ N TL H G++IQYN+ GN FEV + Y PP+ P+G+GAYGIVC A NS+TNE
Sbjct: 183 PESEKANSKGTLIHDGKYIQYNVLGNPFEVYSNYIPPLQPVGRGAYGIVCCATNSDTNEG 362
Query: 81 VAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMD 140
VA+KKI +AFDN+IDAKRTLREIKLL HMDH+NV+ I+DI+ P +E FNDVYI YELMD
Sbjct: 363 VAIKKIGDAFDNRIDAKRTLREIKLLCHMDHDNVIKIKDIIKPADKEKFNDVYIVYELMD 542
Query: 141 TDLHQIIRSNQALSEEHCQYFLYQILRGLKY 171
TDLHQII+SNQAL++EH Q FLYQ+LRGLKY
Sbjct: 543 TDLHQIIQSNQALTDEH*Q*FLYQLLRGLKY 635
>TC88954 similar to PIR|S40470|S40470 mitogen-activated protein kinase 4 (EC
2.7.1.-) - Arabidopsis thaliana, partial (34%)
Length = 863
Score = 207 bits (526), Expect = 7e-54
Identities = 92/132 (69%), Positives = 116/132 (87%)
Frame = +3
Query: 28 IPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKIA 87
+ +H GR++QY+++GN+FEV++KY PP+ PIG+GAYGIVC+A NS+T+E VA+KKIA
Sbjct: 204 VKKVFTHKGRYVQYSLYGNLFEVSSKYVPPLRPIGRGAYGIVCAAVNSDTHEEVAIKKIA 383
Query: 88 NAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQII 147
N FDN IDAKRTLREIKLLRHMDHEN++AI+DI+ PPQ++ FNDVYI YELMDTDLHQII
Sbjct: 384 NTFDNIIDAKRTLREIKLLRHMDHENIIAIKDIIRPPQKDNFNDVYIVYELMDTDLHQII 563
Query: 148 RSNQALSEEHCQ 159
RSNQ L+ +HCQ
Sbjct: 564 RSNQPLNPDHCQ 599
>TC78173 SP|P24923|CC21_MEDSA Cell division control protein 2 homolog 1 (EC
2.7.1.-) (Fragment). [Alfalfa] {Medicago sativa},
complete
Length = 1268
Score = 206 bits (523), Expect = 2e-53
Identities = 115/302 (38%), Positives = 170/302 (56%), Gaps = 4/302 (1%)
Frame = +2
Query: 46 NIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKL 105
+ F T + + IG+G YG+V A + TNE +A+KKI +++ +REI L
Sbjct: 110 SFFFFTMEQYEKVEKIGEGTYGVVYKARDRVTNETIALKKIRLEQEDEGVPSTAIREISL 289
Query: 106 LRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQY--FLY 163
L+ M H N+V ++D+V +R +Y+ +E +D DL + + S+ ++ Q FLY
Sbjct: 290 LKEMQHRNIVRLQDVVHSDKR-----LYLVFEYLDLDLKKHMDSSPEFIKDPRQVKMFLY 454
Query: 164 QILRGLKYIHSANVLHRDLKPSNLLLNANCD-LKICDFGLARVTS-ETDFMTEYVVTRWY 221
Q+L G+ Y HS VLHRDLKP NLL++ + LK+ DFGLAR T VVT WY
Sbjct: 455 QMLCGIAYCHSHRVLHRDLKPQNLLIDRRTNSLKLADFGLARAFGIPVRTFTHEVVTLWY 634
Query: 222 RAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLG 281
RAPE+LL S Y+ +DVWSVGCIF E+ +R+PLFPG + +L + ++GTP+ED
Sbjct: 635 RAPEILLGSRHYSTPVDVWSVGCIFAEMANRRPLFPGDSEIDELFKIFRILGTPNEDTWP 814
Query: 282 FLNENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLT 341
+ + + P + + P + P +DL+ ML DP KRIT A+ H Y
Sbjct: 815 GVT-SLPDFKSTFPRWPSKDLATVVPNLEPAGLDLLNSMLCLDPTKRITARSAVEHEYFK 991
Query: 342 SL 343
+
Sbjct: 992 DI 997
>TC87305 homologue to SP|Q05006|CC22_MEDSA Cell division control protein 2
homolog 2 (EC 2.7.1.-). [Alfalfa] {Medicago sativa},
complete
Length = 1378
Score = 182 bits (463), Expect(2) = 1e-51
Identities = 96/222 (43%), Positives = 141/222 (63%), Gaps = 4/222 (1%)
Frame = +2
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IG+G YG+V A + TNE +A+KKI +++ +REI LL+ M H N+V ++D+
Sbjct: 128 IGEGTYGVVYKARDRATNETIALKKIRLEQEDEGVPSTAIREISLLKEMQHRNIVRLQDV 307
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQY--FLYQILRGLKYIHSANVL 178
V +R +Y+ +E +D DL + + S+ +++ Q FLYQIL G+ Y HS VL
Sbjct: 308 VHSEKR-----LYLVFEYLDLDLKKFMDSSPEFAKDQRQIKMFLYQILCGIAYCHSHRVL 472
Query: 179 HRDLKPSNLLLNANCD-LKICDFGLARVTS-ETDFMTEYVVTRWYRAPELLLNSSDYTAA 236
HRDLKP NLL++ + + LK+ DFGLAR T VVT WYRAPE+LL S Y+
Sbjct: 473 HRDLKPQNLLIDRSSNALKLADFGLARAFGIPVRTFTHEVVTLWYRAPEILLGSRHYSTP 652
Query: 237 IDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSED 278
+DVWSVGCIF E+++++PLFPG + +L + + GTP+E+
Sbjct: 653 VDVWSVGCIFAEMINQRPLFPGDSEIDELFKIFRITGTPNEE 778
Score = 38.9 bits (89), Expect(2) = 1e-51
Identities = 22/80 (27%), Positives = 37/80 (45%), Gaps = 9/80 (11%)
Frame = +3
Query: 295 PPYRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDI------SD 348
P + + + P + P +DL+ ML DP +RIT AL H Y + + ++
Sbjct: 825 PKWPAKDLATQVPNLEPAGLDLLSNMLCLDPTRRITARGALEHEYFKDIKFVP*VLGFTE 1004
Query: 349 EP---VCMTPFSFDFEQHAL 365
E +C++ FD E A+
Sbjct: 1005EVSILLCVSFMGFDSEMGAI 1064
>BF649380 similar to GP|9294059|dbj MAP kinase {Arabidopsis thaliana},
partial (35%)
Length = 664
Score = 192 bits (487), Expect = 2e-49
Identities = 97/209 (46%), Positives = 140/209 (66%), Gaps = 8/209 (3%)
Frame = +1
Query: 187 LLLNANCDLKICDFGLARVT----SETDFMTEYVVTRWYRAPELLLNS-SDYTAAIDVWS 241
+L NA+C LKICDFGLARV+ F T+YV TRWYRAPEL + S YT AID+WS
Sbjct: 1 ILANADCKLKICDFGLARVSFNDAPSAIFWTDYVATRWYRAPELCGSFFSKYTPAIDIWS 180
Query: 242 VGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQ 300
+GCIF E++ KPLFPG++ VHQL L+ +L+GTP + + + NE A+RY+ + +
Sbjct: 181 IGCIFAEMLSGKPLFPGKNVVHQLDLMTDLLGTPPTESISKIRNEKARRYLSSMRKKQPV 360
Query: 301 SFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEP--VCMTPFSF 358
F +KFP V P A++L+E++L FDP+ R T E+AL+ PY L ++ EP ++ F
Sbjct: 361 PFTKKFPNVDPLALNLLERLLAFDPKDRPTAEEALSDPYFHGLSNVDREPSTQAISKLEF 540
Query: 359 DFEQHALTEEQMKELIYREALAFNPEYQQ 387
+FE+ LT++ ++ LIYRE L ++P+ Q
Sbjct: 541 EFERRKLTKDDVRXLIYREILXYHPQMLQ 627
>TC92891 similar to GP|4580577|gb|AAD24428.1| MAP protein kinase MPKA
{Emericella nidulans}, partial (51%)
Length = 804
Score = 188 bits (478), Expect = 3e-48
Identities = 107/226 (47%), Positives = 139/226 (61%), Gaps = 8/226 (3%)
Frame = +1
Query: 41 YNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTL 100
+ + F V KY I +G+GAYG+VC+A N T E VA+KK+ N F+ I KR L
Sbjct: 139 FPVLNQQFLVDEKYNF-IRELGQGAYGVVCAARNKHTGEGVAIKKVTNVFNKTILTKRAL 315
Query: 101 REIKLLRHM-DHENVVAI--RDIVPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEH 157
RE+KLL+H H+N+ + DI+ P +N++Y+ ELM+ DLH IIRS Q L++ H
Sbjct: 316 REVKLLQHFRGHKNITCLYDMDIIDPHD---YNEIYLYEELMEADLHAIIRSGQTLTDAH 486
Query: 158 CQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVTSE-----TDFM 212
Q F YQIL GLKYIHSANVLHRDLKP NLL+NA+C+LKICDFGLAR SE + FM
Sbjct: 487 YQSFSYQILSGLKYIHSANVLHRDLKPGNLLVNADCELKICDFGLARGFSEDPARNSGFM 666
Query: 213 TEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPG 258
TEYV T+ ++ S + F L+ +PLF G
Sbjct: 667 TEYVATKVVQSSRNYAQFSKLYKSY*FMVCKMYFGRLLGGRPLFKG 804
>TC87466 homologue to GP|10185114|emb|CAC08564. wound-induced GSK-3-like
protein {Medicago sativa}, complete
Length = 1934
Score = 188 bits (478), Expect = 3e-48
Identities = 106/292 (36%), Positives = 170/292 (57%), Gaps = 7/292 (2%)
Frame = +3
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
+G G++G+V A ETNE VA+KK+ D + RE++++R +DH N+V ++
Sbjct: 456 VGTGSFGVVFQAKCLETNEAVAIKKVLQ------DKRYKNRELQVMRMVDHPNIVKLKHC 617
Query: 121 V--PPPQREVFNDVYIAY--ELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSA- 175
+ E++ ++ + Y E + IR +Q + H Q + YQILRGL Y+H
Sbjct: 618 FYSTTEKDELYLNLVLEYVPETVYKVSKNYIRMHQHMPIIHVQLYTYQILRGLNYLHEVI 797
Query: 176 NVLHRDLKPSNLLLNANC-DLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYT 234
V HRD+KP NLL+NA LKICDFG A++ + Y+ +R+YRAPEL+ +++YT
Sbjct: 798 GVCHRDIKPQNLLVNAQTHQLKICDFGSAKMLVPGEPNISYICSRYYRAPELIFGATEYT 977
Query: 235 AAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQL 294
AID+WSVGC+ EL+ + +FPG V QL +++++GTP+ +++ +N N + +
Sbjct: 978 TAIDMWSVGCVLAELLLGQAMFPGESGVDQLVEIIKVLGTPTREEIRCMNPNYNEF--KF 1151
Query: 295 PPYRRQSFQEKFPQVHP-EAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHD 345
P + + + F + P EA+DLV ++L + P R T A AHP+ L D
Sbjct: 1152PQIKAHPWHKLFHKRMPSEAVDLVSRLLQYSPHLRCTALAACAHPFFNDLRD 1307
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,065,240
Number of Sequences: 36976
Number of extensions: 173737
Number of successful extensions: 1601
Number of sequences better than 10.0: 500
Number of HSP's better than 10.0 without gapping: 1269
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1308
length of query: 387
length of database: 9,014,727
effective HSP length: 98
effective length of query: 289
effective length of database: 5,391,079
effective search space: 1558021831
effective search space used: 1558021831
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC124964.6


