

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
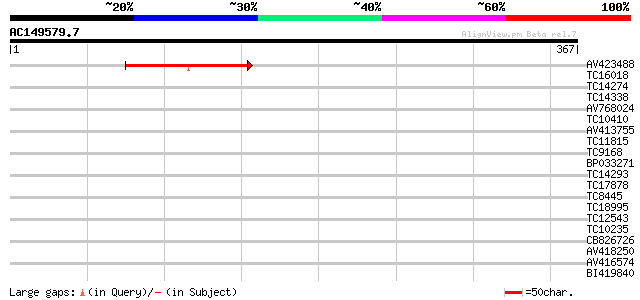
Query= AC149579.7 - phase: 0 /pseudo
(367 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV423488 105 1e-23
TC16018 UP|Q8MZB8 (Q8MZB8) AT15479p (CG31542-PA), partial (7%) 35 0.020
TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.3... 32 0.13
TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%) 32 0.22
AV768024 31 0.28
TC10410 weakly similar to UP|Q8H5W8 (Q8H5W8) OJ1123_B01.10 prote... 31 0.28
AV413755 31 0.37
TC11815 weakly similar to UP|Q8MLU0 (Q8MLU0) CG13503-PA, partial... 31 0.37
TC9168 similar to GB|AAP21369.1|30102902|BT006561 At4g19200 {Ara... 31 0.37
BP033271 30 0.49
TC14293 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding pr... 30 0.49
TC17878 similar to GB|AAK91488.1|15215889|AY050475 AT5g02770/F9G... 30 0.49
TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial ... 30 0.83
TC18995 weakly similar to UP|Q8PB28 (Q8PB28) Serine protease, pa... 30 0.83
TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 29 1.1
TC10235 29 1.4
CB826726 28 1.8
AV418250 28 1.8
AV416574 28 2.4
BI419840 28 2.4
>AV423488
Length = 487
Score = 105 bits (262), Expect = 1e-23
Identities = 51/84 (60%), Positives = 65/84 (76%), Gaps = 2/84 (2%)
Frame = +3
Query: 76 NGSMPMENDNFHNVGATQYPEFSTQITPSGMAIADEVTP--EDSTPKSKRSKEPAWNTQQ 133
+G M M N+NF +V A ++PEFSTQ+T GM A++ TP ED+TPKSK++ PAW+T Q
Sbjct: 120 SGYMAMVNENFSSVDAPEFPEFSTQMTLGGMTAANQFTPNQEDTTPKSKKTHSPAWSTAQ 299
Query: 134 NLVLISAWIKYGTSSVVGRNQRGE 157
NLVLIS WI GTSSVVG+NQ+GE
Sbjct: 300 NLVLISRWINRGTSSVVGKNQKGE 371
>TC16018 UP|Q8MZB8 (Q8MZB8) AT15479p (CG31542-PA), partial (7%)
Length = 425
Score = 35.0 bits (79), Expect = 0.020
Identities = 24/87 (27%), Positives = 35/87 (39%), Gaps = 13/87 (14%)
Frame = +2
Query: 4 NNNHFNNQNYSNYPFNYENPNNYPNPNQFFNQR----PQNIPNFGFPP--------NFNQ 51
N+N + + +PFN N++ N R + PN+ PP +FN
Sbjct: 5 NSNPLASSSTMEFPFNQGRNNSHATHTHHHNNRRDDDEEQHPNYPPPPGSTGSPYSSFNN 184
Query: 52 SSSVPNFHP-YYGSMMKYPSQTPPFNG 77
P P +YG + P Q PPF G
Sbjct: 185PYPPPQQQPPFYGEPHRPPPQQPPFYG 265
>TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.35
{Arabidopsis thaliana;} , partial (16%)
Length = 1208
Score = 32.3 bits (72), Expect = 0.13
Identities = 18/56 (32%), Positives = 27/56 (48%)
Frame = +2
Query: 17 PFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYPSQT 72
P +Y NP++ PNPN N P + G+ P+ +QS PY S Y + +
Sbjct: 464 PLDYPNPSSNPNPNPNPNCNPSFLYYRGYTPSPSQSQPP---SPYSSSYTSYAADS 622
>TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%)
Length = 1751
Score = 31.6 bits (70), Expect = 0.22
Identities = 26/87 (29%), Positives = 30/87 (33%), Gaps = 18/87 (20%)
Frame = +2
Query: 22 NPNNYPNPNQFFNQRPQN-----IPNF---GFPPNFNQSSSVPNFHP----------YYG 63
NP YP NQ PQN PN G+PPN PN Y
Sbjct: 923 NPGGYPPNNQAGGYHPQNQAGGYPPNSQAGGYPPNSQAGGYPPNSQAGVYPPNNQGVYPP 1102
Query: 64 SMMKYPSQTPPFNGSMPMENDNFHNVG 90
+ Y +TPP G P + N G
Sbjct: 1103 NQNNYAGRTPPNVGGAPPNSAYGPNAG 1183
>AV768024
Length = 439
Score = 31.2 bits (69), Expect = 0.28
Identities = 26/105 (24%), Positives = 39/105 (36%)
Frame = +2
Query: 23 PNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYPSQTPPFNGSMPME 82
PNN ++ P N PN P+ P+ HP+ P TPP S+P
Sbjct: 38 PNNKDLSFDAKSKPPTNFPN----PSMLSQMQSPSLHPFKSLKFPIPLITPP---SLPNS 196
Query: 83 NDNFHNVGATQYPEFSTQITPSGMAIADEVTPEDSTPKSKRSKEP 127
F T TP I+++ TP + +S+ + P
Sbjct: 197 PSPFS-------LRVRTSTTPPPPLISEQQTPNSPSTQSRIAAAP 310
>TC10410 weakly similar to UP|Q8H5W8 (Q8H5W8) OJ1123_B01.10 protein, partial
(12%)
Length = 431
Score = 31.2 bits (69), Expect = 0.28
Identities = 24/94 (25%), Positives = 35/94 (36%)
Frame = +3
Query: 2 DPNNNHFNNQNYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPY 61
+PN N + P +NPN NPN P P PP FN++ P
Sbjct: 159 NPNPTPIPAPNPNPNPIQIQNPNPIHNPNPRPTPTP---PQPRPPPTFNRAPPPQQQQP- 326
Query: 62 YGSMMKYPSQTPPFNGSMPMENDNFHNVGATQYP 95
S + S PP + P +F++ + P
Sbjct: 327 --SHFSHFSSLPPSSAPSPSPASSFNSTPSPSIP 422
>AV413755
Length = 424
Score = 30.8 bits (68), Expect = 0.37
Identities = 19/65 (29%), Positives = 31/65 (47%), Gaps = 3/65 (4%)
Frame = +3
Query: 15 NYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYY---GSMMKYPSQ 71
N+PF+ YPNP P + P++ + P+++ PN P Y S PS+
Sbjct: 201 NFPFHPSGGGTYPNP----RFEPSSSPSYNYRPSYS-----PNPSPGYTNTPSPRFEPSR 353
Query: 72 TPPFN 76
+P +N
Sbjct: 354 SPSYN 368
>TC11815 weakly similar to UP|Q8MLU0 (Q8MLU0) CG13503-PA, partial (3%)
Length = 629
Score = 30.8 bits (68), Expect = 0.37
Identities = 28/103 (27%), Positives = 43/103 (41%), Gaps = 14/103 (13%)
Frame = +2
Query: 15 NYPFNYENPNNYPNPNQFFNQRPQNIPNFGFP--PNFNQSSSVPNF-------HPYYGSM 65
N+P NPN PN + Q P + + P P+ NQS+++ + P
Sbjct: 11 NFPTKNSNPNKIPNQHPTNLQPPPSPTHLQSPP*PHPNQSATLLSIIFNLQTPPPSSAFS 190
Query: 66 MKYPSQTPPFNGSMPMENDN-----FHNVGATQYPEFSTQITP 103
++ P+ TPP P++ FH+ TQ QITP
Sbjct: 191 LQAPTPTPP-----PLQCHRTLLLPFHHRTTTQIALIGIQITP 304
>TC9168 similar to GB|AAP21369.1|30102902|BT006561 At4g19200 {Arabidopsis
thaliana;}, partial (12%)
Length = 682
Score = 30.8 bits (68), Expect = 0.37
Identities = 17/43 (39%), Positives = 21/43 (48%)
Frame = +2
Query: 37 PQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYPSQTPPFNGSM 79
PQ G PP + QS S PN YG YP PP+ G++
Sbjct: 128 PQAQKPSGTPPVYGQSQS-PNTTGGYGQTGGYPPSHPPYGGAV 253
>BP033271
Length = 541
Score = 30.4 bits (67), Expect = 0.49
Identities = 18/73 (24%), Positives = 32/73 (43%)
Frame = +3
Query: 17 PFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYPSQTPPFN 76
PF NPN +P +N P+ P N ++ + +PY GS ++ Q
Sbjct: 171 PFP*ANPNXVLHPLNL-----KNFPSQNSPSNPEKNQGLHRVYPYRGSKTRHTDQNRGRR 335
Query: 77 GSMPMENDNFHNV 89
G+ +D+ ++V
Sbjct: 336 GTTSQHSDSSNHV 374
>TC14293 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding protein-like
(At5g04170), partial (18%)
Length = 540
Score = 30.4 bits (67), Expect = 0.49
Identities = 30/132 (22%), Positives = 44/132 (32%), Gaps = 18/132 (13%)
Frame = +2
Query: 6 NHFNNQNYSNYP--FNYENPNNY--------------PNPNQFFNQRPQNIPNFGFPPNF 49
+HFN Y N P + Y P Y P P Q + P + P PP+
Sbjct: 89 SHFNMSGYPNKPSGYGYGAPPPYQPYGAAPPSQSYGAPPPPQPYGAAPPSQPYGAPPPSQ 268
Query: 50 NQSSSVPNFHPYYGSMMKYPS--QTPPFNGSMPMENDNFHNVGATQYPEFSTQITPSGMA 107
+ P PY S PS P+ E+ + G + YP + +
Sbjct: 269 SYGGPPPPSQPYSASPYGQPSAPYAAPYQKPPKDESHSHGGGGGSGYPPPPSAYGSPFAS 448
Query: 108 IADEVTPEDSTP 119
+ V P + P
Sbjct: 449 LVPSVFPPGTDP 484
>TC17878 similar to GB|AAK91488.1|15215889|AY050475 AT5g02770/F9G14_80
{Arabidopsis thaliana;}, partial (17%)
Length = 465
Score = 30.4 bits (67), Expect = 0.49
Identities = 18/51 (35%), Positives = 22/51 (42%)
Frame = +3
Query: 12 NYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYY 62
N P N +PN PNPNQ Q+ PN PN NQ+ P +
Sbjct: 123 NPKKKPKNPPHPNPNPNPNQI-----QSPPNLKLNPNQNQNPPAMLLSPMF 260
>TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial (7%)
Length = 841
Score = 29.6 bits (65), Expect = 0.83
Identities = 20/68 (29%), Positives = 32/68 (46%), Gaps = 2/68 (2%)
Frame = +2
Query: 1 MDPNNNHFNNQNYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNF--NQSSSVPNF 58
+ PN N F+N N++ P NPN + N N N N NF N S++ +
Sbjct: 119 LSPNPNTFSNLNHNLSP----NPNTFSNLNH--NLSSNTFSNLNHNRNFRINIFSTLEHI 280
Query: 59 HPYYGSMM 66
P++G ++
Sbjct: 281 LPHHGLLL 304
>TC18995 weakly similar to UP|Q8PB28 (Q8PB28) Serine protease, partial (4%)
Length = 473
Score = 29.6 bits (65), Expect = 0.83
Identities = 22/89 (24%), Positives = 38/89 (41%), Gaps = 8/89 (8%)
Frame = +1
Query: 52 SSSVPNFH-----PYYGSMMKYPSQTPPFNGSMPMENDNFH---NVGATQYPEFSTQITP 103
SSS P+ H YG P +T + + + N NF +G+ + Q TP
Sbjct: 97 SSSSPSLHFREMNSNYGRSNSNPQKTASSSPYVSLNNFNFDFDLGIGSNRPKSLIDQKTP 276
Query: 104 SGMAIADEVTPEDSTPKSKRSKEPAWNTQ 132
+ + ++ + S +SK+P+W Q
Sbjct: 277 TSNSNSNPSYSYSNPNSSTQSKKPSWTHQ 363
>TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(52%)
Length = 510
Score = 29.3 bits (64), Expect = 1.1
Identities = 16/42 (38%), Positives = 19/42 (45%)
Frame = +3
Query: 19 NYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHP 60
NY P+N P + NIP F F NQ VP+F P
Sbjct: 39 NYMLPSNPPPYPANYTMIQNNIPTFQFQRFSNQLFQVPDFSP 164
>TC10235
Length = 1116
Score = 28.9 bits (63), Expect = 1.4
Identities = 23/91 (25%), Positives = 37/91 (40%), Gaps = 11/91 (12%)
Frame = +3
Query: 8 FNNQNYSNYPFNYENPNNYP---NPNQFFNQRPQ-------NIPNFGFPPNFNQSSSVP- 56
FN+ ++N+ NY +P+N NP+ FN N FP N+ + P
Sbjct: 78 FNHGRHTNH-LNYHSPSNKSQQLNPDNPFNPMGSPTTQP*INPTRSPFPQTQNRPPNPPP 254
Query: 57 NFHPYYGSMMKYPSQTPPFNGSMPMENDNFH 87
HP++ S + PP + P + H
Sbjct: 255 QTHPFHHSHLLPTFHRPPHHHKPPRQQQQHH 347
>CB826726
Length = 505
Score = 28.5 bits (62), Expect = 1.8
Identities = 18/44 (40%), Positives = 21/44 (46%), Gaps = 11/44 (25%)
Frame = +2
Query: 4 NNNHFNNQNYSN-YPFNYE----------NPNNYPNPNQFFNQR 36
NNN+ NN N +N YP + PN PNPNQ F R
Sbjct: 242 NNNNNNNVNRANRYPLHPPPPGIGYGRGGGPNPNPNPNQGFQPR 373
>AV418250
Length = 420
Score = 28.5 bits (62), Expect = 1.8
Identities = 19/75 (25%), Positives = 31/75 (41%), Gaps = 2/75 (2%)
Frame = +1
Query: 11 QNYSNYPFNYENPNNYPNPNQFFN--QRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKY 68
+ + ++ F + + P P F+ P++ P FPP F +S P P
Sbjct: 91 EEWLSFLFFVPHRSQIPQPIATFSPISEPKSNP-LPFPPTFAPKNSTPRCSPSPNPSTMM 267
Query: 69 PSQTPPFNGSMPMEN 83
S PP + S+P N
Sbjct: 268 SSGPPPLSASVPSIN 312
>AV416574
Length = 362
Score = 28.1 bits (61), Expect = 2.4
Identities = 16/47 (34%), Positives = 27/47 (57%)
Frame = +3
Query: 52 SSSVPNFHPYYGSMMKYPSQTPPFNGSMPMENDNFHNVGATQYPEFS 98
+SS PN H GS P PF+ S+P +D F ++ ++++P+ S
Sbjct: 72 ASSKPNPH---GSSRSSPWAFSPFSDSLPFSSDGFTSI-SSEHPKTS 200
>BI419840
Length = 484
Score = 28.1 bits (61), Expect = 2.4
Identities = 13/33 (39%), Positives = 20/33 (60%)
Frame = +1
Query: 263 QRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDY 295
Q + +I GSGSR +RDH G ++R + +Y
Sbjct: 139 QSQTRIRRGSGSRKGS*EERDHQGCSERHLWNY 237
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.333 0.143 0.475
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,482,710
Number of Sequences: 28460
Number of extensions: 101638
Number of successful extensions: 926
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 894
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 916
length of query: 367
length of database: 4,897,600
effective HSP length: 92
effective length of query: 275
effective length of database: 2,279,280
effective search space: 626802000
effective search space used: 626802000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149579.7


