

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121244.7 - phase: 0
(418 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
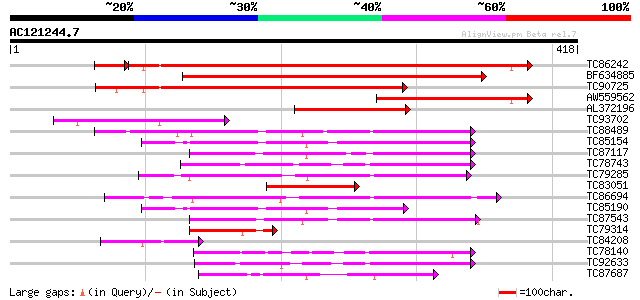
Score E
Sequences producing significant alignments: (bits) Value
TC86242 similar to GP|14486705|gb|AAK63247.1 phosphatidylinosito... 362 e-102
BF634885 similar to GP|11994666|db phosphatidylinositol/phosphat... 262 2e-70
TC90725 similar to GP|14335006|gb|AAK59767.1 AT4g39170/T22F8_70 ... 229 1e-60
AW559562 similar to GP|14486705|gb phosphatidylinositol transfer... 141 4e-34
AL372196 homologue to GP|14486705|gb| phosphatidylinositol trans... 111 6e-25
TC93702 similar to GP|16209642|gb|AAL14382.1 At2g21540/F2G1.19 {... 103 1e-22
TC88489 similar to GP|14334978|gb|AAK59666.1 unknown protein {Ar... 74 1e-13
TC85154 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 72 5e-13
TC87117 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 71 9e-13
TC78743 similar to PIR|T05953|T05953 polyphosphoinositide bindin... 70 1e-12
TC79285 similar to PIR|T05949|T05949 phosphatidylinositol-phosph... 69 4e-12
TC83051 similar to GP|21537211|gb|AAM61552.1 thylakoid lumen pro... 68 7e-12
TC86694 similar to PIR|T05949|T05949 phosphatidylinositol-phosph... 66 2e-11
TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 66 3e-11
TC87543 similar to PIR|A96745|A96745 probable cytosolic factor T... 60 2e-09
TC79314 similar to GP|16930483|gb|AAL31927.1 AT3g51670/T18N14_50... 58 8e-09
TC84208 similar to GP|8778498|gb|AAF79506.1| F20N2.11 {Arabidops... 54 9e-08
TC78140 similar to PIR|H96781|H96781 unknown protein F22H5.20 [i... 52 4e-07
TC92633 similar to PIR|T05953|T05953 polyphosphoinositide bindin... 49 4e-06
TC87687 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 49 4e-06
>TC86242 similar to GP|14486705|gb|AAK63247.1 phosphatidylinositol
transfer-like protein III {Lotus japonicus}, partial
(79%)
Length = 2271
Score = 362 bits (930), Expect(2) = e-102
Identities = 184/346 (53%), Positives = 238/346 (68%), Gaps = 48/346 (13%)
Frame = +2
Query: 88 KFKHSFKKKR-------------VDELQAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMM 134
KF+HS +KK +++++ V + AV DAFR LI D LLP +HDDYHM+
Sbjct: 314 KFRHSLRKKSSRRKNPSRSNSLAIEDVRDVKELQAV-DAFRHSLISDCLLPSRHDDYHML 490
Query: 135 LRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEG 194
LRFLKARKFDI KAK MWA+M++WRKE+G DTIMEDFEF+EL+EV++Y PHGYHGVDKEG
Sbjct: 491 LRFLKARKFDIEKAKLMWANMIQWRKEYGTDTIMEDFEFSELDEVLQYYPHGYHGVDKEG 670
Query: 195 RPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILD 254
RPV+IER K+D NKLMQVTT++RY+KYH Q E+ A+KFPAC+IA+KRHIDSS TILD
Sbjct: 671 RPVYIERLGKVDPNKLMQVTTMERYLKYHVQGFEKTFAVKFPACSIAAKRHIDSSTTILD 850
Query: 255 LQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVA 314
+ G+GF N + RE+++R KI D YP+T + FIIN G + L + + F+DPK +
Sbjct: 851 VHGVGFKNFSKIARELIQRLQKIDGDYYPETLCRMFIINAGPGFKLLWNTVKTFLDPKTS 1030
Query: 315 SKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPWNNPEILK------ 368
SK++V+G+++Q KL ++ID SELP FLGG+CTC +QGGC+RSDKGPW +P ILK
Sbjct: 1031SKINVLGNKFQSKLFEIIDVSELPEFLGGSCTCIDQGGCMRSDKGPWQDPNILKMVLNGE 1210
Query: 369 -----------------------------VKGSDTLTAESSSEAED 385
++ SDT TAES SE ED
Sbjct: 1211IQCSRQIVTISNDEGTVIECDKASFPMVQIRSSDTSTAESGSEVED 1348
Score = 28.9 bits (63), Expect(2) = e-102
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +1
Query: 63 FLTGSDENRKERRSDFKKKLLNAFSK 88
F D+NR+ R KKK++NA SK
Sbjct: 238 FENSEDDNRRTRNGSLKKKVINASSK 315
>BF634885 similar to GP|11994666|db phosphatidylinositol/phosphatidylcholine
transfer protein-like {Arabidopsis thaliana}, partial
(35%)
Length = 684
Score = 262 bits (669), Expect = 2e-70
Identities = 119/224 (53%), Positives = 163/224 (72%)
Frame = +3
Query: 128 HDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGY 187
HD+YH MLRFLK RKFD+ K MWADML WRKE+GAD+I++DF + E EV +Y PHG
Sbjct: 9 HDNYHTMLRFLKGRKFDLDKTVQMWADMLHWRKEYGADSILQDFVYKEFEEVQRYYPHGM 188
Query: 188 HGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHID 247
HGVDKEGRPV+IER K+D KLM VTTIDR+++YH Q E++ KFPAC+IA+KRHID
Sbjct: 189 HGVDKEGRPVYIERLGKVDPIKLMNVTTIDRFLRYHVQGFEKLFQEKFPACSIAAKRHID 368
Query: 248 SSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEY 307
+ TI+D+ G+ + + + E++ R KI DNYP+T Q FI+N G + + + +
Sbjct: 369 KTTTIIDVHGVNWLSFSKIAHELVMRMQKIDGDNYPETLNQMFIVNAGSGFKLVWNTAKG 548
Query: 308 FMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQG 351
F+DPK +K+ V+G ++Q +LL+VID+ +LP FLGG+C+C N G
Sbjct: 549 FLDPKTTAKIIVLGSKFQSRLLEVIDSCQLPDFLGGSCSCPNDG 680
>TC90725 similar to GP|14335006|gb|AAK59767.1 AT4g39170/T22F8_70
{Arabidopsis thaliana}, partial (39%)
Length = 831
Score = 229 bits (585), Expect = 1e-60
Identities = 125/251 (49%), Positives = 167/251 (65%), Gaps = 21/251 (8%)
Frame = +1
Query: 64 LTGSDENRKERRSD-------FKKKLLNAFSKFKHSFKKKR-------------VDELQA 103
L G+DE R+ D KKK +NA K +HS KKKR +++++
Sbjct: 79 LLGNDERRESSEDDKWKRIGSLKKKAINASCKLRHSLKKKRSSKSSGSRSNSLSIEDVRQ 258
Query: 104 VNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFG 163
V + AV DAFR+ L++D LLP HDDYHM+LRFLKARKFDI KAK MWA+M++WRK++G
Sbjct: 259 VEELNAV-DAFRQTLLLDNLLPSIHDDYHMLLRFLKARKFDIEKAKLMWANMIQWRKDYG 435
Query: 164 ADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVF-IERFEKLDRNKLMQVTTIDRYVKY 222
DTI+EDF+F EL EV KY G+ +F IER K+D N+LMQVTT++RY++Y
Sbjct: 436 TDTIIEDFDFKELQEVQKYLSTWLSWCG*GGKTLFYIERLGKVDPNRLMQVTTMERYLRY 615
Query: 223 HAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNY 282
H Q E+ ++KFPAC+IA+KR I SS TILD+QG+GF N ++ RE++ + KI D Y
Sbjct: 616 HVQGFEKTFSVKFPACSIAAKRRIGSSTTILDVQGVGFKNFTKSARELIIQLQKIDSDYY 795
Query: 283 PQTGGQSFIIN 293
P+T Q FIIN
Sbjct: 796 PETLHQMFIIN 828
>AW559562 similar to GP|14486705|gb phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (34%)
Length = 668
Score = 141 bits (356), Expect = 4e-34
Identities = 77/148 (52%), Positives = 91/148 (61%), Gaps = 33/148 (22%)
Frame = +3
Query: 271 MKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLK 330
++R K+ DNY +T Q FIIN G R L S + F+DPK SK+HV+G++YQ KLL+
Sbjct: 3 IQRLQKVDCDNYFETFCQMFIINAGPGFRLLWSTVKSFLDPKTTSKIHVLGNKYQSKLLE 182
Query: 331 VIDASELPTFLGGTCTCANQGGCLRSDKGPWNNPEILK---------------------- 368
VIDASELP FLGGTC+CA++GGCLRSDKGPW NPEILK
Sbjct: 183 VIDASELPEFLGGTCSCADEGGCLRSDKGPWKNPEILKMVLNGEPRRARQVVKVLNSEGK 362
Query: 369 -----------VKGSDTLTAESSSEAED 385
VKGSDT TAES SEAED
Sbjct: 363 VIAYAKPRYPMVKGSDTSTAESGSEAED 446
>AL372196 homologue to GP|14486705|gb| phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (14%)
Length = 264
Score = 111 bits (277), Expect = 6e-25
Identities = 54/85 (63%), Positives = 66/85 (77%)
Frame = +1
Query: 211 MQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREI 270
MQVTT+DRYVKYH Q E+ AIKFPACTIA+KRHIDSS TILD+QG+G N ++ RE+
Sbjct: 1 MQVTTMDRYVKYHVQEFEKSFAIKFPACTIAAKRHIDSSTTILDVQGVGLKNFTKSAREL 180
Query: 271 MKRFLKILIDNYPQTGGQSFIINVG 295
++R K+ DNYP+T Q FIIN G
Sbjct: 181 IQRLQKVDGDNYPETLCQMFIINAG 255
>TC93702 similar to GP|16209642|gb|AAL14382.1 At2g21540/F2G1.19 {Arabidopsis
thaliana}, partial (19%)
Length = 473
Score = 103 bits (257), Expect = 1e-22
Identities = 60/141 (42%), Positives = 82/141 (57%), Gaps = 11/141 (7%)
Frame = +1
Query: 33 GCILFIHSSEMLRMIF--VFGLVIDSSIVMFWFLTGSDENRKERRSDFKKKLLNAFSKFK 90
GCI S +R I V +D + + + S++ +K++ KK L+A SKFK
Sbjct: 49 GCI*SFELSCNIRSIMCDVMPAPLDRPVKIGHEMEHSEDEKKKKVGSIKKVALSASSKFK 228
Query: 91 HSFKKKRVDELQAVNVVPA---------VVDAFRKLLIMDELLPQKHDDYHMMLRFLKAR 141
+SF KK + +++ VDA R+ LI++ELLP KHDD HMMLRFL+AR
Sbjct: 229 NSFTKKGRKHSRVMSICIEDSFDAEELQAVDALRQTLILEELLPSKHDDPHMMLRFLRAR 408
Query: 142 KFDIGKAKHMWADMLEWRKEF 162
K+DI K K MW DML+WRKEF
Sbjct: 409 KYDIEKTKQMWTDMLKWRKEF 471
>TC88489 similar to GP|14334978|gb|AAK59666.1 unknown protein {Arabidopsis
thaliana}, partial (47%)
Length = 1187
Score = 73.9 bits (180), Expect = 1e-13
Identities = 75/305 (24%), Positives = 136/305 (44%), Gaps = 24/305 (7%)
Frame = +3
Query: 63 FLTGSDENRKERRSDFKKKLLNAFSKFKHSFKKKRVDELQAVNVVPAVVDAFRKLLIMDE 122
FL+ E K+ +F+ K+ A ++ +K+ +E + V +P + + ++ +E
Sbjct: 234 FLSDLKEFEKKALIEFRSKVEEAV--LGNTLFEKKEEETKKVETLPEGGEESSEKVVKEE 407
Query: 123 -------------------LLPQKHDDYH--MMLRFLKARKFDIGKAKHMWADMLEWRKE 161
LLP K ++ ++L+FL+AR+F + +A M L+WRKE
Sbjct: 408 EEKRVEEVEEKDLSLWGVSLLPSKGNEGVDVVLLKFLRAREFKVNEAFEMLKKTLKWRKE 587
Query: 162 FGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVT--TIDRY 219
D+++E+ ++L N GVD+EG PV + D ++ Q T T ++
Sbjct: 588 MKIDSVLEEDFGSDLASAAYMN-----GVDREGHPVCYNIYSVFDGEEIYQKTFGTEEKR 752
Query: 220 VKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILI 279
++ RC M K+ + S + I DL+ + +E R K+ + +L
Sbjct: 753 KEFLRWRCSVME--KWIQKLNLKPGGVSSLLQINDLKNSPGPSKKEL-RIATKQTVTMLQ 923
Query: 280 DNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGD-RYQRKLLKVIDASELP 338
DNYP+ ++ INV +L ++ F+ + SK V + L+K I E+P
Sbjct: 924 DNYPELVAKNIFINVPFWYYALNALLSPFLTQRTKSKFVVARPAKVTETLIKYIPIEEIP 1103
Query: 339 TFLGG 343
GG
Sbjct: 1104VNYGG 1118
>TC85154 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (83%)
Length = 1811
Score = 71.6 bits (174), Expect = 5e-13
Identities = 65/253 (25%), Positives = 118/253 (45%), Gaps = 7/253 (2%)
Frame = +3
Query: 98 VDELQAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLE 157
V E++ +VP V+ + L+ DE D ++L+FL+AR F + +A M +
Sbjct: 399 VAEVEVTPIVPEEVEIWGIPLLADE-----RSDV-ILLKFLRARDFKVKEAFTMIKQTVL 560
Query: 158 WRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTID 217
WRKEFG + ++++ + ++V+ + G DKEG PV+ F + + L Q T D
Sbjct: 561 WRKEFGVEALLQEDLGTDWDKVVFTD-----GTDKEGHPVYYNVFGEFEDKDLYQKTFSD 725
Query: 218 -----RYVKYHAQRCEE-MHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIM 271
++V++ Q E+ + + F + I + + I DL+ ++ R+ +
Sbjct: 726 EEKRTKFVRWWIQSLEKSVRKLDF------APSGISTLVQINDLKNSPGLLGKKELRQSI 887
Query: 272 KRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIG-DRYQRKLLK 330
K+ L++L DNYP+ + INV + + F+ + SK G + L K
Sbjct: 888 KQTLQLLQDNYPEFVAKQIFINVPWWYLAFSRMISPFLTQRTKSKFVFAGSSKSAETLFK 1067
Query: 331 VIDASELPTFLGG 343
I ++P GG
Sbjct: 1068YIAPEQVPVKYGG 1106
>TC87117 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (72%)
Length = 1662
Score = 70.9 bits (172), Expect = 9e-13
Identities = 59/218 (27%), Positives = 101/218 (46%), Gaps = 7/218 (3%)
Frame = +3
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDK 192
++L+FL+AR F + +A M + + WRKEFG + +M++ +EL +V+ HG DK
Sbjct: 531 ILLKFLRARDFKVKEAFTMIKNTILWRKEFGIEELMDEKLGDELEKVVY-----MHGFDK 695
Query: 193 EGRPVFIERFEKLDRNKLMQVTTID-----RYVKYHAQRCEE-MHAIKFPACTIASKRHI 246
EG PV + + +L T D ++K+ Q E+ + + F + + H+
Sbjct: 696 EGHPVCYNIYGEFQNKELYNKTFSDEEKRHNFLKWRIQFLEKSIRNLDFNHGGVCTIVHV 875
Query: 247 DSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICE 306
+ + D G G L R+ K+ L++ DNYP+ + INV ++ +
Sbjct: 876 ND---LKDSPGPGKWEL----RQATKQALQLFQDNYPEFVAKQVFINVPWWYLAVNRMIS 1034
Query: 307 YFMDPKVASKVHVIG-DRYQRKLLKVIDASELPTFLGG 343
F+ + SK G + LL I +LP GG
Sbjct: 1035PFLTQRTKSKFVFAGPSKSTETLLSYIAPEQLPVKYGG 1148
>TC78743 similar to PIR|T05953|T05953 polyphosphoinositide binding protein
Ssh2 - soybean, partial (94%)
Length = 1232
Score = 70.5 bits (171), Expect = 1e-12
Identities = 60/218 (27%), Positives = 99/218 (44%), Gaps = 1/218 (0%)
Frame = +1
Query: 127 KHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHG 186
K D M+ RFL+AR D+ KA M+ ++WRK F + +E+ + +
Sbjct: 385 KEVDDLMIRRFLRARDLDVDKASAMFLKYMKWRKSFVPSGSVSP---SEIADDLAQEKIY 555
Query: 187 YHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHI 246
G+DK+GRP+ + K +NK +D + +Y E++ + P
Sbjct: 556 VQGLDKKGRPIIVAFAAKHFQNK----NGLDAFKRYVVFALEKLISRMPPG--------E 699
Query: 247 DSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICE 306
+ ++I D++G G+ N +D L IL D YP+ G+ FI++ + I
Sbjct: 700 EKFVSIADIKGWGYAN---SDIRGYLGALTILQDYYPERLGKLFIVHAPYMFMKVWKIIY 870
Query: 307 YFMDPKVASK-VHVIGDRYQRKLLKVIDASELPTFLGG 343
F+D K V V + + LL+ ID S+LP GG
Sbjct: 871 PFIDDNTKKKIVFVENKKLKATLLEEIDESQLPEIYGG 984
>TC79285 similar to PIR|T05949|T05949
phosphatidylinositol-phosphatidylcholine transfer
protein SEC14 homolog Ssh1 - soybean, partial (68%)
Length = 842
Score = 68.6 bits (166), Expect = 4e-12
Identities = 62/253 (24%), Positives = 112/253 (43%), Gaps = 8/253 (3%)
Frame = +3
Query: 96 KRVDELQAVNVVPAVVDAFRKLLIMDELLPQKHDDY--HMMLRFLKARKFDIGKAKHMWA 153
K++D ++ + + VD +K+ + H Y M+ RFLKAR ++ KA+ M
Sbjct: 129 KKMDAIKQLQSLMENVDEQQKIAFKN-----LHQGYPTEMLARFLKARDGNVAKAQKMLI 293
Query: 154 DMLEWRKEFGADTIMEDFEFNELNEVIKYNP-HGYHGVDKEGRPVFIERFEKLDRNKLMQ 212
D L WR E D ++ +L + ++ + G G KEG PV +
Sbjct: 294 DCLHWRVENEIDKVLAKPIPADLYKPVRDSQLIGMSGYTKEGLPVIAVG---------VG 446
Query: 213 VTTIDR-----YVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEAD 267
++T D+ Y++ H Q E + P T R+I + + +LD+ G+ F L +
Sbjct: 447 LSTYDKASDKYYIQSHIQVNEYRDRVILPTATKKHGRYIGTCVKVLDMTGLKFSALNQL- 623
Query: 268 REIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRK 327
++ I NYP+ +I+N + + + + + K+ V+ + +
Sbjct: 624 -RLLTAISTIDDLNYPEKTDIYYIVNAPYVFSACWKVVKPLLQERTRKKIQVLQGCGKEE 800
Query: 328 LLKVIDASELPTF 340
LLKV+D + LP F
Sbjct: 801 LLKVMDYAFLPHF 839
>TC83051 similar to GP|21537211|gb|AAM61552.1 thylakoid lumen protein
chloroplast precursor {Arabidopsis thaliana}, partial
(56%)
Length = 915
Score = 67.8 bits (164), Expect = 7e-12
Identities = 29/69 (42%), Positives = 48/69 (69%)
Frame = +2
Query: 190 VDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSS 249
+ ++ R V+IER +D NKL Q+TT +R +K+H E+ +++PAC++A+KRHI S+
Sbjct: 659 IQRQDRSVYIERIGMIDVNKLWQITTQERLIKHHVSEQEKTLRVRYPACSLAAKRHIAST 838
Query: 250 ITILDLQGI 258
+ILD G+
Sbjct: 839 TSILDGNGV 865
>TC86694 similar to PIR|T05949|T05949
phosphatidylinositol-phosphatidylcholine transfer
protein SEC14 homolog Ssh1 - soybean, partial (98%)
Length = 1433
Score = 66.2 bits (160), Expect = 2e-11
Identities = 74/305 (24%), Positives = 129/305 (42%), Gaps = 13/305 (4%)
Frame = +3
Query: 71 RKERRSDFKKKLLNAFSKFKHSFKKKRVDELQAVNVVPAVVDAFRKLLIMDELLPQKHDD 130
RK + S F K L +F+ F + D A+N + A++D ++E L + +
Sbjct: 111 RKTKSSVFFKVLAFSFAVFLITMGIVSQD---ALNQLQALIDQ------VEEPLQKTFQN 263
Query: 131 YHM------MLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIM-EDFEFNELNEVIKYN 183
H ++RFLKAR+++ KA M D L WR + D I+ + +L ++ +
Sbjct: 264 VHQGHVTETLIRFLKAREWNASKAHKMLIDSLNWRVQNEIDKILSKPIIPQDLYRGLRDS 443
Query: 184 P-HGYHGVDKEGRPVF-----IERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPA 237
G G +EG PVF + F+K ++ YV+ H Q E + P+
Sbjct: 444 QLIGLSGYSREGLPVFAIGVGLSTFDK---------ASVHYYVQSHIQINEYRDRVILPS 596
Query: 238 CTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLE 297
+ R I + + +LD+ G+ L + +++ I NYP+ +I+N
Sbjct: 597 ASKKHGRPITTCVKVLDMTGLKLSALNQI--KLLTIISSIDDLNYPEKTNTYYIVNAPYI 770
Query: 298 LRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSD 357
+ + + + KV V+ + +LLK++D + LP F C G S
Sbjct: 771 FSGCWKVVKPLLQERTRKKVQVLQGCGRDELLKIMDYACLPHF----CKKEGSGSSKHSG 938
Query: 358 KGPWN 362
G N
Sbjct: 939 SGSEN 953
>TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (50%)
Length = 1496
Score = 65.9 bits (159), Expect = 3e-11
Identities = 55/203 (27%), Positives = 101/203 (49%), Gaps = 6/203 (2%)
Frame = +3
Query: 98 VDELQAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLE 157
V E++ +VP V+ + L+ DE D ++L+FL+AR F + +A M +
Sbjct: 855 VAEVEVTPIVPEEVEIWGIPLLADE-----RSDV-ILLKFLRARDFKVKEAFTMIKQTVL 1016
Query: 158 WRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTID 217
WRKEFG + ++++ + ++V+ + G DKEG PV+ F + + L Q T D
Sbjct: 1017 WRKEFGVEALLQEDLGTDWDKVVFTD-----GTDKEGHPVYYNVFGEFEDKDLYQKTFSD 1181
Query: 218 -----RYVKYHAQRCEE-MHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIM 271
++V++ Q E+ + + F + I + + I DL+ ++ R+ +
Sbjct: 1182 EEKRTKFVRWWIQSLEKSVRKLDF------APSGISTLVQINDLKNSPGLLGKKELRQSI 1343
Query: 272 KRFLKILIDNYPQTGGQSFIINV 294
K+ L++L DNYP+ + INV
Sbjct: 1344 KQTLQLLQDNYPEFVAKQIFINV 1412
>TC87543 similar to PIR|A96745|A96745 probable cytosolic factor T9N14.8
[imported] - Arabidopsis thaliana, partial (74%)
Length = 1718
Score = 59.7 bits (143), Expect = 2e-09
Identities = 55/226 (24%), Positives = 102/226 (44%), Gaps = 11/226 (4%)
Frame = +1
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDK 192
++L+FL+AR F ++ M + L+WRK F D ++++ ++L++V+ HG +
Sbjct: 511 ILLKFLRARDFKPKESHTMLKNTLQWRKSFNIDALLDEDLGDDLDKVVFM-----HGFSR 675
Query: 193 EGRPVFIERFEKLDRNKLMQVT-----TIDRYVKYHAQRCEE-MHAIKFPACTIASKRHI 246
EG PV + + +L + T +R++++ Q E+ + + F S +
Sbjct: 676 EGHPVCYNVYGEFQNKELYEKTFGSEEKRERFLRWRVQFLEKSIRKLDF------SPGGV 837
Query: 247 DSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICE 306
++ + DL+ +E R K L++L DNYP+ + INV + +I
Sbjct: 838 NTLFQVNDLKNSPGPAKKEL-RVATKMALELLQDNYPEFVAKQVFINVPWWYLAFYTILN 1014
Query: 307 YFMDPKVASKVHVIG-DRYQRKLLKVIDASELPTFLGGT----CTC 347
F+ + SK G + L K I ++P GG C C
Sbjct: 1015PFLTQRTKSKFVFAGTSKSPDTLFKYITPEQVPVQYGGLSVDFCDC 1152
>TC79314 similar to GP|16930483|gb|AAL31927.1 AT3g51670/T18N14_50
{Arabidopsis thaliana}, partial (33%)
Length = 868
Score = 57.8 bits (138), Expect = 8e-09
Identities = 32/67 (47%), Positives = 42/67 (61%), Gaps = 2/67 (2%)
Frame = +1
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMED--FEFNELNEVIKYNPHGYHGV 190
++L+FL+AR F + +A +M L WRKEF ADTI+E+ +F EL VI Y G
Sbjct: 493 ILLKFLRARDFRVNEALNMITKCLAWRKEFNADTIVEEDLSQFKELEGVIAY----MQGY 660
Query: 191 DKEGRPV 197
DKEG PV
Sbjct: 661 DKEGHPV 681
>TC84208 similar to GP|8778498|gb|AAF79506.1| F20N2.11 {Arabidopsis
thaliana}, partial (13%)
Length = 664
Score = 54.3 bits (129), Expect = 9e-08
Identities = 31/88 (35%), Positives = 47/88 (53%), Gaps = 12/88 (13%)
Frame = +1
Query: 68 DENRKERRSDFKKKLLNAFSKFKHSFKKK------------RVDELQAVNVVPAVVDAFR 115
D + + +KK +NA SKF HS KK+ +++++ AV++ R
Sbjct: 403 DRRQYSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPSVAIEDVRDAREETAVLE-LR 579
Query: 116 KLLIMDELLPQKHDDYHMMLRFLKARKF 143
+ L+ LP +HDDYH +LRFLKAR F
Sbjct: 580 QRLVERGSLPSRHDDYHTLLRFLKARDF 663
>TC78140 similar to PIR|H96781|H96781 unknown protein F22H5.20 [imported] -
Arabidopsis thaliana, partial (83%)
Length = 1253
Score = 52.0 bits (123), Expect = 4e-07
Identities = 52/211 (24%), Positives = 89/211 (41%), Gaps = 3/211 (1%)
Frame = +2
Query: 136 RFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGR 195
R+L+AR +++ K+K M + L+WR + + I D E E K G+H D++GR
Sbjct: 293 RYLEARNWNVDKSKKMLKETLKWRSVYKPEEIRWD-EVAVEGETGKMYRAGFH--DRQGR 463
Query: 196 PVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDL 255
V I R + ++ID +K H E + P + ++D
Sbjct: 464 TVLILR------PGMQNTSSIDNQIK-HLVYLLENAMLNLPPGQ-------EQMAWLIDF 601
Query: 256 QGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVAS 315
G N + + + IL ++YP+ G +F+ N + I +YF+D K
Sbjct: 602 TGWSITN--NVPLKSARETISILQNHYPERLGIAFLYNPPRIFEAFWKIVKYFLDNKTFH 775
Query: 316 KVHVIGDRYQ---RKLLKVIDASELPTFLGG 343
KV + + + + D LP+ LGG
Sbjct: 776 KVKFVYPKNKDSVELMRSYFDDENLPSELGG 868
>TC92633 similar to PIR|T05953|T05953 polyphosphoinositide binding protein
Ssh2 - soybean, partial (89%)
Length = 806
Score = 48.9 bits (115), Expect = 4e-06
Identities = 53/212 (25%), Positives = 87/212 (41%), Gaps = 5/212 (2%)
Frame = +1
Query: 137 FLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRP 196
F R D+ KA M+ L+WR F + +++ I + G DK GRP
Sbjct: 157 FYVPRDLDVEKASAMFLKYLKWRHSFVPNG---SISLSQVPNQIAQDKAFAQGHDKIGRP 327
Query: 197 VFI----ERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITI 252
+ + F+K D +D + ++ +++ A P + + I
Sbjct: 328 ILLVFGGRHFQKKDG--------LDEFKRFAVYILDKLCASMPPG--------EEKFVGI 459
Query: 253 LDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPK 312
+L+G G+ N +D L IL D YP+ G+ FI++ + I F+D K
Sbjct: 460 AELKGWGYSN---SDLRGYIGALSILQDYYPERLGKLFILHAPYIFMKVWKIVYPFIDNK 630
Query: 313 VASK-VHVIGDRYQRKLLKVIDASELPTFLGG 343
K V V ++ + LL ID S+LP G
Sbjct: 631 TKKKIVFVENNKLKSTLLXDIDESQLPEIYXG 726
>TC87687 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (67%)
Length = 1969
Score = 48.9 bits (115), Expect = 4e-06
Identities = 46/195 (23%), Positives = 80/195 (40%), Gaps = 18/195 (9%)
Frame = +1
Query: 140 ARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFI 199
AR F + +A M + + WRKEF + ++ D + E ++ + HG DKEG PV
Sbjct: 766 ARDFKVKEAFTMIKNTIRWRKEFKIEELLLDENLGD--EYLEKTVY-MHGYDKEGHPVCY 936
Query: 200 ERFEKLDRNKLMQVTTID-----RYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILD 254
+ + D ++ + T D R++K+ E+ SI LD
Sbjct: 937 NIYGEFDNKEVYKKTFSDEEKRERFLKWRILVLEK-------------------SIRKLD 1059
Query: 255 LQGIGFCNLEEAD-------------REIMKRFLKILIDNYPQTGGQSFIINVGLELRSL 301
G C + + + R+ K+ L++L DNYP+ + INV ++
Sbjct: 1060FTPGGICTIVQVNDLKNSPGPTKWELRQATKQALQLLQDNYPEFVAKQVFINVPWWYLAV 1239
Query: 302 RSICEYFMDPKVASK 316
+ F+ + SK
Sbjct: 1240NRMISPFLTQRTKSK 1284
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.140 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,212,420
Number of Sequences: 36976
Number of extensions: 182882
Number of successful extensions: 1304
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 1271
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1284
length of query: 418
length of database: 9,014,727
effective HSP length: 99
effective length of query: 319
effective length of database: 5,354,103
effective search space: 1707958857
effective search space used: 1707958857
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121244.7


