

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
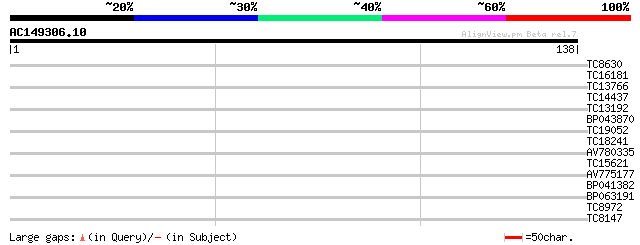
Query= AC149306.10 - phase: 0 /pseudo
(138 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8630 weakly similar to GB|AAH14051.1|15559365|BC014051 small i... 28 0.74
TC16181 similar to UP|Q8W577 (Q8W577) At1g20960/F9H16_5, partial... 27 0.97
TC13766 weakly similar to AAQ83299 (AAQ83299) QRT3, partial (9%) 26 2.2
TC14437 UP|CLPP_LOTJA (Q9BBQ9) ATP-dependent Clp protease proteo... 26 2.8
TC13192 weakly similar to UP|Q8M0B6 (Q8M0B6) Orf102, partial (22%) 26 2.8
BP043870 24 3.2
TC19052 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/M... 25 3.7
TC18241 25 4.8
AV780335 25 6.3
TC15621 similar to UP|O81340 (O81340) 26S proteasome regulatory ... 25 6.3
AV775177 25 6.3
BP041382 25 6.3
BP063191 25 6.3
TC8972 similar to UP|RK3B_ARATH (Q9LRN8) 50S ribosomal protein L... 24 8.2
TC8147 similar to UP|BCCP_SOYBN (Q42783) Biotin carboxyl carrier... 24 8.2
>TC8630 weakly similar to GB|AAH14051.1|15559365|BC014051 small inducible
cytokine subfamily E, member 1 {Homo sapiens;} , partial
(33%)
Length = 1258
Score = 27.7 bits (60), Expect = 0.74
Identities = 25/73 (34%), Positives = 32/73 (43%), Gaps = 15/73 (20%)
Frame = +1
Query: 51 C*KNC------C*IVVVEP*ISIF-WWPSGSFEICPILSSGLLSFLF-------QGSYM- 95
C KNC C +V++ *+ F WW SG + GL SFL Y+
Sbjct: 1024 CVKNCFTCSQFCLMVLIYI*VVFFFWWKSGFHNKMINMKYGLSSFLIPCFTPKVH*DYLG 1203
Query: 96 YNLFP*IFFFFLN 108
Y LF IF F+ N
Sbjct: 1204 YGLFVPIFHFYHN 1242
>TC16181 similar to UP|Q8W577 (Q8W577) At1g20960/F9H16_5, partial (26%)
Length = 1036
Score = 27.3 bits (59), Expect = 0.97
Identities = 16/37 (43%), Positives = 20/37 (53%)
Frame = -3
Query: 80 ILSSGLLSFLFQGSYMYNLFP*IFFFFLNIRGVVKFL 116
I S +LSF F S +++F IF FLNI FL
Sbjct: 563 IFSI*VLSFPFAASRNFSIFATIFISFLNINSETIFL 453
>TC13766 weakly similar to AAQ83299 (AAQ83299) QRT3, partial (9%)
Length = 543
Score = 26.2 bits (56), Expect = 2.2
Identities = 15/41 (36%), Positives = 24/41 (57%)
Frame = +2
Query: 76 EICPILSSGLLSFLFQGSYMYNLFP*IFFFFLNIRGVVKFL 116
E+C ILS + + Y++F *IF+FFL++ + FL
Sbjct: 347 ELC-ILSHIFILHFSVSLFFYHIFY*IFYFFLSLSFLSLFL 466
>TC14437 UP|CLPP_LOTJA (Q9BBQ9) ATP-dependent Clp protease proteolytic
subunit (Endopeptidase Clp) , partial (58%)
Length = 1013
Score = 25.8 bits (55), Expect = 2.8
Identities = 16/42 (38%), Positives = 21/42 (49%)
Frame = -2
Query: 68 FWWPSGSFEICPILSSGLLSFLFQGSYMYNLFP*IFFFFLNI 109
++ S +F C I S + FLF S Y F IFF +NI
Sbjct: 622 YYGGSVAFLFCLICYSAIQKFLFFFSNSYKKFFFIFFHNINI 497
>TC13192 weakly similar to UP|Q8M0B6 (Q8M0B6) Orf102, partial (22%)
Length = 550
Score = 25.8 bits (55), Expect = 2.8
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = -2
Query: 67 IFWWPSGSFEICPILSSGLLSFLFQGS 93
+FWWP S C SSGL + +GS
Sbjct: 132 LFWWPWSSISFC---SSGLANSSNKGS 61
>BP043870
Length = 562
Score = 23.9 bits (50), Expect(2) = 3.2
Identities = 8/27 (29%), Positives = 15/27 (54%)
Frame = -1
Query: 8 TLWYMGCQTVVFSVEL*NRYYSVCLFG 34
T+W++GC +V + + S C+ G
Sbjct: 493 TMWHLGCSCLVNKFQALEKRSSACVAG 413
Score = 20.0 bits (40), Expect(2) = 3.2
Identities = 7/11 (63%), Positives = 7/11 (63%)
Frame = -3
Query: 46 FLETCC*KNCC 56
FL CC NCC
Sbjct: 359 FLFCCCN*NCC 327
>TC19052 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/MPE11_6),
partial (14%)
Length = 507
Score = 25.4 bits (54), Expect = 3.7
Identities = 11/21 (52%), Positives = 16/21 (75%)
Frame = +3
Query: 16 TVVFSVEL*NRYYSVCLFGVA 36
T++F EL NR YS+C+ G+A
Sbjct: 57 TLIFQSEL-NRIYSLCISGIA 116
>TC18241
Length = 574
Score = 25.0 bits (53), Expect = 4.8
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = +3
Query: 66 SIFWWPSGSFEICPILSSGLLSFLFQGSYMY 96
+I W S +F C I +S S LF+ Y+Y
Sbjct: 249 TIATWES*TFSFCLIQASSSESILFKLKYLY 341
>AV780335
Length = 398
Score = 24.6 bits (52), Expect = 6.3
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -2
Query: 89 LFQGSYMYNLFP*IFFFFLN 108
LFQG+ N+F FF FLN
Sbjct: 298 LFQGNLFINMFIIFFFQFLN 239
>TC15621 similar to UP|O81340 (O81340) 26S proteasome regulatory subunit
S5A, partial (80%)
Length = 1109
Score = 24.6 bits (52), Expect = 6.3
Identities = 11/27 (40%), Positives = 16/27 (58%)
Frame = -2
Query: 72 SGSFEICPILSSGLLSFLFQGSYMYNL 98
+G+F C IL +G SFL G + +L
Sbjct: 949 TGTFSHCCILGAGFTSFLLVGRLLCSL 869
>AV775177
Length = 271
Score = 24.6 bits (52), Expect = 6.3
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = -3
Query: 63 P*ISIFWWPSGSFEICPILSSG 84
P ISI WP G F + +LS G
Sbjct: 68 PIISILGWPIGLFTVSVLLSFG 3
>BP041382
Length = 493
Score = 24.6 bits (52), Expect = 6.3
Identities = 15/37 (40%), Positives = 16/37 (42%)
Frame = +3
Query: 75 FEICPILSSGLLSFLFQGSYMYNLFP*IFFFFLNIRG 111
F I S G S FQ + LFP FFFF G
Sbjct: 222 FSIPRHFSPGSRSRPFQHPQRHQLFPFFFFFFFFYNG 332
>BP063191
Length = 438
Score = 24.6 bits (52), Expect = 6.3
Identities = 8/14 (57%), Positives = 9/14 (64%)
Frame = +3
Query: 27 YYSVCLFGVAYWWR 40
YY V L+G WWR
Sbjct: 231 YYQVMLYGETPWWR 272
>TC8972 similar to UP|RK3B_ARATH (Q9LRN8) 50S ribosomal protein L3-2,
chloroplast precursor, partial (31%)
Length = 643
Score = 24.3 bits (51), Expect = 8.2
Identities = 8/16 (50%), Positives = 13/16 (81%)
Frame = +1
Query: 102 IFFFFLNIRGVVKFLR 117
+ FF+LN+ G ++FLR
Sbjct: 397 VLFFYLNMDGSIRFLR 444
>TC8147 similar to UP|BCCP_SOYBN (Q42783) Biotin carboxyl carrier protein
of acetyl-CoA carboxylase, chloroplast precursor (BCCP)
, partial (42%)
Length = 731
Score = 24.3 bits (51), Expect = 8.2
Identities = 18/54 (33%), Positives = 27/54 (49%)
Frame = +3
Query: 85 LLSFLFQGSYMYNLFP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYLWIVVV*GL 138
L FL YM +F *IFF+F + RKL+ I I + ++++ GL
Sbjct: 471 LFFFLIFHFYMIGMF**IFFWF-------QLSRKLMCVIFISCLLFFLLIFEGL 611
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.368 0.170 0.704
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,120,069
Number of Sequences: 28460
Number of extensions: 47898
Number of successful extensions: 772
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 766
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 772
length of query: 138
length of database: 4,897,600
effective HSP length: 81
effective length of query: 57
effective length of database: 2,592,340
effective search space: 147763380
effective search space used: 147763380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 50 (23.9 bits)
Medicago: description of AC149306.10


