

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146746.4 - phase: 0
(472 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
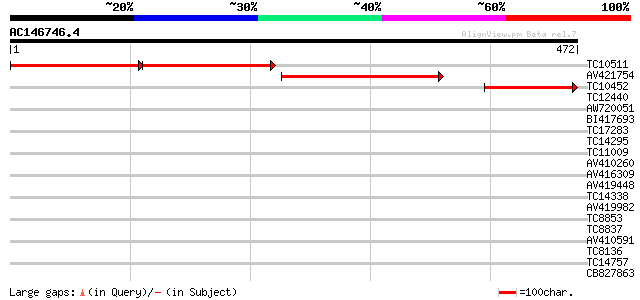
Score E
Sequences producing significant alignments: (bits) Value
TC10511 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associate... 213 3e-91
AV421754 231 1e-61
TC10452 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associate... 137 3e-33
TC12440 36 0.016
AW720051 34 0.059
BI417693 34 0.059
TC17283 similar to UP|O65760 (O65760) Extensin (Fragment), parti... 34 0.059
TC14295 similar to UP|O03990 (O03990) RAD23, isoform I, partial ... 33 0.077
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 33 0.13
AV410260 33 0.13
AV416309 32 0.17
AV419448 32 0.17
TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%) 32 0.17
AV419982 32 0.17
TC8853 similar to PIR|T09217|T09217 protein sam2B - spinach {Spi... 32 0.17
TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein ki... 32 0.22
AV410591 32 0.22
TC8136 similar to GB|AAB60729.1|2160166|F21M12 F21M12.13 {Arabid... 32 0.29
TC14757 similar to UP|O04361 (O04361) Prohibitin, complete 32 0.29
CB827863 32 0.29
>TC10511 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associated protein,
partial (41%)
Length = 782
Score = 213 bits (542), Expect(2) = 3e-91
Identities = 98/111 (88%), Positives = 103/111 (92%)
Frame = +2
Query: 111 KASSLTEGRRSDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKV 170
K+S++TEGRRSDFFNHLKAA DSL+ALAWIAFTGKDCGMSMPIAHVEESWQM+EFY NKV
Sbjct: 449 KSSAMTEGRRSDFFNHLKAAADSLTALAWIAFTGKDCGMSMPIAHVEESWQMSEFYCNKV 628
Query: 171 LVEYRNKDPNHVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSK 221
LVEYRNKDPNHVEW KALKELYLPGLRDYVK FHPLGPVWS TG APSK
Sbjct: 629 LVEYRNKDPNHVEWAKALKELYLPGLRDYVKGFHPLGPVWSPTGSTIAPSK 781
Score = 139 bits (350), Expect(2) = 3e-91
Identities = 72/111 (64%), Positives = 89/111 (79%)
Frame = +1
Query: 1 MDETLIKRLESAVTRLEALSTGIHHSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAAD 60
M+E LI RLESAV RLEALS G D +DAA+DPS++AF DL+ QYV+R+S AA+
Sbjct: 118 MEEKLIGRLESAVARLEALSVGFQGGGGGGDVADAATDPSIIAFDDLMGQYVARVSAAAE 297
Query: 61 IIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMK 111
IGG VL+VT V+EAF+VQK+LLIK+KQ+QKP+ AGLAEFLKPLN+VI K
Sbjct: 298 KIGGPVLEVTKVVQEAFAVQKQLLIKVKQSQKPNNAGLAEFLKPLNDVITK 450
Score = 26.9 bits (58), Expect = 7.2
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = -1
Query: 222 VNASAAPAAPSAPPPPPASLFSSESSQASS 251
++ S A +A S PPPPP + S S +A++
Sbjct: 239 MDGSVAASATSPPPPPPWNPTDSASKRATA 150
>AV421754
Length = 416
Score = 231 bits (590), Expect = 1e-61
Identities = 114/135 (84%), Positives = 123/135 (90%)
Frame = +2
Query: 227 APAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKNR 286
AP AP+ PPPPPASLFSSESSQ SSSKPKVGMSAVF+EI TGNVT+GLRKV+DDMKTKNR
Sbjct: 11 APTAPAVPPPPPASLFSSESSQTSSSKPKVGMSAVFREISTGNVTSGLRKVSDDMKTKNR 190
Query: 287 ADRSGVVGNSVKESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVY 346
DR+GVVG+S KES+A RAFSK GPPK ELQMGRKWVVENQI +K LVIEDCDSKQSVY
Sbjct: 191 TDRTGVVGSSEKESRAGSRAFSKTGPPKFELQMGRKWVVENQIGKKDLVIEDCDSKQSVY 370
Query: 347 VYGCKNSVLQIQGKV 361
VYGCK+SVLQIQGKV
Sbjct: 371 VYGCKDSVLQIQGKV 415
>TC10452 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associated protein,
partial (15%)
Length = 516
Score = 137 bits (346), Expect = 3e-33
Identities = 67/77 (87%), Positives = 70/77 (90%)
Frame = +2
Query: 396 QGSAPTISVDNTSGCQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIH 455
QGSAPTISVDNTSGCQ+YLS+DSLETSI TAKSSEINVLVP E DGD VEHSLPQ YIH
Sbjct: 2 QGSAPTISVDNTSGCQLYLSQDSLETSIFTAKSSEINVLVPGAEPDGDLVEHSLPQPYIH 181
Query: 456 LFKEGRFETTPASHSGG 472
FK+GRFETTPASHSGG
Sbjct: 182 AFKDGRFETTPASHSGG 232
>TC12440
Length = 629
Score = 35.8 bits (81), Expect = 0.016
Identities = 21/38 (55%), Positives = 27/38 (70%), Gaps = 2/38 (5%)
Frame = +3
Query: 219 PSKVNASA-APAAPSAPPPPPASLFSSESSQA-SSSKP 254
PS +N+ A AP A SAPPPPP+SL SS + ++SKP
Sbjct: 105 PSFLNSHATAPLAQSAPPPPPSSLSSSSPWRCPTASKP 218
>AW720051
Length = 564
Score = 33.9 bits (76), Expect = 0.059
Identities = 27/77 (35%), Positives = 34/77 (44%), Gaps = 2/77 (2%)
Frame = +1
Query: 222 VNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRK-VTDD 280
+N + P+AP PPPP S SS + S K M A Q G G V T
Sbjct: 304 INKNTVPSAPPPPPPPEPQSSSGGSSTLTESTSK--MFAADQSGGGGGVFNEAESFTTSS 477
Query: 281 MKTKNRA-DRSGVVGNS 296
M + N A D++ GNS
Sbjct: 478 MSSTNEAIDQAFGFGNS 528
>BI417693
Length = 500
Score = 33.9 bits (76), Expect = 0.059
Identities = 19/39 (48%), Positives = 22/39 (55%), Gaps = 1/39 (2%)
Frame = -1
Query: 217 FAPSKV-NASAAPAAPSAPPPPPASLFSSESSQASSSKP 254
F P V N+S +P+APS PPP FSS SS S P
Sbjct: 380 FPPQFVANSSLSPSAPSEPPPAAEDSFSSPSSSPSC*NP 264
>TC17283 similar to UP|O65760 (O65760) Extensin (Fragment), partial (16%)
Length = 513
Score = 33.9 bits (76), Expect = 0.059
Identities = 15/31 (48%), Positives = 19/31 (60%)
Frame = +1
Query: 219 PSKVNASAAPAAPSAPPPPPASLFSSESSQA 249
P + +A PA PS PPPPP + S E+ QA
Sbjct: 52 PRRGSAPPLPAIPSPPPPPPTTKHSPETLQA 144
>TC14295 similar to UP|O03990 (O03990) RAD23, isoform I, partial (29%)
Length = 622
Score = 33.5 bits (75), Expect = 0.077
Identities = 18/43 (41%), Positives = 22/43 (50%)
Frame = +3
Query: 208 PVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQAS 250
P + TG AP A AAPAA AP P PA + S + + S
Sbjct: 465 PTPTATGAPLAPVTAAAPAAPAATPAPSPAPAPISSGTAVEGS 593
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 32.7 bits (73), Expect = 0.13
Identities = 18/53 (33%), Positives = 27/53 (49%)
Frame = -2
Query: 208 PVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSA 260
P S + + S ++S++ +P P PP SS SS +SSS P G S+
Sbjct: 237 PKSSSSSSSSSSSSSSSSSSSGSPPPPRPPSTPPSSSSSSSSSSSSPPPGASS 79
Score = 28.5 bits (62), Expect = 2.5
Identities = 16/38 (42%), Positives = 24/38 (63%)
Frame = -2
Query: 223 NASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSA 260
++S++ ++ S+ PPP AS SS SS SSS P S+
Sbjct: 135 SSSSSSSSSSSSPPPGAS--SSSSSSPSSSSPSSSSSS 28
Score = 28.5 bits (62), Expect = 2.5
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = -2
Query: 219 PSKVNASAAPAAPSAPPPPPASLFSSESSQASSS 252
PS +S++ ++ S+ PPP + SS SS +SSS
Sbjct: 150 PSTPPSSSSSSSSSSSSPPPGASSSSSSSPSSSS 49
>AV410260
Length = 373
Score = 32.7 bits (73), Expect = 0.13
Identities = 20/44 (45%), Positives = 26/44 (58%), Gaps = 2/44 (4%)
Frame = +2
Query: 220 SKVNASAAP--AAPSAPPPPPASLFSSESSQASSSKPKVGMSAV 261
SK +SA P ++PSAP P P S +S SS AS+ P SA+
Sbjct: 221 SKTPSSATPTSSSPSAPSPTPTSRSASFSSAASAGTPPPTASAL 352
>AV416309
Length = 277
Score = 32.3 bits (72), Expect = 0.17
Identities = 16/64 (25%), Positives = 29/64 (45%), Gaps = 4/64 (6%)
Frame = +2
Query: 204 HPLGPVWSQTGK----VFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMS 259
HP P + G+ + AP + + PPPPP + S+ +S A+ + P ++
Sbjct: 26 HPFAPSATHMGRETQLILAPPTTTTTTTVESQPPPPPPPLAAPSASASAAAVAAPPTSLA 205
Query: 260 AVFQ 263
F+
Sbjct: 206 PGFR 217
>AV419448
Length = 382
Score = 32.3 bits (72), Expect = 0.17
Identities = 14/37 (37%), Positives = 23/37 (61%)
Frame = +2
Query: 218 APSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKP 254
+PS ++ +A P+ PPPP + S+ SS A++S P
Sbjct: 239 SPSPISNPSASLPPTPPPPPTKTPSSTSSSPAATSSP 349
Score = 28.9 bits (63), Expect = 1.9
Identities = 18/42 (42%), Positives = 22/42 (51%), Gaps = 5/42 (11%)
Frame = +2
Query: 218 APSKVNASAAP-----AAPSAPPPPPASLFSSESSQASSSKP 254
A S +ASA P A PSAP PP ++ S S+ SS P
Sbjct: 119 ASSWTSASAPPPSSPTATPSAPTPPLSAALSLASTATSSPSP 244
>TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%)
Length = 1751
Score = 32.3 bits (72), Expect = 0.17
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Frame = +3
Query: 218 APSKVNASAAPA-APSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRK 276
+P +S PA +P APPPPP+++ + KP M+A TG+++ K
Sbjct: 183 SPPAATSSPPPAPSPPAPPPPPSTILTLTGPTVPRRKPFCSMAAT---SSTGSLSWRSPK 353
Query: 277 VT 278
VT
Sbjct: 354 VT 359
>AV419982
Length = 415
Score = 32.3 bits (72), Expect = 0.17
Identities = 16/44 (36%), Positives = 24/44 (54%), Gaps = 6/44 (13%)
Frame = +1
Query: 218 APSKVNASAAPAAPSA------PPPPPASLFSSESSQASSSKPK 255
APS ++ P+ P + PPPPP++ SS SS S+ P+
Sbjct: 247 APSTFTSTTTPSIPLSLSPPPPPPPPPSASTSSPSSSGKSTAPR 378
>TC8853 similar to PIR|T09217|T09217 protein sam2B - spinach {Spinacia
oleracea;}, partial (43%)
Length = 1157
Score = 32.3 bits (72), Expect = 0.17
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = -3
Query: 215 KVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKP 254
K+F P++ + P AP AP PP +L S SS S P
Sbjct: 507 KLFGPNQTSTEPWPFAPPAP*PPSQALSSLSSSSLLQSPP 388
>TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(15%)
Length = 606
Score = 32.0 bits (71), Expect = 0.22
Identities = 17/41 (41%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Frame = -3
Query: 218 APSKVNASAAPAAPSAPPPPPASL----FSSESSQASSSKP 254
AP ++ ++P +PS+PP PPAS S SS ++SS P
Sbjct: 313 APPQIPPPSSPPSPSSPPSPPASAP*TPSSP*SSNSASSPP 191
Score = 27.7 bits (60), Expect = 4.2
Identities = 14/35 (40%), Positives = 18/35 (51%)
Frame = -3
Query: 225 SAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMS 259
SA+ P A PPPP S +S A+SS + S
Sbjct: 208 SASSPPPQASPPPPTPTVSR*TSTATSSSSQTNSS 104
>AV410591
Length = 429
Score = 32.0 bits (71), Expect = 0.22
Identities = 18/33 (54%), Positives = 22/33 (66%), Gaps = 2/33 (6%)
Frame = +2
Query: 224 ASAAPAAPSAPPPPPASL--FSSESSQASSSKP 254
ASA P +P A PPPP+SL S S+ +S SKP
Sbjct: 128 ASAKP*SPPASPPPPSSLPPPPSNSTTSSPSKP 226
Score = 27.3 bits (59), Expect = 5.5
Identities = 20/59 (33%), Positives = 25/59 (41%), Gaps = 4/59 (6%)
Frame = +2
Query: 206 LGPVWSQTGKVFAPSKVNASAAPAAP----SAPPPPPASLFSSESSQASSSKPKVGMSA 260
L P S + P+ + PA+P S PPPP S SS S AS P S+
Sbjct: 86 LYPTLSTSFGAPTPASAKP*SPPASPPPPSSLPPPPSNSTTSSPSKPASLPTPTPASSS 262
>TC8136 similar to GB|AAB60729.1|2160166|F21M12 F21M12.13 {Arabidopsis
thaliana;}, partial (70%)
Length = 1659
Score = 31.6 bits (70), Expect = 0.29
Identities = 16/28 (57%), Positives = 21/28 (74%)
Frame = +2
Query: 210 WSQTGKVFAPSKVNASAAPAAPSAPPPP 237
WS T AP++++ S APAAP+APPPP
Sbjct: 404 WSSTP---APTRLS-SRAPAAPAAPPPP 475
>TC14757 similar to UP|O04361 (O04361) Prohibitin, complete
Length = 1141
Score = 31.6 bits (70), Expect = 0.29
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 3/58 (5%)
Frame = +1
Query: 218 APSKVNASAAPAAPSAPPPPP---ASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTA 272
+P+ +ASA P PS PP P AS SS + A+SS + G+ N T+
Sbjct: 76 SPAPPSASALPPPPSTPPSTPSMEASAPSSSTDSAASSTTPSAKEPISSSHGSRNPTS 249
Score = 31.2 bits (69), Expect = 0.38
Identities = 21/52 (40%), Positives = 26/52 (49%), Gaps = 2/52 (3%)
Frame = +1
Query: 210 WSQTGKVFAPSKVNASAAPAAPSAPPPP--PASLFSSESSQASSSKPKVGMS 259
W T K +PS + A P+A + PPPP P S S E+S SSS S
Sbjct: 37 WVAT-KPQSPSSQTSPAPPSASALPPPPSTPPSTPSMEASAPSSSTDSAASS 189
>CB827863
Length = 571
Score = 31.6 bits (70), Expect = 0.29
Identities = 16/40 (40%), Positives = 21/40 (52%), Gaps = 2/40 (5%)
Frame = -1
Query: 210 WSQTGKVFAPS--KVNASAAPAAPSAPPPPPASLFSSESS 247
W G PS K+ + A P +APPPPP ++ S SS
Sbjct: 322 WKALGFTADPSGLKLKSLADPLEAAAPPPPPDTIISKSSS 203
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.128 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,563,259
Number of Sequences: 28460
Number of extensions: 105479
Number of successful extensions: 2600
Number of sequences better than 10.0: 223
Number of HSP's better than 10.0 without gapping: 1730
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2275
length of query: 472
length of database: 4,897,600
effective HSP length: 94
effective length of query: 378
effective length of database: 2,222,360
effective search space: 840052080
effective search space used: 840052080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146746.4


