

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146550.5 - phase: 0
(337 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
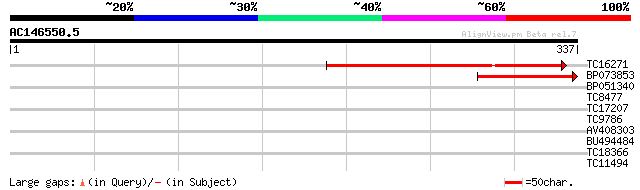
Score E
Sequences producing significant alignments: (bits) Value
TC16271 similar to GB|AAM15909.1|20257479|AF492660 purple acid p... 181 1e-46
BP073853 64 4e-11
BP051340 28 2.9
TC8477 similar to UP|R12C_ARATH (Q9SKZ3) 40S ribosomal protein S... 27 3.7
TC17207 27 6.4
TC9786 27 6.4
AV408303 27 6.4
BU494484 27 6.4
TC18366 homologue to UP|Q8W2E3 (Q8W2E3) 3-hydroxy-3-methylglutar... 26 8.3
TC11494 similar to UP|Q7X9B2 (Q7X9B2) 9/13 hydroperoxide lyase, ... 26 8.3
>TC16271 similar to GB|AAM15909.1|20257479|AF492660 purple acid phosphatase
{Arabidopsis thaliana;} , partial (32%)
Length = 653
Score = 181 bits (460), Expect = 1e-46
Identities = 86/143 (60%), Positives = 109/143 (76%)
Frame = +1
Query: 189 VLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNV 248
VLP + Y + LL++++ AL STAKWKIVV HHPI+S G HG+T+EL QLLPIL+ NNV
Sbjct: 1 VLPRDKYLSNLLRDLEIALKVSTAKWKIVVGHHPIRSIGHHGDTKELIRQLLPILEENNV 180
Query: 249 DAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQIS 308
D YINGHDHCLEHI S I F TSGGGSKAW GDI+ +K Y+DGQGF+S+++
Sbjct: 181 DMYINGHDHCLEHITSTNSQILFLTSGGGSKAWKGDIQN-KTNGVKFYYDGQGFMSLELQ 357
Query: 309 KTNASIVFYDVFGKVLHTWSMSK 331
+ NA +V+YD+FGKVLH ++SK
Sbjct: 358 QINAKVVYYDIFGKVLHLQNLSK 426
>BP073853
Length = 386
Score = 63.9 bits (154), Expect = 4e-11
Identities = 29/59 (49%), Positives = 44/59 (74%)
Frame = -3
Query: 279 KAWGGDIKPWDPKELKLYHDGQGFVSMQISKTNASIVFYDVFGKVLHTWSMSKKLKEAA 337
KAW GDI+ + + L ++DGQGF+S+Q+++T+A+I FYDV G VLH SK+LK ++
Sbjct: 384 KAWRGDIQEKNKRGLNFFYDGQGFMSVQLTQTDANIEFYDVSGNVLHRIISSKQLKHSS 208
>BP051340
Length = 537
Score = 27.7 bits (60), Expect = 2.9
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = -2
Query: 140 ILRQKDNRWVCLRSFILDTDNVEFFFV 166
I+ K RWVCL+ FI + +FFF+
Sbjct: 86 IVETK*LRWVCLQVFIYNVLTEKFFFI 6
>TC8477 similar to UP|R12C_ARATH (Q9SKZ3) 40S ribosomal protein S12-3,
partial (94%)
Length = 533
Score = 27.3 bits (59), Expect = 3.7
Identities = 16/43 (37%), Positives = 20/43 (46%)
Frame = -1
Query: 5 LNQHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQS 47
L HML + II L +L NH+ A LP F + Q S
Sbjct: 491 LGFHMLQNNIIGFMFLPKILDNHTGAASYLPSFSF*INLAQSS 363
>TC17207
Length = 751
Score = 26.6 bits (57), Expect = 6.4
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = -1
Query: 173 EEYFTDPGKHTYDWKGVLPLESYRAELLKE 202
E +F DP K+ WK PL Y LLK+
Sbjct: 445 EGHFNDPSKN---WKPTFPLSYYVMHLLKQ 365
>TC9786
Length = 1242
Score = 26.6 bits (57), Expect = 6.4
Identities = 18/38 (47%), Positives = 20/38 (52%), Gaps = 4/38 (10%)
Frame = -1
Query: 13 VIIAVSVLLCL----LSNHSSIAEKLPRFEHHLKPQQQ 46
VII+ LC+ LSNH IA P E HLKP Q
Sbjct: 375 VIISNLFFLCIEPDTLSNH*-IASSTPYVERHLKPYDQ 265
>AV408303
Length = 362
Score = 26.6 bits (57), Expect = 6.4
Identities = 15/30 (50%), Positives = 17/30 (56%), Gaps = 4/30 (13%)
Frame = +3
Query: 73 IVGKNLNIEFVVSTGD----NFYDDGLKGV 98
IVGKNL I F STGD N G++ V
Sbjct: 69 IVGKNLYIRFCCSTGDAMGMNMVSKGVQNV 158
>BU494484
Length = 386
Score = 26.6 bits (57), Expect = 6.4
Identities = 15/49 (30%), Positives = 23/49 (46%)
Frame = -2
Query: 43 PQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGKNLNIEFVVSTGDNFY 91
PQ+ ++F WG N+ LVA + I+G F+ T NF+
Sbjct: 232 PQKALISF*RTIRWGNHSNFK*KLVAFCLPIIGATKLYPFINPT*LNFF 86
>TC18366 homologue to UP|Q8W2E3 (Q8W2E3) 3-hydroxy-3-methylglutaryl coenzyme
A, partial (30%)
Length = 516
Score = 26.2 bits (56), Expect = 8.3
Identities = 16/35 (45%), Positives = 18/35 (50%), Gaps = 4/35 (11%)
Frame = +1
Query: 70 QMGIVGKNLNIEFVVSTGD----NFYDDGLKGVDD 100
Q I GKNL I F STGD N G++ V D
Sbjct: 214 QPAIAGKNLYIRFRCSTGDAMGMNIVSKGVQNVLD 318
>TC11494 similar to UP|Q7X9B2 (Q7X9B2) 9/13 hydroperoxide lyase, partial
(27%)
Length = 541
Score = 26.2 bits (56), Expect = 8.3
Identities = 10/28 (35%), Positives = 17/28 (60%)
Frame = -2
Query: 249 DAYINGHDHCLEHIIDKESGIHFFTSGG 276
D+ + G + H+ ++SG+ FF SGG
Sbjct: 336 DSRVGGDEGAGRHVGSEDSGVVFFDSGG 253
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,761,869
Number of Sequences: 28460
Number of extensions: 99008
Number of successful extensions: 433
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 432
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 432
length of query: 337
length of database: 4,897,600
effective HSP length: 91
effective length of query: 246
effective length of database: 2,307,740
effective search space: 567704040
effective search space used: 567704040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146550.5


