

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
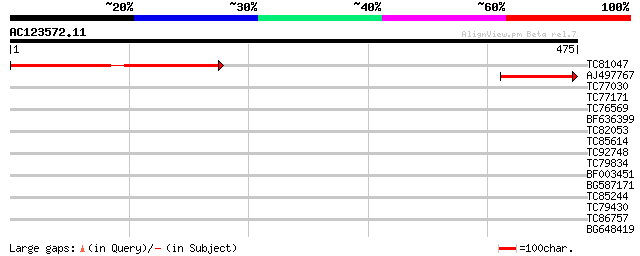
Query= AC123572.11 + phase: 0
(475 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC81047 weakly similar to GP|20268776|gb|AAM14091.1 unknown prot... 285 3e-77
AJ497767 weakly similar to GP|20268776|gb| unknown protein {Arab... 147 7e-36
TC77030 similar to GP|7406669|emb|CAB85628.1 putative ripening-r... 33 0.31
TC77171 similar to PIR|T06562|T06562 CP12 protein precursor chl... 31 0.91
TC76569 homologue to GP|10241560|emb|CAB71128. cationic peroxida... 30 2.0
BF636399 similar to GP|15294065|gb| 40S ribosomal protein S26-2 ... 30 2.0
TC82053 similar to GP|18071664|gb|AAL55431.1 formylglycinamide r... 29 4.5
TC85614 SP|P06452|ATPI_PEA ATP synthase A chain precursor (EC 3.... 29 4.5
TC92748 similar to SP|P53393|SUT3_STYHA Low affinity sulphate tr... 29 4.5
TC79834 similar to GP|16323504|gb|AAL15246.1 unknown protein {Ar... 28 5.9
BF003451 28 5.9
BG587171 SP|P56760|ATPH_ ATP synthase C chain (EC 3.6.3.14) (Lip... 28 7.7
TC85244 similar to GP|2677824|gb|AAB97140.1| abscisic stress rip... 28 7.7
TC79430 similar to PIR|F86358|F86358 Similar to peptide transpor... 28 7.7
TC86757 auxin influx carrier protein [Medicago truncatula] 28 7.7
BG648419 similar to GP|19571124|dbj contains EST AU065194(E60541... 28 7.7
>TC81047 weakly similar to GP|20268776|gb|AAM14091.1 unknown protein
{Arabidopsis thaliana}, partial (35%)
Length = 626
Score = 285 bits (729), Expect = 3e-77
Identities = 147/179 (82%), Positives = 152/179 (84%)
Frame = +3
Query: 1 MANFKCAVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVAS 60
MANFKCAVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVAS
Sbjct: 99 MANFKCAVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVAS 278
Query: 61 FNLLLSIMTISSNLKRAIGGSHVTCGQVSFHNHTLCLFPVWGEGPLGFGLLLSSRITQAF 120
FNLLLSIM + K +C L+ VWGEGPLGFGLLLSSRITQAF
Sbjct: 279 FNLLLSIMMSHNFFKFEKSHWWQSC----------YLWAVWGEGPLGFGLLLSSRITQAF 428
Query: 121 QLYFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHIKKPLSNRCHMSVQWTIPVVCLHA 179
QLYFIFVK+RLPLIRSFLLIPLILLPWIIGAAVIHIKKPL+N CHMSVQW IPVVCLHA
Sbjct: 429 QLYFIFVKKRLPLIRSFLLIPLILLPWIIGAAVIHIKKPLNNPCHMSVQWNIPVVCLHA 605
>AJ497767 weakly similar to GP|20268776|gb| unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 435
Score = 147 bits (372), Expect = 7e-36
Identities = 64/64 (100%), Positives = 64/64 (100%)
Frame = +1
Query: 412 KDYWSSMFFLKFQEECDMRCNGYELEQMTGWNYSPRLSSVHGTDDPFHHDHLLKNSECGN 471
KDYWSSMFFLKFQEECDMRCNGYELEQMTGWNYSPRLSSVHGTDDPFHHDHLLKNSECGN
Sbjct: 1 KDYWSSMFFLKFQEECDMRCNGYELEQMTGWNYSPRLSSVHGTDDPFHHDHLLKNSECGN 180
Query: 472 DTDS 475
DTDS
Sbjct: 181 DTDS 192
>TC77030 similar to GP|7406669|emb|CAB85628.1 putative ripening-related
protein {Vitis vinifera}, partial (51%)
Length = 753
Score = 32.7 bits (73), Expect = 0.31
Identities = 21/88 (23%), Positives = 38/88 (42%)
Frame = +3
Query: 297 PISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYM 356
P + P+ L LLL+ +R+ AD A V LS++ DC +Y
Sbjct: 441 PYENLKPDPILRNLLLSLPYRKLIFTNADKVHA---------VKALSRLGLEDCFEGLYA 593
Query: 357 ARHIIEKYMVAGAAMEINISHRSKQEIL 384
RH+I+ + M++ + +Q ++
Sbjct: 594 LRHLIQSTRIVSLMMKMTLKL*GQQALV 677
>TC77171 similar to PIR|T06562|T06562 CP12 protein precursor chloroplast -
garden pea, partial (93%)
Length = 660
Score = 31.2 bits (69), Expect = 0.91
Identities = 21/68 (30%), Positives = 34/68 (49%), Gaps = 4/68 (5%)
Frame = +3
Query: 285 GIPDSGVLTQSEPISRVDPNEPLDKLLLNKKFRQSFMGFADSC----LAGESVHFFDEVH 340
G+ SG +T P+ R P + ++KK +S ++C ++GE V +DEV
Sbjct: 153 GVAGSGRMTVVRPV-RAAPEQ------ISKKVEESIKSAQETCADDPVSGECVAAWDEVE 311
Query: 341 ELSKISEH 348
ELS + H
Sbjct: 312 ELSAAASH 335
>TC76569 homologue to GP|10241560|emb|CAB71128. cationic peroxidase {Cicer
arietinum}, partial (98%)
Length = 1638
Score = 30.0 bits (66), Expect = 2.0
Identities = 19/62 (30%), Positives = 30/62 (47%)
Frame = -1
Query: 163 RCHMSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTA 222
RC VQW +P +C A AT GV+ + R E+ LW G S+ S +++ +
Sbjct: 879 RCTNFVQWVLPTLC--APSKATTPGVSIPMAPNLSRTAEMDSLWSGRYASNRSTLLFLPS 706
Query: 223 YI 224
+
Sbjct: 705 LL 700
>BF636399 similar to GP|15294065|gb| 40S ribosomal protein S26-2 {Ictalurus
punctatus}, partial (74%)
Length = 534
Score = 30.0 bits (66), Expect = 2.0
Identities = 16/35 (45%), Positives = 21/35 (59%), Gaps = 2/35 (5%)
Frame = -2
Query: 280 MGQALGIPDSGVL--TQSEPISRVDPNEPLDKLLL 312
+GQ +G+ D G+L T I V NEPLD L+L
Sbjct: 239 LGQGVGVDDGGILDITDGGGIHDVPHNEPLDGLVL 135
>TC82053 similar to GP|18071664|gb|AAL55431.1 formylglycinamide
ribonucleotide amidotransferase {Vigna unguiculata},
partial (29%)
Length = 1251
Score = 28.9 bits (63), Expect = 4.5
Identities = 11/36 (30%), Positives = 16/36 (43%)
Frame = +3
Query: 431 CNGYELEQMTGWNYSPRLSSVHGTDDPFHHDHLLKN 466
CNG +L + GW P++ VHG + N
Sbjct: 624 CNGCQLMALLGWVPGPQVGGVHGAGGDLSQPRFIHN 731
>TC85614 SP|P06452|ATPI_PEA ATP synthase A chain precursor (EC 3.6.3.14)
(Subunit IV). [Garden pea] {Pisum sativum}, partial (98%)
Length = 3791
Score = 28.9 bits (63), Expect = 4.5
Identities = 15/36 (41%), Positives = 22/36 (60%)
Frame = +2
Query: 122 LYFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHIK 157
LYFI+ +R L I FLL+ L LL W G ++ ++
Sbjct: 2762 LYFIWNRRNLSWIH*FLLLLLSLLGWPSGLLLLDLE 2869
>TC92748 similar to SP|P53393|SUT3_STYHA Low affinity sulphate transporter
3. {Stylosanthes hamata}, partial (20%)
Length = 536
Score = 28.9 bits (63), Expect = 4.5
Identities = 22/82 (26%), Positives = 36/82 (43%), Gaps = 8/82 (9%)
Frame = +2
Query: 22 SILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVASFNLLLSIMTISSNLKRAI--- 78
S L+F+L + IF + + R K FW+P I + S L I+ +S K+ I
Sbjct: 71 SPLNFVLGCSFLIFLLVTRFIARKKKKLFWLPAIAPLLSVILSTLIVYLSKADKQGINII 250
Query: 79 -----GGSHVTCGQVSFHNHTL 95
G + + Q+ FH +
Sbjct: 251 KHVKGGLNQSSVHQLQFHGQNV 316
>TC79834 similar to GP|16323504|gb|AAL15246.1 unknown protein {Arabidopsis
thaliana}, partial (49%)
Length = 731
Score = 28.5 bits (62), Expect = 5.9
Identities = 14/36 (38%), Positives = 21/36 (57%)
Frame = -3
Query: 63 LLLSIMTISSNLKRAIGGSHVTCGQVSFHNHTLCLF 98
LL+ I+ + NLKR+ + Q+S HN +CLF
Sbjct: 555 LLVGILYVL*NLKRSKSHTLFPITQLSHHNFVICLF 448
>BF003451
Length = 724
Score = 28.5 bits (62), Expect = 5.9
Identities = 22/60 (36%), Positives = 27/60 (44%)
Frame = -2
Query: 28 LLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVASFNLLLSIMTISSNLKRAIGGSHVTCGQ 87
LLL+ S+ F + VPR GS FW P + +LLL M I SH C Q
Sbjct: 177 LLLVLSLDLFPHYSVPRQTGSRFWFPKMFFSNLRHLLLREML-------QIQQSHAKC*Q 19
>BG587171 SP|P56760|ATPH_ ATP synthase C chain (EC 3.6.3.14) (Lipid-binding
protein) (Subunit III). [Mouse-ear cress], partial (96%)
Length = 657
Score = 28.1 bits (61), Expect = 7.7
Identities = 13/35 (37%), Positives = 21/35 (59%)
Frame = -3
Query: 122 LYFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHI 156
L+ F R L I FLL+ L+LL W++G ++ +
Sbjct: 268 LFLYFGVRNLS*IHWFLLLRLLLLGWLLGLLLLDL 164
>TC85244 similar to GP|2677824|gb|AAB97140.1| abscisic stress ripening
protein homolog {Prunus armeniaca}, partial (55%)
Length = 1270
Score = 28.1 bits (61), Expect = 7.7
Identities = 22/73 (30%), Positives = 35/73 (47%), Gaps = 3/73 (4%)
Frame = -1
Query: 10 GGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPV---IQVVASFNLLLS 66
G C T + + ++F+LLLI ++F I+ T GSGF + V V AS
Sbjct: 793 GSCST*LTKMLMFAVAFLLLLIINVFTTII--TITTTGSGFGVRVTISTTVAASSGCFSV 620
Query: 67 IMTISSNLKRAIG 79
+TI++ + G
Sbjct: 619 RVTITTTVAATSG 581
>TC79430 similar to PIR|F86358|F86358 Similar to peptide transporter
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1303
Score = 28.1 bits (61), Expect = 7.7
Identities = 15/40 (37%), Positives = 23/40 (57%)
Frame = +2
Query: 220 VTAYILNEIHDNISWLQVVSRFLLLVLASILVLAFFSISS 259
VT ILN + DN+ W V F + +A I+ L+ FS+ +
Sbjct: 608 VTVSILNYVQDNVGW---VLGFGIPWIAMIIALSLFSLGT 718
>TC86757 auxin influx carrier protein [Medicago truncatula]
Length = 1590
Score = 28.1 bits (61), Expect = 7.7
Identities = 45/202 (22%), Positives = 88/202 (43%), Gaps = 15/202 (7%)
Frame = +1
Query: 34 IFPFIVHKVPRTKGSGFWIP----VIQVVASFNLLLSIMTISSNLKRAIGGSHVTCGQVS 89
++ F H V W P +I ++A+ ++ + ++ + A G + +T
Sbjct: 805 LYTFGGHAVTVEIMHAMWKPQKFKMIYLIATLYVMTLTLPSAAAVYWAFGDNLLT----- 969
Query: 90 FHNHTLCLFPVWGEGPLGFGLLLSSR-ITQAFQ---LYFIFVK------RRLPLIRSFLL 139
H++ L L P G L+L + IT F LYF++ K + L R+ +
Sbjct: 970 -HSNALSLLPRTGFRDTAVILMLIHQFITFGFACTPLYFVWEKFLGVHETKSLLKRALVR 1146
Query: 140 IPLILLPWIIGAAVIHIKKPLSNRC-HMSVQWTIPVVCLHALYVATLVGVTAAVHHIEFR 198
+P+++ W + A + P+++ + V +T+ ++ A ++ T A + +E R
Sbjct: 1147LPVVIPIWFL-AIIFPFFGPINSTVGSLLVSFTVYIIPALA-HMVTFASAPARENAVE-R 1317
Query: 199 FDELRDLWRGILVSSVSVAVWV 220
W G+ +V VAVWV
Sbjct: 1318PPSFLGGWVGLYSVNVFVAVWV 1383
>BG648419 similar to GP|19571124|dbj contains EST AU065194(E60541)~similar to
oligopeptide transporter, partial (16%)
Length = 673
Score = 28.1 bits (61), Expect = 7.7
Identities = 15/40 (37%), Positives = 23/40 (57%)
Frame = +3
Query: 220 VTAYILNEIHDNISWLQVVSRFLLLVLASILVLAFFSISS 259
VT ILN + DN+ W V F + +A I+ L+ FS+ +
Sbjct: 348 VTVSILNYVQDNVGW---VLGFGIPWIAMIIALSLFSLGT 458
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.140 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,447,949
Number of Sequences: 36976
Number of extensions: 309955
Number of successful extensions: 2122
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2089
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2120
length of query: 475
length of database: 9,014,727
effective HSP length: 100
effective length of query: 375
effective length of database: 5,317,127
effective search space: 1993922625
effective search space used: 1993922625
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC123572.11


