

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
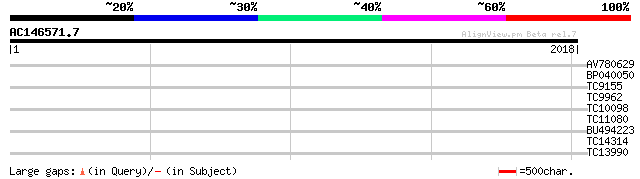
Query= AC146571.7 + phase: 0 /pseudo
(2018 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV780629 39 0.011
BP040050 33 0.44
TC9155 homologue to UP|Q949G5 (Q949G5) Mob1-like protein, complete 32 1.3
TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial... 30 2.9
TC10098 weakly similar to UP|Q9LZX8 (Q9LZX8) Guanine nucleotide ... 30 3.8
TC11080 similar to UP|Q84TQ7 (Q84TQ7) GIA/RGA-like gibberellin r... 30 4.9
BU494223 29 8.4
TC14314 similar to UP|Q9FPI4 (Q9FPI4) AT5g16110 (AT5g16110/T21H1... 29 8.4
TC13990 similar to UP|PLSB_PHAVU (Q43822) Glycerol-3-phosphate a... 29 8.4
>AV780629
Length = 512
Score = 38.5 bits (88), Expect = 0.011
Identities = 19/52 (36%), Positives = 27/52 (51%)
Frame = -1
Query: 1947 IGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKEL 1998
+ + + LE L + CQ C G L V CTS C +FY R+K QK++
Sbjct: 479 VAQVSDLEMLFGRLWTQCQECQGS---LHQDVLCTSRDCPIFYRRKKAQKDM 333
>BP040050
Length = 552
Score = 33.1 bits (74), Expect = 0.44
Identities = 19/67 (28%), Positives = 35/67 (51%)
Frame = +1
Query: 902 VLLTDKVDNQKLDKNLLCKTVGSEPIVDDLKSNHMKLTKVVIGNSSLVDKNLKSHLSLPT 961
+ LT ++ + ++K LC ++ +++N KLT + S +K ++HL P+
Sbjct: 220 IKLTKRI--KPINKTNLCTKKNKTNLI*LIQTNSPKLTTTPSNSQSTKNKTFRNHLHHPS 393
Query: 962 FPNNLHL 968
F N LHL
Sbjct: 394 FFNYLHL 414
>TC9155 homologue to UP|Q949G5 (Q949G5) Mob1-like protein, complete
Length = 1087
Score = 31.6 bits (70), Expect = 1.3
Identities = 17/55 (30%), Positives = 28/55 (50%)
Frame = -2
Query: 978 ALDVFLPISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGSHLYLLTPNI 1032
++DVFL +S S+ W KC + ++ T G + YQ + L+L PN+
Sbjct: 318 SIDVFLKLSTLASRWSTLFWTKCFLVSASKTEETHGGLSQYQ---ASLFLEDPNL 163
>TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial (21%)
Length = 585
Score = 30.4 bits (67), Expect = 2.9
Identities = 17/57 (29%), Positives = 29/57 (50%), Gaps = 3/57 (5%)
Frame = +3
Query: 1055 AEDQDVPKCTSGPLRHTPDQMRQEPGAKD---KDISKCASGPTLRPELHQDTEKKLP 1108
A Q P P++ PD R EPG+ + + + SGP+L +Q+ +++LP
Sbjct: 303 APPQPQPPSQPSPIQRAPDHPRPEPGSDAISLRPLGRTGSGPSLAVP-NQEKDRELP 470
>TC10098 weakly similar to UP|Q9LZX8 (Q9LZX8) Guanine nucleotide exchange
factor-like protein, partial (5%)
Length = 666
Score = 30.0 bits (66), Expect = 3.8
Identities = 23/65 (35%), Positives = 30/65 (45%), Gaps = 4/65 (6%)
Frame = +3
Query: 240 WSLPSESGGSSNDNTHSP----KRQSICELEGDISVDEILNQQFKMFSSLSQTSSNVNMV 295
WS+P SG SP Q+IC L GDIS ++ L+ F + SSL N V
Sbjct: 102 WSIPLGSGKRRELAARSPLVVATLQAICSL-GDISFEKNLSHFFPLLSSLVSCEHGSNDV 278
Query: 296 QSLVS 300
+S
Sbjct: 279 PVALS 293
>TC11080 similar to UP|Q84TQ7 (Q84TQ7) GIA/RGA-like gibberellin response
modulator, partial (33%)
Length = 556
Score = 29.6 bits (65), Expect = 4.9
Identities = 13/39 (33%), Positives = 21/39 (53%)
Frame = -1
Query: 533 VDGSNDDEFSSPCASLAEISSAVEINSEYKRASENHLLH 571
+DG +D SP SL EI S + + +R+ ++LH
Sbjct: 154 IDGVREDSIDSPSKSLCEIGSNLPHRAGLRRSDHPYVLH 38
>BU494223
Length = 495
Score = 28.9 bits (63), Expect = 8.4
Identities = 27/85 (31%), Positives = 37/85 (42%), Gaps = 4/85 (4%)
Frame = +2
Query: 1215 SLDGLSDCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLL 1274
S+ L DCK F K ++ +K+ L WE L LAE + GLL
Sbjct: 38 SISILPDCKADVFNTAKVRVRT-SKVQMIPVNLKLFSWETYDEDLSSLAESSRITAPGLL 214
Query: 1275 NDLSRTPSNS----WINSQDIKTSE 1295
L+ T +S +I S DI +SE
Sbjct: 215 EQLNVTRDSSDYLWYITSVDISSSE 289
>TC14314 similar to UP|Q9FPI4 (Q9FPI4) AT5g16110 (AT5g16110/T21H19_30),
partial (12%)
Length = 1145
Score = 28.9 bits (63), Expect = 8.4
Identities = 18/72 (25%), Positives = 36/72 (50%), Gaps = 2/72 (2%)
Frame = +3
Query: 915 KNLL--CKTVGSEPIVDDLKSNHMKLTKVVIGNSSLVDKNLKSHLSLPTFPNNLHLDEDD 972
KN+L C+ + EP+V K ++ + N+S +D ++ + P ++ ++ED
Sbjct: 159 KNILTSCEEMRMEPVVCP------KPRRLSLLNNSSIDNQIRPIMRPPMINSHPEIEEDS 320
Query: 973 EMPGNALDVFLP 984
+ LD+ LP
Sbjct: 321 GVRAELLDIILP 356
>TC13990 similar to UP|PLSB_PHAVU (Q43822) Glycerol-3-phosphate
acyltransferase, chloroplast precursor (GPAT) , partial
(50%)
Length = 789
Score = 28.9 bits (63), Expect = 8.4
Identities = 14/42 (33%), Positives = 25/42 (59%)
Frame = +3
Query: 721 EVRSRDNLVCRKQSFPSDSDSFLNISKDECLVQHERHYLEAG 762
+V+S+D++V + P S +FLN ++ L+ R +EAG
Sbjct: 120 KVKSKDSVVSASMAAPVSSRTFLNAQNEQELLSGIRKEVEAG 245
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,458,243
Number of Sequences: 28460
Number of extensions: 511409
Number of successful extensions: 2427
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 2396
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2426
length of query: 2018
length of database: 4,897,600
effective HSP length: 104
effective length of query: 1914
effective length of database: 1,937,760
effective search space: 3708872640
effective search space used: 3708872640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146571.7


