

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
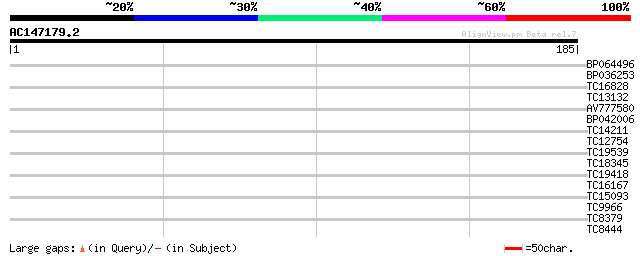
Query= AC147179.2 + phase: 0
(185 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP064496 31 0.15
BP036253 28 0.95
TC16828 27 1.6
TC13132 similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement... 27 2.1
AV777580 26 3.6
BP042006 26 3.6
TC14211 26 4.7
TC12754 similar to UP|Q9FHM5 (Q9FHM5) Similarity to DNA-binding ... 26 4.7
TC19539 25 6.2
TC18345 similar to UP|Q84VU1 (Q84VU1) 65 kDa microtubule associa... 25 6.2
TC19418 weakly similar to UP|Q9FZ10 (Q9FZ10) Serine threonine ki... 25 6.2
TC16167 similar to UP|Q9FGH3 (Q9FGH3) Dihydroflavonol 4-reductas... 25 8.1
TC15093 25 8.1
TC9966 25 8.1
TC8379 similar to UP|Q9SNJ3 (Q9SNJ3) ESTs AU031435(E61570), part... 25 8.1
TC8444 homologue to UP|RUBB_PEA (P08927) RuBisCO subunit binding... 25 8.1
>BP064496
Length = 526
Score = 30.8 bits (68), Expect = 0.15
Identities = 14/45 (31%), Positives = 26/45 (57%)
Frame = -1
Query: 19 IHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTI 63
I+ +F+FC+YVLL C+ L L ++ F ++ L++F +
Sbjct: 154 IYPVFMFCSYVLLLGPQCCLILALCSSVL-----FHVLFLYLFPV 35
>BP036253
Length = 501
Score = 28.1 bits (61), Expect = 0.95
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = -3
Query: 133 LIFCLEWVVLTLAFFLKYYACV 154
L FCL +V LA+ L YY+CV
Sbjct: 238 LFFCLACIVKILAWELHYYSCV 173
>TC16828
Length = 557
Score = 27.3 bits (59), Expect = 1.6
Identities = 11/34 (32%), Positives = 18/34 (52%)
Frame = -1
Query: 49 SVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAH 82
S+ F+ +L+H+F S C + GA R +H
Sbjct: 449 SLLSFYFVLIHLFPTHHMSSNCQSCGAERQSYSH 348
>TC13132 similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement pol
polyprotein, partial (3%)
Length = 591
Score = 26.9 bits (58), Expect = 2.1
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = +3
Query: 37 CIFLTLSLRLVPSVCGFFLILL 58
CI TL L + P+ C FFL+LL
Sbjct: 342 CITETLGLWMRPASCNFFLLLL 407
>AV777580
Length = 494
Score = 26.2 bits (56), Expect = 3.6
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = +3
Query: 35 SSCIFLTLSLRLVPSVCGFFLILLH 59
S C F L L L+ ++CG F+ILL+
Sbjct: 171 SKCKFPFLKLILLITICGTFVILLY 245
>BP042006
Length = 469
Score = 26.2 bits (56), Expect = 3.6
Identities = 11/31 (35%), Positives = 19/31 (60%), Gaps = 1/31 (3%)
Frame = +1
Query: 22 LFLFCN-YVLLGAASSCIFLTLSLRLVPSVC 51
LF+FCN ++L A +FL L ++ ++C
Sbjct: 271 LFIFCNCFLLAQTAQQAVFLYFFLLILTAIC 363
>TC14211
Length = 488
Score = 25.8 bits (55), Expect = 4.7
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = +3
Query: 42 LSLRLVPSVCGFFLILLH 59
LS +PS+ GFFL+++H
Sbjct: 93 LSCSFLPSISGFFLLVIH 146
>TC12754 similar to UP|Q9FHM5 (Q9FHM5) Similarity to DNA-binding protein,
partial (24%)
Length = 932
Score = 25.8 bits (55), Expect = 4.7
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +2
Query: 19 IHKLFLFCNYVLLGAASSCIFLTLSLRLVPS 49
I KLF+F + VL+ A + IFL +L+ S
Sbjct: 44 ISKLFVFSSMVLISAC*AAIFLETKAQLLQS 136
>TC19539
Length = 565
Score = 25.4 bits (54), Expect = 6.2
Identities = 14/42 (33%), Positives = 21/42 (49%)
Frame = +2
Query: 16 HYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLIL 57
HY + + L C ++ + A+ IF T+ V C FLIL
Sbjct: 404 HYFVVLM*LLCLFICIFLAAFIIFRTIMQCTVVFFCMLFLIL 529
>TC18345 similar to UP|Q84VU1 (Q84VU1) 65 kDa microtubule associated
protein, partial (8%)
Length = 521
Score = 25.4 bits (54), Expect = 6.2
Identities = 9/21 (42%), Positives = 16/21 (75%)
Frame = +2
Query: 51 CGFFLILLHIFTIAGAVSGCA 71
CG+FL+L++I I ++GC+
Sbjct: 287 CGYFLVLIYISVI*KWLAGCS 349
>TC19418 weakly similar to UP|Q9FZ10 (Q9FZ10) Serine threonine kinase
homolog COK-4, partial (11%)
Length = 523
Score = 25.4 bits (54), Expect = 6.2
Identities = 21/72 (29%), Positives = 35/72 (48%), Gaps = 5/72 (6%)
Frame = -1
Query: 25 FCNYVLL-GAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAH- 82
FC Y + SC+ +SL + S C F+ I LH++ + C+++ YSA
Sbjct: 505 FCLYHQVPNLKKSCMIFKISLFSLSSPCFFYCI*LHVWM---HNNNCSSLFRQNEYSAED 335
Query: 83 ---MVATVLTAI 91
M+ ++L AI
Sbjct: 334 FNFMLYSLLLAI 299
>TC16167 similar to UP|Q9FGH3 (Q9FGH3) Dihydroflavonol 4-reductase-like
(Cinnamoyl-CoA reductase-like protein), partial (44%)
Length = 535
Score = 25.0 bits (53), Expect = 8.1
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +2
Query: 53 FFLILLHIFTIAGAVSGCAAV 73
F + LLH T+ AV+GCA V
Sbjct: 263 FQIDLLHYDTVLAAVNGCAGV 325
>TC15093
Length = 611
Score = 25.0 bits (53), Expect = 8.1
Identities = 14/44 (31%), Positives = 22/44 (49%)
Frame = +2
Query: 128 GGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQ 171
G + ILI+ ++L + FF AC+ N C +V+ S Q
Sbjct: 395 GYVIILIYGFICLILQIYFFRNLAACIIFCNACIHIVMYSTLCQ 526
>TC9966
Length = 862
Score = 25.0 bits (53), Expect = 8.1
Identities = 10/27 (37%), Positives = 13/27 (48%)
Frame = +2
Query: 60 IFTIAGAVSGCAAVGANRWYSAHMVAT 86
+ TI C G NRW+ AH+ T
Sbjct: 305 LLTIKMP*PSCLHFGENRWFCAHLCVT 385
>TC8379 similar to UP|Q9SNJ3 (Q9SNJ3) ESTs AU031435(E61570), partial (5%)
Length = 809
Score = 25.0 bits (53), Expect = 8.1
Identities = 18/40 (45%), Positives = 25/40 (62%)
Frame = +2
Query: 22 LFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIF 61
LF+F LL AASSC+FL SL S+ F++L+ I+
Sbjct: 656 LFIF----LL*AASSCVFLCKSLH-Y*SISFCFVMLIWIY 760
>TC8444 homologue to UP|RUBB_PEA (P08927) RuBisCO subunit binding-protein
beta subunit, chloroplast precursor (60 kDa chaperonin
beta subunit) (CPN-60 beta), partial (53%)
Length = 949
Score = 25.0 bits (53), Expect = 8.1
Identities = 15/49 (30%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Frame = +3
Query: 89 TAIFQGSVSVLVFTRTSDFLGEL----QSYVREEDGSVILKLSGGLAIL 133
T + GS V R + ++ Q Y +E+ I KLSGG+A++
Sbjct: 603 TIVGDGSTQEAVNKRVAQIKNQIEAAEQDYEKEKLSERIAKLSGGVAVI 749
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.142 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,985,288
Number of Sequences: 28460
Number of extensions: 60733
Number of successful extensions: 488
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 488
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 488
length of query: 185
length of database: 4,897,600
effective HSP length: 85
effective length of query: 100
effective length of database: 2,478,500
effective search space: 247850000
effective search space used: 247850000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147179.2


