

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122161.8 - phase: 0 /pseudo
(473 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
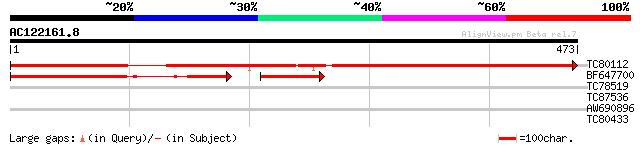
Sequences producing significant alignments: (bits) Value
TC80112 weakly similar to GP|20161371|dbj|BAB90295. hypothetical... 539 e-153
BF647700 similar to GP|20466316|gb| unknown protein {Arabidopsis... 170 2e-60
TC78519 similar to GP|9967495|emb|CAC03535.2 putative protein {A... 30 2.0
TC87536 similar to GP|21553686|gb|AAM62779.1 unknown {Arabidopsi... 29 3.4
AW690896 similar to GP|18087676|gb SRPK4 {Oryza sativa}, partial... 28 5.8
TC80433 similar to GP|8778710|gb|AAF79718.1| T1N15.2 {Arabidopsi... 28 7.6
>TC80112 weakly similar to GP|20161371|dbj|BAB90295. hypothetical
protein~similar to Arabidopsis thaliana chromosome 1
NM_102624.1, partial (31%)
Length = 2068
Score = 539 bits (1388), Expect = e-153
Identities = 309/483 (63%), Positives = 328/483 (66%), Gaps = 10/483 (2%)
Frame = +2
Query: 1 MDDDELELEFSIDDPSISIPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSELPPDSN 60
MDDDELELE SIDDPSISIPRGGPI+VSNMSGPIVRVPLFQDSILTQLHSLQSELPPDSN
Sbjct: 539 MDDDELELECSIDDPSISIPRGGPIFVSNMSGPIVRVPLFQDSILTQLHSLQSELPPDSN 718
Query: 61 HDISVDDLKVFTEDDLMDMALKQVFQGRDNNQDPPNAELVTSLPLFFFIL*YH*LIMMDV 120
HDISVDDLKVFTEDDLMDMALKQVFQGRDNNQDPPNAEL
Sbjct: 719 HDISVDDLKVFTEDDLMDMALKQVFQGRDNNQDPPNAEL--------------------- 835
Query: 121 VVLNNID*ILNLVAGVRRSNSGESLV*QINPFLMLQSNCKEKVEEVVRIKQKQEEDKAQV 180
N+ G R ++ +L SNCKEKVEEVVRIKQKQEEDKAQV
Sbjct: 836 ----------NIGFGCRGQKKQLRRKSRLTNKPILDSNCKEKVEEVVRIKQKQEEDKAQV 985
Query: 181 RLHSFQLVFLIFIHFIMC----KITSI*SNSAIFL*S*LQDQSICQQINQNSKDDVS*VH 236
RLHSF I ++ S+ S S+ + L Q + V H
Sbjct: 986 RLHSFHPDCRINKSANKSIKTQRMMSLKSTSSARKVNTLGLQEHIPVHDSEVVLSVEIFH 1165
Query: 237 KFRQKGQHTGSSGAHT----CAGFRSCSLCRDFP*FSKRCQAIQC-QKKGQHDPSGYFLI 291
FR KG ++ T G ++ S+ RD C Q QK GQHDPSGYFLI
Sbjct: 1166NFR-KGVKLSNAKKKTQELLVLGGQNLSVLRD----KINCSTDQVMQKAGQHDPSGYFLI 1330
Query: 292 EDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLP 351
EDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLP
Sbjct: 1331EDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLP 1510
Query: 352 RFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQ 411
RFASFEMHKIHFCDLGFRLGAGYLYCHQG L + F
Sbjct: 1511RFASFEMHKIHFCDLGFRLGAGYLYCHQG*LYPYLSNTRYEVNSCR*CTELGCLSNCHFS 1690
Query: 412 -LKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTDFIEYD 470
+ F+ CGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTDFIEYD
Sbjct: 1691TARYVFRXCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTDFIEYD 1870
Query: 471 YNH 473
YNH
Sbjct: 1871YNH 1879
>BF647700 similar to GP|20466316|gb| unknown protein {Arabidopsis thaliana},
partial (4%)
Length = 660
Score = 170 bits (431), Expect(2) = 2e-60
Identities = 103/187 (55%), Positives = 122/187 (65%), Gaps = 2/187 (1%)
Frame = +1
Query: 1 MDDDELELEFSIDDPSISIPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSELPPD-S 59
M+DDELE ID+PSISIPRGGPIYVSNMSGPIVRVPLFQDS+L+QLHSLQSELPP
Sbjct: 67 MEDDELECSIGIDNPSISIPRGGPIYVSNMSGPIVRVPLFQDSLLSQLHSLQSELPPQHH 246
Query: 60 NHDIS-VDDLKVFTEDDLMDMALKQVFQGRDNNQDPPNAELVTSLPLFFFIL*YH*LIMM 118
++DI+ VDDLKVFTED+LMDM+LKQVFQ + N +D + + LF
Sbjct: 247 DYDITVVDDLKVFTEDELMDMSLKQVFQNQPNAEDHHHQK----KKLF------------ 378
Query: 119 DVVVLNNID*ILNLVAGVRRSNSGESLV*QINPFLMLQSNCKEKVEEVVRIKQKQEEDKA 178
RR + ++ +L SNCKEKV++VVRIK KQEEDKA
Sbjct: 379 ------------------RRKS-------RLTNKSILDSNCKEKVDKVVRIKHKQEEDKA 483
Query: 179 QVRLHSF 185
QV LHSF
Sbjct: 484 QVTLHSF 504
Score = 80.5 bits (197), Expect(2) = 2e-60
Identities = 40/53 (75%), Positives = 43/53 (80%)
Frame = +3
Query: 210 FL*S*LQDQSICQQINQNSKDDVS*VHKFRQKGQHTGSSGAHTCAGFRSCSLC 262
F+ S LQDQSICQQINQNSKDDVS VH F QH+ SS AHTCAGF+SCSLC
Sbjct: 498 FISSSLQDQSICQQINQNSKDDVSXVH*FCH*AQHSRSSXAHTCAGFKSCSLC 656
>TC78519 similar to GP|9967495|emb|CAC03535.2 putative protein {Arabidopsis
thaliana}, partial (74%)
Length = 2160
Score = 30.0 bits (66), Expect = 2.0
Identities = 15/42 (35%), Positives = 21/42 (49%)
Frame = +1
Query: 215 LQDQSICQQINQNSKDDVS*VHKFRQKGQHTGSSGAHTCAGF 256
L + +I QQI + DD V K +H+G +G H C F
Sbjct: 379 LDEITILQQIAEGDTDDKKCVVKLLDHFKHSGPNGQHVCMVF 504
>TC87536 similar to GP|21553686|gb|AAM62779.1 unknown {Arabidopsis
thaliana}, partial (94%)
Length = 1358
Score = 29.3 bits (64), Expect = 3.4
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = -1
Query: 256 FRSCSLCRDFP*FSKRCQAIQCQKKGQHDPSGYFLIEDVFYT 297
FR+ S CR F++ + Q KG H P G+FLI +T
Sbjct: 677 FRTDSTCRRSFYFNRSFWCSRRQPKGLHCPKGFFLIGSDLHT 552
>AW690896 similar to GP|18087676|gb SRPK4 {Oryza sativa}, partial (23%)
Length = 590
Score = 28.5 bits (62), Expect = 5.8
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = +2
Query: 215 LQDQSICQQINQNSKDDVS*VHKFRQKGQHTGSSGAHTCAGF 256
+ + I +QI + DD V K +H+G +G H C F
Sbjct: 389 MDEIKILKQIEEGDPDDKKCVVKLLDHFKHSGPNGVHVCMVF 514
>TC80433 similar to GP|8778710|gb|AAF79718.1| T1N15.2 {Arabidopsis
thaliana}, partial (13%)
Length = 794
Score = 28.1 bits (61), Expect = 7.6
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +2
Query: 424 IFRATKVTVDDKWTPDNPCYFC 445
++R TK T+ + P NPC FC
Sbjct: 272 LWRPTKATLIPRAAPSNPCIFC 337
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.334 0.146 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,800,794
Number of Sequences: 36976
Number of extensions: 243497
Number of successful extensions: 1735
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 1717
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1734
length of query: 473
length of database: 9,014,727
effective HSP length: 100
effective length of query: 373
effective length of database: 5,317,127
effective search space: 1983288371
effective search space used: 1983288371
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC122161.8


