

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
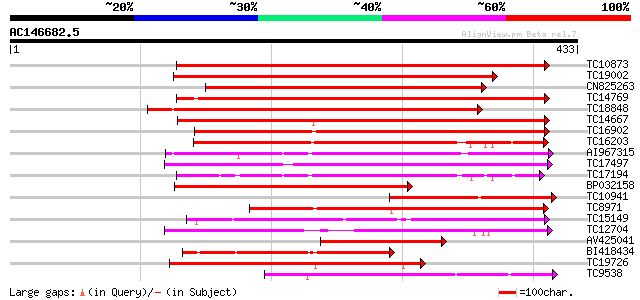
Query= AC146682.5 - phase: 0
(433 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 266 6e-72
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 241 2e-64
CN825263 235 1e-62
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 234 3e-62
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 232 8e-62
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 227 3e-60
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 213 4e-56
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 211 1e-55
AI967315 206 5e-54
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 194 2e-50
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 193 5e-50
BP032158 191 3e-49
TC10941 homologue to UP|Q7Y229 (Q7Y229) At1g56720, partial (27%) 185 1e-47
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 179 8e-46
TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like pro... 174 2e-44
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 174 3e-44
AV425041 171 3e-43
BI418434 167 4e-42
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 164 3e-41
TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein k... 157 2e-39
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 266 bits (679), Expect = 6e-72
Identities = 140/287 (48%), Positives = 188/287 (64%), Gaps = 2/287 (0%)
Frame = +1
Query: 128 YSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEA 187
+ +E+ ATR F+E N+IGEGG+G VY+G L G VAVK L ++ Q +EF +EV
Sbjct: 415 FGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVMEVLM 594
Query: 188 IGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGT 247
+ + H NLVRL+GYC +G +R+LVYEY+ G+LE L PL W RMK+A+G
Sbjct: 595 LSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGA 774
Query: 248 AKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKL-LGSEKTHVTTRVMGTF 306
A+GL YLH +P V++RD+KS+NILLD +N K+SDFGLAKL + THV+TRVMGT+
Sbjct: 775 ARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVSTRVMGTY 954
Query: 307 GYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRS 366
GY +PEYA +G L +SD+YSFGV+L+E++TGR ID SR PGE NLV W + S RR
Sbjct: 955 GYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWARPYFSDRRR 1134
Query: 367 -DELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
+VDPL++ R L + + I C+ RP + IV LE
Sbjct: 1135FGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDIVVALE 1275
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 241 bits (614), Expect = 2e-64
Identities = 118/247 (47%), Positives = 163/247 (65%)
Frame = +1
Query: 126 RWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEV 185
R +SLKE+ AT F N +GEGG+G VY G L DG +AVK L +A+ EF VEV
Sbjct: 22 RVFSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEV 201
Query: 186 EAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAI 245
E + +VRHKNL+ L GYCAEG R++VY+Y+ N +L LHG L W+ RM IAI
Sbjct: 202 EILARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAI 381
Query: 246 GTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGT 305
G+A+G+ YLH P ++HRDIK+SN+LLD ++ A+V+DFG AKL+ THVTTRV GT
Sbjct: 382 GSAEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVTTRVKGT 561
Query: 306 FGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRR 365
GY++PEYA G NE DV+SFG+LL+E+ +G+ P++ + ++ DW + +++
Sbjct: 562 LGYLAPEYAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDWALPLACAKK 741
Query: 366 SDELVDP 372
E DP
Sbjct: 742 FTEFADP 762
>CN825263
Length = 663
Score = 235 bits (599), Expect = 1e-62
Identities = 113/216 (52%), Positives = 155/216 (71%), Gaps = 1/216 (0%)
Frame = +1
Query: 150 GYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARR 209
G+G+VY+G+L DG VAVK L + + +EF EVE + ++ H+NLV+L+G C E R
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 210 MLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKS 269
L+YE V NG++E LHG T PL W+ RMKIA+G A+GL YLHE P V+HRD KS
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 270 SNILLDKNWNAKVSDFGLAK-LLGSEKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSF 328
SNILL+ ++ KVSDFGLA+ L H++T VMGTFGY++PEYA TG L +SDVYS+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSY 540
Query: 329 GVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSR 364
GV+L+E++TG P+D S+PPG+ NLV W + +++S+
Sbjct: 541 GVVLLELLTGTKPVDLSQPPGQENLVTWARPILTSK 648
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 234 bits (596), Expect = 3e-62
Identities = 124/286 (43%), Positives = 181/286 (62%), Gaps = 1/286 (0%)
Frame = +3
Query: 128 YSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEA 187
++L+ +E+AT ++ +IGEGG+G VYRG L DG VAVK + Q +EF E+
Sbjct: 2133 FTLEYIEVATERYK--TLIGEGGFGSVYRGTLNDGQEVAVKVRSSTSTQGTREFDNELNL 2306
Query: 188 IGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGT 247
+ ++H+NLV L+GYC E +++LVY ++ NG+L+ L+G L W R+ IA+G
Sbjct: 2307 LSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPTRLSIALGA 2486
Query: 248 AKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSE-KTHVTTRVMGTF 306
A+GL YLH V+HRDIKSSNILLD + AKV+DFG +K E ++V+ V GT
Sbjct: 2487 ARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLEVRGTA 2666
Query: 307 GYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRS 366
GY+ PEY T L+E+SDV+SFGV+L+EI++GR P++ RP E +LV+W + +
Sbjct: 2667 GYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWSLVEWATPYIRGSKV 2846
Query: 367 DELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
DE+VDP I+ A+ RV+ + L+C++ RP M IV LE
Sbjct: 2847 DEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSMVAIVRELE 2984
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 232 bits (592), Expect = 8e-62
Identities = 116/256 (45%), Positives = 164/256 (63%)
Frame = +2
Query: 106 GVRKGDGGGHMMEDPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVV 165
G K + G N W R ++ KE+ AT GF + N +GEGG+G VY G DG +
Sbjct: 203 GSEKVEEGPTSFGSVNNSW-RIFTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQI 379
Query: 166 AVKNLHNNKGQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWL 225
AVK L +AE EF VEVE +G+VRHKNL+ L GYC +R++VY+Y+ N +L L
Sbjct: 380 AVKKLKAMNSKAEMEFAVEVEVLGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHL 559
Query: 226 HGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDF 285
HG L W RMKIAIG+A+G+ YLH + P ++HRDIK+SN+LL+ ++ V+DF
Sbjct: 560 HGQFAVEVQLNWQKRMKIAIGSAEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADF 739
Query: 286 GLAKLLGSEKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYS 345
G AKL+ +H+TTRV GT GY++PEYA G ++E DVYSFG+LL+E++TGR PI+
Sbjct: 740 GFAKLIPEGVSHMTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLELVTGRKPIEKL 919
Query: 346 RPPGEMNLVDWFKAMV 361
+ + +W + ++
Sbjct: 920 PGGVKRTITEWAEPLI 967
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 227 bits (578), Expect = 3e-60
Identities = 115/291 (39%), Positives = 177/291 (60%), Gaps = 7/291 (2%)
Frame = +2
Query: 129 SLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLH-NNKGQAEKEFKVEVEA 187
SL E++ T F +IGEG YG VY L DG VAVK L +++ + EF +V
Sbjct: 326 SLDELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFLTQVSM 505
Query: 188 IGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVG-----PTSPLTWDIRMK 242
+ ++++ N V L GYC EG R+L YE+ G+L LHG G P L W R++
Sbjct: 506 VSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVR 685
Query: 243 IAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHV-TTR 301
IA+ A+GL YLHE ++P ++HRDI+SSN+L+ +++ AK++DF L+ + +TR
Sbjct: 686 IAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTR 865
Query: 302 VMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMV 361
V+GTFGY +PEYA TG L ++SDVYSFGV+L+E++TGR P+D++ P G+ +LV W +
Sbjct: 866 VLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRL 1045
Query: 362 SSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
S + + VDP ++ P+ + ++ + C+ + RP M +V L+
Sbjct: 1046SEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALQ 1198
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 213 bits (543), Expect = 4e-56
Identities = 108/272 (39%), Positives = 167/272 (60%), Gaps = 1/272 (0%)
Frame = +3
Query: 142 EGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKNLVRLVG 201
E ++G GG+G VY G + G VA+K + Q EF+ E+E + K+RH++LV L+G
Sbjct: 3 EALLLGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIG 182
Query: 202 YCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLEPK 261
YC E +LVY+++ G L + L+ P PL W R++I IG A+GL YLH G +
Sbjct: 183 YCEENTEMILVYDHMAYGTLREHLYKTQKP--PLPWKQRLEICIGAARGLHYLHTGAKYT 356
Query: 262 VVHRDIKSSNILLDKNWNAKVSDFGLAKLLGS-EKTHVTTRVMGTFGYVSPEYASTGMLN 320
++HRD+K++NILLD+ W AKVSDFGL+K + + THV+T V G+FGY+ PEY L
Sbjct: 357 IIHRDVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLT 536
Query: 321 ERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSDELVDPLIETPPSP 380
++SDVYSFGV+L EI+ R ++ S +++L +W + D+++DP ++ +P
Sbjct: 537 DKSDVYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAP 716
Query: 381 RALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
K+ ++C+ I+RP MG ++ LE
Sbjct: 717 ECFKKFAETAMKCVSDQGIERPSMGDVLWNLE 812
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 211 bits (538), Expect = 1e-55
Identities = 116/284 (40%), Positives = 178/284 (61%), Gaps = 13/284 (4%)
Frame = +2
Query: 141 EEGNVIGEGGYGVVYRGVLQDGCVVAVKNL-HNNKGQAEKEFKVEVEAIGKVRHKNLVRL 199
+E N+IG+GG G+VYRG + +G VA+K L G+ + F+ E+E +GK+RH+N++RL
Sbjct: 2195 KEENIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRL 2374
Query: 200 VGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLE 259
+GY + +L+YEY+ NG+L +WLHG G L W++R KIA+ A+GL Y+H
Sbjct: 2375 LGYVSNKDTNLLLYEYMPNGSLGEWLHGAKG--GHLRWEMRYKIAVEAARGLCYMHHDCS 2548
Query: 260 PKVVHRDIKSSNILLDKNWNAKVSDFGLAKLL-GSEKTHVTTRVMGTFGYVSPEYASTGM 318
P ++HRD+KS+NILLD ++ A V+DFGLAK L + + + G++GY++PEYA T
Sbjct: 2549 PLIIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLK 2728
Query: 319 LNERSDVYSFGVLLMEIITGRSPIDYSRPPGE----MNLVDWFKAMVS--SRRSD----- 367
++E+SDVYSFGV+L+E+I GR P+ GE +++V W +S S+ SD
Sbjct: 2729 VDEKSDVYSFGVVLLELIIGRKPV------GEFGDGVDIVGWVNKTMSELSQPSDTALVL 2890
Query: 368 ELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHML 411
+VDP + P + + I + C+ RP M ++VHML
Sbjct: 2891 AVVDPRLSGYPLTSVI-HMFNIAMMCVKEMGPARPTMREVVHML 3019
>AI967315
Length = 1308
Score = 206 bits (525), Expect = 5e-54
Identities = 115/299 (38%), Positives = 178/299 (59%), Gaps = 3/299 (1%)
Frame = +1
Query: 120 PNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNN--KGQA 177
P W + +S +E+ AT GF N++G+GGY VY+G L+ G +AVK L +
Sbjct: 91 PRPSW-KCFSYEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERK 267
Query: 178 EKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTW 237
EKEF E+ IG V H N++ L+G C + LV+E G++ +H +PL W
Sbjct: 268 EKEFLTEIGTIGHVCHSNVMPLLGCCIDNGL-YLVFELSTVGSVASLIHDE--KMAPLDW 438
Query: 238 DIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTH 297
R KI +GTA+GL YLH+G + +++HRDIK+SNILL +++ ++SDFGLAK L S+ TH
Sbjct: 439 KTRYKIVLGTARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTH 618
Query: 298 VT-TRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDW 356
+ + GTFG+++PEY G+++E++DV++FGV L+E+I+GR P+D S +L W
Sbjct: 619 HSIAPIEGTFGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPVDGS----HQSLHTW 786
Query: 357 FKAMVSSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLESDD 415
K ++S ++LVDP +E RV CI RP M +++ ++E +
Sbjct: 787 AKPILSKWEIEKLVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVLEVMEEGE 963
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 194 bits (493), Expect = 2e-50
Identities = 109/297 (36%), Positives = 168/297 (55%), Gaps = 1/297 (0%)
Frame = +1
Query: 119 DPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAE 178
D +I + + AT F N +GEGG+G VY+G+L +G +AVK L N GQ
Sbjct: 1609 DEDIDLATIFDFSTISSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGM 1788
Query: 179 KEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWD 238
+EFK E++ I +++H+NLV+L G + N + + + + + W+
Sbjct: 1789 EEFKNEIKLIARLQHRNLVKLFGCSVHQDEN-------SHANKKMKILLDSTRSKLVDWN 1947
Query: 239 IRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKL-LGSEKTH 297
R++I G A+GL YLH+ +++HRD+K+SNILLD N K+SDFGLA++ +G +
Sbjct: 1948 KRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEA 2127
Query: 298 VTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWF 357
T RVMGT+GY+ PEYA G + +SDV+SFGV+++EII+G+ + P +NL+
Sbjct: 2128 RTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSHA 2307
Query: 358 KAMVSSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLESD 414
+ R ELVD L++ P P + R + + L C+ RP M IV ML +
Sbjct: 2308 WRLWIEERPLELVDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSIVLMLNGE 2478
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 193 bits (490), Expect = 5e-50
Identities = 120/293 (40%), Positives = 175/293 (58%), Gaps = 12/293 (4%)
Frame = +1
Query: 128 YSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEA 187
+S +E+ AT F N IG+GG+G VY L+ G A+K + QA EF E++
Sbjct: 1051 FSYQELAKATNNFSLDNKIGQGGFGAVYYAELR-GKKTAIKKMDV---QASTEFLCELKV 1218
Query: 188 IGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGT 247
+ V H NLVRL+GYC EG+ LVYE+++NGNL Q+LHG+ PL W R++IA+
Sbjct: 1219 LTHVHHLNLVRLIGYCVEGSL-FLVYEHIDNGNLGQYLHGS--GKEPLPWSSRVQIALDA 1389
Query: 248 AKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGTFG 307
A+GL Y+HE P +HRD+KS+NIL+DKN KV+DFGL KL+ + + TR++GTFG
Sbjct: 1390 ARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQTRLVGTFG 1569
Query: 308 YVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEM-----NLVDWFKAMVS 362
Y+ PEYA G ++ + DVY+FGV+L E+I+ ++ + GE+ LV F+ ++
Sbjct: 1570 YMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAV---LKTGELVAESKGLVALFEEALN 1740
Query: 363 SRRSD------ELVDP-LIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIV 408
+SD +LVDP L E P LK + + C + + RP M +V
Sbjct: 1741 --KSDPCDALRKLVDPRLGENYPIDSVLK-IAQLGRACTRDNPLLRPSMRSLV 1890
>BP032158
Length = 555
Score = 191 bits (484), Expect = 3e-49
Identities = 93/181 (51%), Positives = 125/181 (68%)
Frame = +2
Query: 127 WYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVE 186
++SL++++ AT F+ N IGEGG+G VY+GVL +G V+AVK L + Q +EF E+
Sbjct: 11 YFSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFINEIG 190
Query: 187 AIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIG 246
I ++H NLV+L G C EG + +LVYEY+EN +L + L GN L W RMKI +G
Sbjct: 191 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVG 370
Query: 247 TAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGTF 306
AKGL YLHE K+VHRDIK++N+LLDK+ AK+SDFGLAKL E TH++TR+ GT
Sbjct: 371 IAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHISTRIAGTI 550
Query: 307 G 307
G
Sbjct: 551 G 553
>TC10941 homologue to UP|Q7Y229 (Q7Y229) At1g56720, partial (27%)
Length = 831
Score = 185 bits (469), Expect = 1e-47
Identities = 90/127 (70%), Positives = 106/127 (82%)
Frame = +2
Query: 291 LGSEKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGE 350
LG+ K+HVTTRVMGTFGYV+PEYA+TG+LNE+SDVYSFGVLL+E ITGR P+DY RP E
Sbjct: 2 LGAGKSHVTTRVMGTFGYVAPEYANTGLLNEKSDVYSFGVLLLEGITGRDPVDYGRPTNE 181
Query: 351 MNLVDWFKAMVSSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHM 410
+NLVDW K MV SRRS+E+VDP IE PS RALKR LL LRC+D D KRPKM Q+V M
Sbjct: 182 VNLVDWLK-MVGSRRSEEVVDPNIEVKPSTRALKRALLTALRCVDPDSEKRPKMSQVVRM 358
Query: 411 LESDDFP 417
LES+++P
Sbjct: 359 LESEEYP 379
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 179 bits (454), Expect = 8e-46
Identities = 96/230 (41%), Positives = 142/230 (61%), Gaps = 2/230 (0%)
Frame = +3
Query: 184 EVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKI 243
EVE + K+ H+NLV+L+G+ +G R+L+ EYV NG L + L G G L ++ R++I
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKI--LDFNQRLEI 179
Query: 244 AIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKL--LGSEKTHVTTR 301
AI A GLTYLH E +++HRD+KSSNILL ++ AKV+DFG A+L + ++TH++T+
Sbjct: 180 AIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTK 359
Query: 302 VMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMV 361
V GT GY+ PEY T L +SDVYSFG+LL+EI+TGR P++ + E + W
Sbjct: 360 VKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY 539
Query: 362 SSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHML 411
+ EL+DPL+E + L ++L + +C RP M + L
Sbjct: 540 NEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 689
>TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like protein kinase
protein (AT3g17840/MEB5_6), partial (45%)
Length = 1489
Score = 174 bits (442), Expect = 2e-44
Identities = 106/286 (37%), Positives = 165/286 (57%), Gaps = 9/286 (3%)
Frame = +3
Query: 136 ATRGFE-------EGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAI 188
+ RGF+ V+G+G +G Y+ VL+ G VVAVK L + +EKEFK ++E +
Sbjct: 243 SARGFDLEDLLRASAEVLGKGTFGTAYKAVLETGLVVAVKRL-KDVTISEKEFKDKIETV 419
Query: 189 GKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGP-TSPLTWDIRMKIAIGT 247
G + H++LV L Y ++LVY+Y+ G+L LHGN G +PL W+IR IA+G
Sbjct: 420 GAMDHQSLVPLRAYYFSRDEKLLVYDYMPMGSLSALLHGNKGAGRTPLNWEIRSGIALGA 599
Query: 248 AKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGTFG 307
A+G+ YLH P V H +IK+SNILL K++ AKVSDFGLA L+G T RV G
Sbjct: 600 ARGIEYLH-SQGPNVSHGNIKASNILLTKSYEAKVSDFGLAHLVGPSST--PNRVA---G 761
Query: 308 YVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSD 367
Y +PE +++++DVYSFGVLL+E++TG++P ++L W +++V +
Sbjct: 762 YRAPEVTDPRKVSQKADVYSFGVLLLELLTGKAPTHALLNEEGVDLPRWVQSLVREEWTS 941
Query: 368 ELVD-PLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
E+ D L+ + ++L + + C +RP + ++ +E
Sbjct: 942 EVFDLELLRYQNVEEEMVQLLQLAVDCAAPYPDRRPSIAEVTRSIE 1079
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 174 bits (440), Expect = 3e-44
Identities = 113/321 (35%), Positives = 167/321 (51%), Gaps = 25/321 (7%)
Frame = +1
Query: 119 DPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAE 178
D +I + + T F E N +GEGG+G VY+GVL +G +AVK L N GQ
Sbjct: 1390 DEDIDLATIFDFSTISSTTNHFSESNKLGEGGFGPVYKGVLANGQEIAVKRLSNTSGQGM 1569
Query: 179 KEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWD 238
+EFK EV+ I +++H+NLV+L+G +L+YE++ N +L+ ++ +D
Sbjct: 1570 EEFKNEVKLIARLQHRNLVKLLGCSIHHDEMLLIYEFMHNRSLDYFI-----------FD 1716
Query: 239 IRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLL-GSEKTH 297
R++I +HRD+K+SNILLD N K+SDFGLA++ G +
Sbjct: 1717 SRLRI-------------------IHRDLKTSNILLDSEMNPKISDFGLARIFTGDQVEA 1839
Query: 298 VTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNL---- 353
T RVMGT+GY+SPEYA G + +SDV+SFGV+++EII+G+ + P NL
Sbjct: 1840 KTKRVMGTYGYMSPEYAVHGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHS 2019
Query: 354 ----VDWFKAM-------VSSR---------RSDELVDPLIETPPSPRALKRVLLICLRC 393
V KA+ V +R R ELVD L++ P + R + I L C
Sbjct: 2020 SNFAVFLIKALRICMFENVKNRKAWRLWIEERPLELVDELLDGLAIPTEILRYIHIALLC 2199
Query: 394 IDLDVIKRPKMGQIVHMLESD 414
+ RP M +V ML +
Sbjct: 2200 VQQRPEYRPDMLSVVLMLNGE 2262
>AV425041
Length = 288
Score = 171 bits (432), Expect = 3e-43
Identities = 81/96 (84%), Positives = 90/96 (93%)
Frame = +1
Query: 238 DIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTH 297
DIRM I +GTAKGLTYLHEGLEPKVVHRDIKSSNILL K W++KVSDFGLAKLLG E ++
Sbjct: 1 DIRMNIILGTAKGLTYLHEGLEPKVVHRDIKSSNILLSKQWHSKVSDFGLAKLLGPESSY 180
Query: 298 VTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLM 333
+TTRVMGTFGYV+PEYASTGMLNERSDVYSFG+L+M
Sbjct: 181 ITTRVMGTFGYVAPEYASTGMLNERSDVYSFGILIM 288
>BI418434
Length = 478
Score = 167 bits (422), Expect = 4e-42
Identities = 86/162 (53%), Positives = 115/162 (70%)
Frame = +2
Query: 133 VEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIGKVR 192
+++ATRGF E +G GG+G V++G L D +VAVK L G EKEF+ EV IG ++
Sbjct: 2 LQLATRGFSEK--VGHGGFGTVFQGELSDSSLVAVKRLER-PGGGEKEFRAEVSTIGNIQ 172
Query: 193 HKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLT 252
H NLVRL G+C+E + R+LVYEY+ NG L +L + GP L+WD+R ++AIGTAKG+
Sbjct: 173 HVNLVRLRGFCSENSHRLLVYEYMHNGALSAYLRKD-GPC--LSWDVRFQVAIGTAKGIA 343
Query: 253 YLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSE 294
YLHE L ++H DIK NILLD ++ AKVSDFGLAKL+G +
Sbjct: 344 YLHEELRCCILHCDIKPENILLDSDFTAKVSDFGLAKLIGRD 469
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 164 bits (415), Expect = 3e-41
Identities = 89/200 (44%), Positives = 129/200 (64%), Gaps = 5/200 (2%)
Frame = +3
Query: 123 GWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVL-QDGCVVAVKNLHNNKGQAEKEF 181
G R ++ E++ AT F+E + +G+GGYGVVYRG+L ++ VAVK +K ++ +F
Sbjct: 117 GTPREFNYVELKKATNNFDEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDDF 296
Query: 182 KVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPT--SPLTWDI 239
E+ I ++RHK+LVRL G+C + +LVY+Y+ NG+L+ + G T +PL+W +
Sbjct: 297 LSELIIINRLRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPL 476
Query: 240 RMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVT 299
R KI G A L YLH + KVVHRD+K+SNI+LD +NAK+ DFGLA+ L +EK T
Sbjct: 477 RYKIISGVASALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYT 656
Query: 300 --TRVMGTFGYVSPEYASTG 317
V GT GY++PE TG
Sbjct: 657 ELEGVHGTMGYIAPECFHTG 716
>TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein kinase 5
precursor , partial (10%)
Length = 1067
Score = 157 bits (398), Expect = 2e-39
Identities = 91/234 (38%), Positives = 138/234 (58%), Gaps = 10/234 (4%)
Frame = +2
Query: 195 NLVRLVGYCAEGARRMLVYEYVENGNLEQWLH---------GNVGPTSPLTWDIRMKIAI 245
N+VRL+ + A +LVYEY+EN +L++WLH G V + L W R+KIAI
Sbjct: 5 NIVRLLCCISNEASMLLVYEYLENHSLDKWLHLKPKSSSVSGVVQQYTVLDWPKRLKIAI 184
Query: 246 GTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAK-LLGSEKTHVTTRVMG 304
G A+GL+Y+H P +VHRD+K+SNILLDK +NAKV+DFGLA+ L+ + ++ + V+G
Sbjct: 185 GAAQGLSYMHHDCSPPIVHRDVKTSNILLDKQFNAKVADFGLARMLIKPGELNIMSTVIG 364
Query: 305 TFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSR 364
TFGY++PEY T ++E+ DVYSFGV+L+E+ TG+ +Y + W ++ S
Sbjct: 365 TFGYIAPEYVQTTRISEKVDVYSFGVVLLELTTGKE-ANYGDQHSSLAEWAWRHILIGSN 541
Query: 365 RSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLESDDFPF 418
D L ++E + V + + C RP M +++ +L S PF
Sbjct: 542 VXDLLXKDVMEASYIDE-MCSVFKLGVMCTATLPATRPSMKEVLPILLSFGEPF 700
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,884,581
Number of Sequences: 28460
Number of extensions: 91215
Number of successful extensions: 1002
Number of sequences better than 10.0: 324
Number of HSP's better than 10.0 without gapping: 815
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 818
length of query: 433
length of database: 4,897,600
effective HSP length: 93
effective length of query: 340
effective length of database: 2,250,820
effective search space: 765278800
effective search space used: 765278800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146682.5


