

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
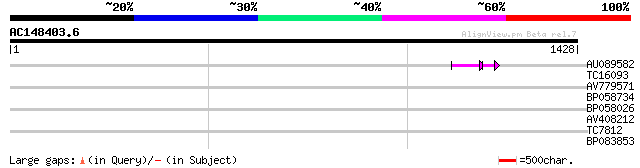
Query= AC148403.6 - phase: 0 /pseudo
(1428 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AU089582 56 9e-10
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 32 0.70
AV779571 31 1.6
BP058734 30 2.7
BP058026 30 2.7
AV408212 29 4.6
TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic me... 29 4.6
BP083853 29 6.0
>AU089582
Length = 383
Score = 56.2 bits (134), Expect(2) = 9e-10
Identities = 32/83 (38%), Positives = 44/83 (52%)
Frame = +1
Query: 1112 CCPVGRDLYQGDCEVAWCSFEHCVR*RSKIYF*ILEKFARGFGFEVEVEFGVSSADRWSV 1171
C + DL DC +AWC+ + R RS I+ LE F+ FG ++ E+ SS++ WSV
Sbjct: 10 CVSIC*DLLG*DCFLAWCTCVYNFRSRSSIHITFLEVFSNCFGNSIKNEYRFSSSN*WSV 189
Query: 1172 GEDNSVTRGFVENLCS*ARRNLG 1194
ED S RG+ LC R G
Sbjct: 190 REDYSDLRGYASCLCVXPERXAG 258
Score = 25.0 bits (53), Expect(2) = 9e-10
Identities = 18/43 (41%), Positives = 23/43 (52%)
Frame = +2
Query: 1190 RRNLG*SSSVDRVHIQ**LSF*YWNGTFRGFIWSEMQNSVVLV 1232
R LG D + I **LS *+ +GT +W EMQ + LV
Sbjct: 245 RGXLGSVFVFDGIRI***LSV*HLDGTI*SLVW*EMQVTYRLV 373
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 32.0 bits (71), Expect = 0.70
Identities = 16/38 (42%), Positives = 25/38 (65%)
Frame = -1
Query: 313 LLLLILVLHIVLLLSIVFLLWVLIFLI*MERWLSKLQL 350
+L L+L+LH++L L L W+L+ L+ + WL KL L
Sbjct: 234 MLFLLLLLHLLLWL----LYWLLLLLLLLLNWLLKLML 133
>AV779571
Length = 538
Score = 30.8 bits (68), Expect = 1.6
Identities = 20/54 (37%), Positives = 37/54 (68%)
Frame = -2
Query: 304 LAILIILL*LLLLILVLHIVLLLSIVFLLWVLIFLI*MERWLSKLQLRVR*LLL 357
+A L+I+ LLL +L+L ++LLL ++ +L + I + M L K++LR++ LL+
Sbjct: 333 IANLLIIYHLLLSLLLLLLLLLLIVIMILSLTIE*VLML*VLLKMRLRLQQLLI 172
>BP058734
Length = 453
Score = 30.0 bits (66), Expect = 2.7
Identities = 23/47 (48%), Positives = 32/47 (67%), Gaps = 4/47 (8%)
Frame = +3
Query: 304 LAILIILL*LLLLILVLHIV-LLLSIVFLL---WVLIFLI*MERWLS 346
L +L+I LLLL+L+L +V LLLS + LL +LIFL+ + WLS
Sbjct: 225 LCVLLIHRRLLLLLLLLMMVLLLLSFLGLLGGFLMLIFLVILWLWLS 365
>BP058026
Length = 389
Score = 30.0 bits (66), Expect = 2.7
Identities = 13/30 (43%), Positives = 26/30 (86%)
Frame = -2
Query: 303 VLAILIILL*LLLLILVLHIVLLLSIVFLL 332
+ ++L++LL LLLL+L+L ++LLL ++F++
Sbjct: 91 IKSLLLLLLLLLLLLLLLLLLLLLLLLFMI 2
Score = 29.6 bits (65), Expect = 3.5
Identities = 14/27 (51%), Positives = 23/27 (84%)
Frame = -2
Query: 313 LLLLILVLHIVLLLSIVFLLWVLIFLI 339
LLLL+L+L ++LLL ++ LL +L+F+I
Sbjct: 82 LLLLLLLLLLLLLLLLLLLLLLLLFMI 2
>AV408212
Length = 362
Score = 29.3 bits (64), Expect = 4.6
Identities = 23/56 (41%), Positives = 33/56 (58%)
Frame = -1
Query: 302 EVLAILIILL*LLLLILVLHIVLLLSIVFLLWVLIFLI*MERWLSKLQLRVR*LLL 357
E+ +I +LL*L LL+L L ++LLL ++ LL +L+ M L L V LLL
Sbjct: 197 EMCSIPTLLL*LWLLLL*LLLLLLLLLLLLLLLLLLESCMPLQQDLLHLHVLLLLL 30
>TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic membrane
protein 1, complete
Length = 1149
Score = 29.3 bits (64), Expect = 4.6
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +1
Query: 307 LIILL*LLLLILVLHIVLLLSIVFLLWVLIFLI*MERWL 345
+I L*+ L ++ H L+L +F W L L *M WL
Sbjct: 862 IIEFL*IFPLCIMFHSTLVLYFIFKNWELFLLC*MNWWL 978
>BP083853
Length = 455
Score = 28.9 bits (63), Expect = 6.0
Identities = 20/62 (32%), Positives = 33/62 (52%), Gaps = 2/62 (3%)
Frame = +3
Query: 445 LRIKQ*LIGYRWCVIFLKFFQMRFLMCLQREKLSFLLILSLE--RSRSRWHLIVCLLLSY 502
L + + + +C F+ +F F+M LQ+ + L LE S+ + HL+VCLLL
Sbjct: 213 LEVTEVAVSVAFC--FVCYFLSSFVMPLQQTLMHIWLHQGLEWIHSQLQEHLLVCLLLHA 386
Query: 503 LS 504
+S
Sbjct: 387 MS 392
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.368 0.165 0.613
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,508,658
Number of Sequences: 28460
Number of extensions: 382074
Number of successful extensions: 5639
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2977
Number of HSP's successfully gapped in prelim test: 207
Number of HSP's that attempted gapping in prelim test: 2310
Number of HSP's gapped (non-prelim): 3556
length of query: 1428
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1326
effective length of database: 1,994,680
effective search space: 2644945680
effective search space used: 2644945680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148403.6


