

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147429.6 - phase: 0 /pseudo
(974 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
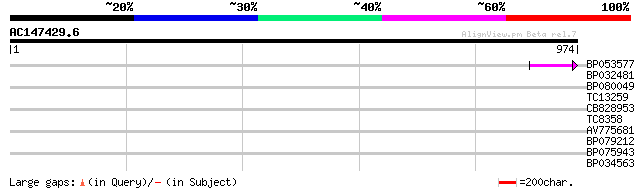
Score E
Sequences producing significant alignments: (bits) Value
BP053577 47 1e-05
BP032481 40 0.002
BP080049 38 0.007
TC13259 37 0.020
CB828953 33 0.17
TC8358 similar to UP|Q84T14 (Q84T14) Vacuolar ATPase subunit E (... 33 0.17
AV775681 29 3.1
BP079212 28 7.0
BP075943 28 7.0
BP034563 28 9.1
>BP053577
Length = 463
Score = 47.4 bits (111), Expect = 1e-05
Identities = 29/82 (35%), Positives = 41/82 (49%)
Frame = +3
Query: 893 LLKDYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDG 952
L D GFR++ E D L + + RSYL +S+ L FD SF+ R G
Sbjct: 135 LATDLGFRRITLETDCLQLFNTWR-SQEEGRSYLSSIISDCRILISAFDYVSVSFVRRTG 311
Query: 953 NHTAHSLAQLAHSEPNLVWIEE 974
N A LA+ + + N+VW+EE
Sbjct: 312 NTVADFLARNSDTYANMVWVEE 377
>BP032481
Length = 447
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/55 (40%), Positives = 29/55 (52%)
Frame = -1
Query: 920 DVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSLAQLAHSEPNLVWIEE 974
+V+ SYL L + L FD+ SF+ R GN A A+ A + NLVW EE
Sbjct: 234 EVE*SYLSSILQDCRLLFTAFDVVSLSFVRRTGNTVAGFFARNAETYANLVWPEE 70
>BP080049
Length = 356
Score = 38.1 bits (87), Expect = 0.007
Identities = 26/82 (31%), Positives = 37/82 (44%)
Frame = -1
Query: 893 LLKDYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDG 952
L + GFR + FE D L + K S L + + L FD+ SF+ R
Sbjct: 332 LASELGFR*IVFETDCLHLYEAWKRRGGC--SVLDTLILDCRNLLSYFDVFQLSFVRRTS 159
Query: 953 NHTAHSLAQLAHSEPNLVWIEE 974
N +LA+L N+VW+EE
Sbjct: 158 NCVVDALAKLCFQLGNVVWVEE 93
>TC13259
Length = 506
Score = 36.6 bits (83), Expect = 0.020
Identities = 24/79 (30%), Positives = 41/79 (51%)
Frame = +1
Query: 896 DYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHT 955
+ GF +V+ E DN+ ++ K G SYL ++++ ++ F +++PR +
Sbjct: 151 ELGFFEVEVEIDNQVVVNSWK-GTKKHVSYLANVIADVVRISQCFRFFSLAYVPRLSHLA 327
Query: 956 AHSLAQLAHSEPNLVWIEE 974
A LAQ A S+ V IEE
Sbjct: 328 ADYLAQFALSDYCFVGIEE 384
>CB828953
Length = 554
Score = 33.5 bits (75), Expect = 0.17
Identities = 14/33 (42%), Positives = 23/33 (69%)
Frame = -2
Query: 930 LSEIHQLQLNFDICHFSFIPRDGNHTAHSLAQL 962
+++I L+ +F C FS+IPR+ N AH +A+L
Sbjct: 337 VADILNLRRHFQWCGFSWIPRNSNKVAHEIAKL 239
>TC8358 similar to UP|Q84T14 (Q84T14) Vacuolar ATPase subunit E (Fragment),
complete
Length = 1321
Score = 33.5 bits (75), Expect = 0.17
Identities = 19/68 (27%), Positives = 34/68 (49%)
Frame = +3
Query: 907 DNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSLAQLAHSE 966
+N L + V +++ + + + L F H S+I ++GN A + +LA+S
Sbjct: 1035 NNNILNISVYVALEIIVVWTACLIKDCKALASLFISFHLSYICQNGNKAADFM*RLAYSH 1214
Query: 967 PNLVWIEE 974
N VWIE+
Sbjct: 1215 NNSVWIED 1238
>AV775681
Length = 466
Score = 29.3 bits (64), Expect = 3.1
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = +3
Query: 925 YLGLFLSEIHQLQLNFDICHFSFIP 949
Y+GLF+ H+L N I +FS IP
Sbjct: 84 YMGLFILSGHELNTNISIIYFSTIP 158
>BP079212
Length = 397
Score = 28.1 bits (61), Expect = 7.0
Identities = 16/37 (43%), Positives = 22/37 (59%)
Frame = +3
Query: 253 FYCSNSFFLWVSPTISLKLLCIVSKLLLFMLSLMVNL 289
FYC ++ ++ I L LC+ S LL MLS M+NL
Sbjct: 120 FYCIPRAWILLTSDIELAFLCLHSS-LLSMLSPMINL 227
>BP075943
Length = 547
Score = 28.1 bits (61), Expect = 7.0
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Frame = -3
Query: 917 VGIDVDRSYLGLFLSEI-HQLQLNFDICHFSFIPRDGNHTAHS 958
+G+++ RS G+ L++ + LQL D HF F PR +H +S
Sbjct: 401 LGLEIARSTSGIVLNQRKYALQLISDSGHFGFQPRFYSHGTNS 273
>BP034563
Length = 501
Score = 27.7 bits (60), Expect = 9.1
Identities = 12/39 (30%), Positives = 20/39 (50%)
Frame = +2
Query: 253 FYCSNSFFLWVSPTISLKLLCIVSKLLLFMLSLMVNLLR 291
F+C +SF LW+S + S C + + L L + +R
Sbjct: 272 FFCCSSFILWISSSFSSNFSC---NIRIISLDLSTSFVR 379
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.363 0.165 0.602
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,224,559
Number of Sequences: 28460
Number of extensions: 250918
Number of successful extensions: 2629
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1934
Number of HSP's successfully gapped in prelim test: 63
Number of HSP's that attempted gapping in prelim test: 658
Number of HSP's gapped (non-prelim): 2049
length of query: 974
length of database: 4,897,600
effective HSP length: 99
effective length of query: 875
effective length of database: 2,080,060
effective search space: 1820052500
effective search space used: 1820052500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147429.6


