

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
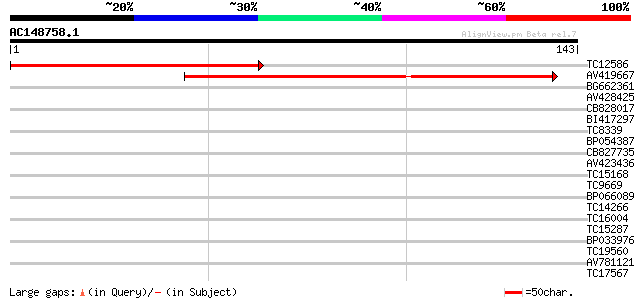
Query= AC148758.1 - phase: 0 /pseudo
(143 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC12586 weakly similar to PIR|C86410|C86410 protein F3M18.18 [im... 100 2e-22
AV419667 97 1e-21
BG662361 36 0.002
AV428425 32 0.042
CB828017 31 0.093
BI417297 28 0.60
TC8339 homologue to UP|Q84L89 (Q84L89) L-asparaginase, complete 28 0.60
BP054387 26 2.3
CB827735 26 3.0
AV423436 26 3.0
TC15168 similar to UP|O04084 (O04084) Serine carboxypeptidase is... 26 3.0
TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%) 25 3.9
BP066089 25 3.9
TC14266 homologue to PIR|C86216|C86216 protein T23G18.6 [importe... 25 3.9
TC16004 25 5.1
TC15287 similar to UP|Q9FMY5 (Q9FMY5) U2 snRNP auxiliary factor,... 25 6.7
BP033976 25 6.7
TC19560 similar to AAQ56838 (AAQ56838) At1g16900, partial (22%) 25 6.7
AV781121 24 8.7
TC17567 similar to UP|Q8RXD3 (Q8RXD3) ABI3-interacting protein 2... 24 8.7
>TC12586 weakly similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(17%)
Length = 622
Score = 99.8 bits (247), Expect = 2e-22
Identities = 46/64 (71%), Positives = 53/64 (81%)
Frame = +1
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV GE+ LG RK Q L+FS LGPKLY+NTTPVDVG +PV
Sbjct: 430 IEELHQFLEFQLPRQWAPVFGELALGPERKSQNTASLQFSFLGPKLYVNTTPVDVGKKPV 609
Query: 61 TGLR 64
TGLR
Sbjct: 610 TGLR 621
>AV419667
Length = 406
Score = 96.7 bits (239), Expect = 1e-21
Identities = 51/94 (54%), Positives = 66/94 (69%)
Frame = +1
Query: 45 KLYINTTPVDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSY 104
+L +N VDVG +PVTGLRL LEG ++N LAIHLQHL+SLPK+ L D N + S
Sbjct: 46 RLTLNYLQVDVGKKPVTGLRLYLEGKKNNCLAIHLQHLSSLPKTFQLMDEPNGNVP-HSS 222
Query: 105 SCTFHKKVKRNCFSYVCTAPVESDDSLSIVTGAQ 138
+++KV+ FS+VCTAPVESD S+VTGA+
Sbjct: 223 ERKYYEKVQWKSFSHVCTAPVESDGENSVVTGAR 324
>BG662361
Length = 330
Score = 36.2 bits (82), Expect = 0.002
Identities = 27/76 (35%), Positives = 36/76 (46%), Gaps = 9/76 (11%)
Frame = +1
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILG-PKLYINTT-------- 51
+EDL FL+FQ+ R WAP Q N + + L P + ++ T
Sbjct: 139 IEDLQYFLDFQITRVWAP------------EQNNLQTKGACLSIPSVQLDGT*AFCSVPD 282
Query: 52 PVDVGNRPVTGLRLQL 67
V VG +PVTGLRL L
Sbjct: 283 QVTVGRKPVTGLRLSL 330
>AV428425
Length = 422
Score = 32.0 bits (71), Expect = 0.042
Identities = 17/42 (40%), Positives = 24/42 (56%)
Frame = +3
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSIL 42
+EDL FL+FQ+ R WAP + RK V L+FS++
Sbjct: 309 IEDLQYFLDFQITRVWAPEQNNLQ----RKEPVCPSLQFSLM 422
>CB828017
Length = 494
Score = 30.8 bits (68), Expect = 0.093
Identities = 18/44 (40%), Positives = 22/44 (49%), Gaps = 3/44 (6%)
Frame = +2
Query: 50 TTP---VDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLP 90
TTP V V RP+ GL + + R RL H H PK+LP
Sbjct: 137 TTPP*WVPVEPRPIMGLHVAIPSHRHRRLHHHRSHPPPHPKTLP 268
>BI417297
Length = 612
Score = 28.1 bits (61), Expect = 0.60
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = -2
Query: 41 ILGPKLYINTTPVDVGNRPVTGLRLQLEGSRSN 73
++GP L + TTPV GN P + L S S+
Sbjct: 203 VVGPPLIVRTTPVRRGNSPNNPVALSSSSSSSH 105
>TC8339 homologue to UP|Q84L89 (Q84L89) L-asparaginase, complete
Length = 1467
Score = 28.1 bits (61), Expect = 0.60
Identities = 19/52 (36%), Positives = 27/52 (51%)
Frame = +1
Query: 19 VLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPVTGLRLQLEGS 70
VL +I L S RK Q N+ L SIL L + T+P + + G Q++ S
Sbjct: 115 VLTQISLSSVRKKQSNSSLAVSILASLLSLPTSPPSMSSNLW*GNWKQIQSS 270
>BP054387
Length = 553
Score = 26.2 bits (56), Expect = 2.3
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -1
Query: 96 NAYLSCDSYSCTFHKKVKRNC 116
+A + CDS+ C FH +V C
Sbjct: 289 SAVVLCDSHGCAFHPEVPFKC 227
>CB827735
Length = 531
Score = 25.8 bits (55), Expect = 3.0
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = +3
Query: 75 LAIHLQHLASLPKSLPLADNANAYLSCDSY 104
LA+ L H+ + KSL A A +L CDSY
Sbjct: 438 LAVILVHV*FIYKSLHTA*GAVPFLDCDSY 527
>AV423436
Length = 459
Score = 25.8 bits (55), Expect = 3.0
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = +3
Query: 29 RKHQVNTWLRFSILGPKLY 47
R H+++ W R + GPKLY
Sbjct: 210 RPHRISNWARKASWGPKLY 266
>TC15168 similar to UP|O04084 (O04084) Serine carboxypeptidase isolog,
30227-33069, partial (55%)
Length = 1005
Score = 25.8 bits (55), Expect = 3.0
Identities = 13/47 (27%), Positives = 21/47 (44%)
Frame = +2
Query: 63 LRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFH 109
+ ++L G N +HLQH L P+++ A S +S H
Sbjct: 713 MTMRLVGGLKNIRGLHLQHFEELVMQFPVSNPATHLHSSPPFSLENH 853
>TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%)
Length = 472
Score = 25.4 bits (54), Expect = 3.9
Identities = 16/48 (33%), Positives = 22/48 (45%), Gaps = 1/48 (2%)
Frame = +2
Query: 5 HRFLEFQLPRQWAPVLGEIHLGSYR-KHQVNTWLRFSILGPKLYINTT 51
HRFL F + Q + + L + KH TW R+ +L P TT
Sbjct: 209 HRFL*FSMLTQESSSISSSWLWRWPWKHSTLTWRRWRLLQPTFITITT 352
>BP066089
Length = 426
Score = 25.4 bits (54), Expect = 3.9
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = -1
Query: 104 YSCTFHKKVKRNCFSYV 120
YSC +K RNCFS++
Sbjct: 321 YSCGLWRK*HRNCFSFI 271
>TC14266 homologue to PIR|C86216|C86216 protein T23G18.6 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, complete
Length = 1607
Score = 25.4 bits (54), Expect = 3.9
Identities = 29/123 (23%), Positives = 45/123 (36%), Gaps = 19/123 (15%)
Frame = +1
Query: 18 PVLGEIHLGSYR-KH--QVNTWLRFSILGPKLYINTTPVDVGNRP--------VTGLRLQ 66
P G IH KH ++ ++ S L L TP D RP + L +
Sbjct: 217 PWAGRIHFHRLNIKHDSRLEGLIKMSDLTINLAAICTPADYNTRPLDTIYSNFIDALPVA 396
Query: 67 LEGSRSNRLAIHLQHL--------ASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFS 118
S +N+ IH + LPK PL + Y+ + S +++ +S
Sbjct: 397 KYCSENNKRLIHFSTCEVYGKTIGSFLPKDSPLRQDPAYYVLKEDESPCIFGSIEKQRWS 576
Query: 119 YVC 121
Y C
Sbjct: 577 YAC 585
>TC16004
Length = 594
Score = 25.0 bits (53), Expect = 5.1
Identities = 23/70 (32%), Positives = 33/70 (46%)
Frame = -2
Query: 49 NTTPVDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTF 108
N P + G PVT Q+EGSR RLA S P+ ++ + ++Y C
Sbjct: 314 N*NPTNKG--PVT---TQVEGSRGKRLACQ-----SHPE-----EDW*CISNVEAYCCYG 180
Query: 109 HKKVKRNCFS 118
H +KR+C S
Sbjct: 179 HHGIKRHCAS 150
>TC15287 similar to UP|Q9FMY5 (Q9FMY5) U2 snRNP auxiliary factor, small
subunit, partial (81%)
Length = 1263
Score = 24.6 bits (52), Expect = 6.7
Identities = 21/68 (30%), Positives = 31/68 (44%), Gaps = 18/68 (26%)
Frame = -2
Query: 78 HLQHLASL----------PKSLPLADN-ANAYL-------SCDSYSCTFHKKVKRNCFSY 119
HL H SL P SLP D+ A ++L S DS+ + +CFSY
Sbjct: 956 HLLHNQSLWYRR*K*EFTPASLPFLDSIARSWLAVLRSLPSPDSHHASSFPHHPYHCFSY 777
Query: 120 VCTAPVES 127
C++ +E+
Sbjct: 776 HCSSSLEN 753
>BP033976
Length = 551
Score = 24.6 bits (52), Expect = 6.7
Identities = 24/81 (29%), Positives = 38/81 (46%), Gaps = 2/81 (2%)
Frame = +3
Query: 49 NTTPVDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSC-- 106
NTT ++ ++P ++L++ S N I + +P SL + + SYS
Sbjct: 57 NTTGQNMNHKP-RRIKLKVPISLQNHKTI----VNIVPLSLH*LSDGPPH*LIKSYSPLK 221
Query: 107 TFHKKVKRNCFSYVCTAPVES 127
KKV+ CFS CT V+S
Sbjct: 222 AITKKVRSTCFSLSCTNHVKS 284
>TC19560 similar to AAQ56838 (AAQ56838) At1g16900, partial (22%)
Length = 516
Score = 24.6 bits (52), Expect = 6.7
Identities = 10/21 (47%), Positives = 12/21 (56%)
Frame = +1
Query: 103 SYSCTFHKKVKRNCFSYVCTA 123
S C+F K K+NCF C A
Sbjct: 454 SCGCSFKKVWKKNCFICTCYA 516
>AV781121
Length = 404
Score = 24.3 bits (51), Expect = 8.7
Identities = 18/55 (32%), Positives = 24/55 (42%), Gaps = 4/55 (7%)
Frame = +1
Query: 8 LEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILG----PKLYINTTPVDVGNR 58
L F L R A V G I +YR N W R P +++++ V VG R
Sbjct: 43 LSFTLQRNRAVVKGSIKQLTYR*SISNIWERIQGCSNPSYPDTFLSSSSVRVGKR 207
>TC17567 similar to UP|Q8RXD3 (Q8RXD3) ABI3-interacting protein 2, partial
(25%)
Length = 511
Score = 24.3 bits (51), Expect = 8.7
Identities = 11/27 (40%), Positives = 18/27 (65%), Gaps = 1/27 (3%)
Frame = -1
Query: 73 NRLAIHLQHLAS-LPKSLPLADNANAY 98
+ L +HLQHL+S LP P+ N++ +
Sbjct: 226 HHLELHLQHLSSLLPLLFPVPSNSHRH 146
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,841,549
Number of Sequences: 28460
Number of extensions: 42000
Number of successful extensions: 240
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 238
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 238
length of query: 143
length of database: 4,897,600
effective HSP length: 82
effective length of query: 61
effective length of database: 2,563,880
effective search space: 156396680
effective search space used: 156396680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148758.1


