

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.9 - phase: 2 /pseudo
(1036 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
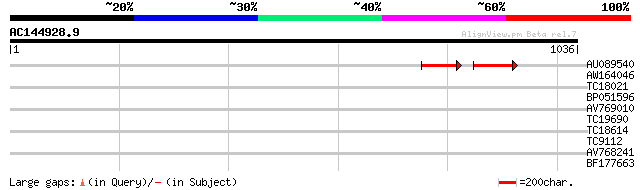
Score E
Sequences producing significant alignments: (bits) Value
AU089540 72 3e-26
AW164046 30 2.6
TC18021 similar to UP|AAH27795 (AAH27795) Zfp294 protein (Fragme... 29 3.4
BP051596 29 4.4
AV769010 28 5.7
TC19690 28 7.5
TC18614 similar to UP|Q94KA9 (Q94KA9) Importin alpha 2, partial ... 28 9.8
TC9112 weakly similar to UP|Q9SI70 (Q9SI70) F23N19.16, partial (... 28 9.8
AV768241 28 9.8
BF177663 28 9.8
>AU089540
Length = 660
Score = 71.6 bits (174), Expect(2) = 3e-26
Identities = 43/81 (53%), Positives = 49/81 (60%)
Frame = +2
Query: 848 LFYAK*RP*DHFLWINCTTGGRVILA*AA*LIFSRKTRWIILIQ*SNQWCIYLCSIS*PI 907
L+YA+ + *DHFLWINC TGGRV L A LIF K + I IQ SN WCI CSI I
Sbjct: 404 LYYARLQL*DHFLWINCNTGGRVHLGSAVLLIFCPKIQLICSIQSSNPWCICPCSIFLAI 583
Query: 908 QDLLSQITILCCFVLYTV*LA 928
Q S++ L TV*LA
Sbjct: 584 QGQASEVITWFWSALCTV*LA 646
Score = 65.1 bits (157), Expect(2) = 3e-26
Identities = 31/73 (42%), Positives = 45/73 (61%)
Frame = +3
Query: 753 KIQGFIQPENSWCIQAI*VLSYTGWEAAITRSQDPGRRLSYFATCWSMLRINYQIE*SNI 812
K QGFI+P+N W AI +L + ++AA+ R + LS F +CW LR +YQ +* NI
Sbjct: 186 KDQGFIKPKNPWRSPAIQILRWADFQAAVARRKITSS*LSPFISCWCYLRHSYQSK**NI 365
Query: 813 WCIWLHLYGDCCI 825
W+H+Y CC+
Sbjct: 366 RVSWIHIYCHCCL 404
>AW164046
Length = 421
Score = 29.6 bits (65), Expect = 2.6
Identities = 19/58 (32%), Positives = 26/58 (44%), Gaps = 4/58 (6%)
Frame = +1
Query: 160 DFSSSSETFC----SAGSYCPTTTTKFPCSSGLRKAEFM*F*HSNPKHACIWSYAHCC 213
+FSSS F Y P +T SS + + + *H N K +W Y+HCC
Sbjct: 244 NFSSSDYRFSLLLEKETEYPPDSTNCSEGSSICKSKDSLFT*HPNDKA**LWQYSHCC 417
>TC18021 similar to UP|AAH27795 (AAH27795) Zfp294 protein (Fragment),
partial (10%)
Length = 646
Score = 29.3 bits (64), Expect = 3.4
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = +1
Query: 561 KEAGKCWIRNGYGTFTFDVR*TNIWPGQCIISAPSKSTK 599
+EAG+C+ +GY F+ + R + P C IS S TK
Sbjct: 388 EEAGQCFATHGYVIFSIEAREMYLIPFSCKISPFSLYTK 504
>BP051596
Length = 505
Score = 28.9 bits (63), Expect = 4.4
Identities = 13/25 (52%), Positives = 18/25 (72%)
Frame = +1
Query: 159 ADFSSSSETFCSAGSYCPTTTTKFP 183
++ SSSSE+ S GSY T+T+ FP
Sbjct: 331 SESSSSSESLTSLGSYSSTSTSLFP 405
>AV769010
Length = 369
Score = 28.5 bits (62), Expect = 5.7
Identities = 11/36 (30%), Positives = 17/36 (46%), Gaps = 1/36 (2%)
Frame = +2
Query: 445 HRYHGPVRGWQDN-ISFSSSWKSTWMLGYWFNSDKW 479
H Y+GP R WQDN + + +W ++ W
Sbjct: 191 HYYYGPCRSWQDNTFGLYTEEQGCCFRSWWDHTGNW 298
>TC19690
Length = 581
Score = 28.1 bits (61), Expect = 7.5
Identities = 21/64 (32%), Positives = 26/64 (39%), Gaps = 1/64 (1%)
Frame = -3
Query: 94 ARTSNCRACCEGFFCPHGITCMIPCPLGSYCPIATLNKTTGVCEPYLYQLPPMQP-NHTC 152
A SN R FC C IP +G CP +T EP Y P++ N C
Sbjct: 279 ALLSNTRPRIRRVFCKVTSLCDIPGSVGPPCP-----RTETSPEPIPYCRGPLRALNEVC 115
Query: 153 GGAN 156
G +N
Sbjct: 114 GASN 103
>TC18614 similar to UP|Q94KA9 (Q94KA9) Importin alpha 2, partial (27%)
Length = 538
Score = 27.7 bits (60), Expect = 9.8
Identities = 20/90 (22%), Positives = 34/90 (37%), Gaps = 21/90 (23%)
Frame = -2
Query: 51 SANYLKPNRNCNLTSWVSGC---------EPGWACSVPSGQKIDLKDS-KEMPARTSN-- 98
SA Y++ C SW+ +P W VP+ + + KE+ A +
Sbjct: 408 SAEYVQSKAACETASWIQAACYCHHSKHPQP-WLGGVPTSSRRQWSERRKELAAEAGDPR 232
Query: 99 ---------CRACCEGFFCPHGITCMIPCP 119
C +C +CPHG+ ++ P
Sbjct: 231 VSSSTSSPPCDSCGSRPWCPHGVCDLLQRP 142
>TC9112 weakly similar to UP|Q9SI70 (Q9SI70) F23N19.16, partial (53%)
Length = 807
Score = 27.7 bits (60), Expect = 9.8
Identities = 20/65 (30%), Positives = 32/65 (48%), Gaps = 2/65 (3%)
Frame = +1
Query: 18 SSCIKKTKGDISNRLCTAAEVKFYLNSLME--KSSSANYLKPNRNCNLTSWVSGCEPGWA 75
+SC K + +L + FY L++ K S+ L+ +R C +TS +S C+ G A
Sbjct: 229 ASCCDAIKKTVETQLTCLCNL-FYSPGLLQSFKISTDQALQLSRRCGVTSDLSSCKSGSA 405
Query: 76 CSVPS 80
S S
Sbjct: 406 PSPAS 420
>AV768241
Length = 434
Score = 27.7 bits (60), Expect = 9.8
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +3
Query: 202 KHACIWSYAHCCFEYFATHYL 222
KH WSYA CC+E + ++ L
Sbjct: 324 KHT*PWSYASCCYE*YQSNLL 386
>BF177663
Length = 283
Score = 27.7 bits (60), Expect = 9.8
Identities = 25/79 (31%), Positives = 29/79 (36%), Gaps = 8/79 (10%)
Frame = +1
Query: 782 TRSQDPGRRLSYFATCWSMLRINYQIE*SNIWCIWLHLYGDCCILLYSTHHSFEEFFVLI 841
TRS R L + CW MLR CIW+ L I L S + FVL+
Sbjct: 10 TRSLANFRVLLCYVHCWEMLRT----------CIWIFL-----IFLQSLKFMHDGVFVLV 144
Query: 842 --------RCCQFQLFYAK 852
C F L Y K
Sbjct: 145 YESRYWFDLCLDFNLRYMK 201
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.354 0.153 0.599
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,789,625
Number of Sequences: 28460
Number of extensions: 447772
Number of successful extensions: 6001
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 4349
Number of HSP's successfully gapped in prelim test: 163
Number of HSP's that attempted gapping in prelim test: 1432
Number of HSP's gapped (non-prelim): 4825
length of query: 1036
length of database: 4,897,600
effective HSP length: 99
effective length of query: 937
effective length of database: 2,080,060
effective search space: 1949016220
effective search space used: 1949016220
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144928.9


