

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.8 + phase: 0 /pseudo
(359 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
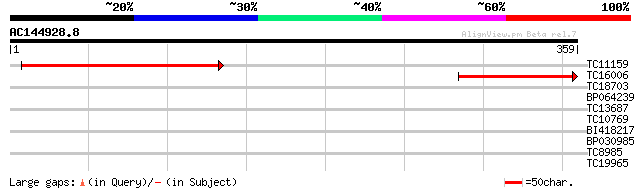
Score E
Sequences producing significant alignments: (bits) Value
TC11159 similar to UP|Q9ST64 (Q9ST64) ATPase (Fragment), partial... 224 1e-59
TC16006 similar to UP|Q9FF47 (Q9FF47) Arsenite translocating ATP... 107 3e-24
TC18703 weakly similar to UP|Q9FF47 (Q9FF47) Arsenite translocat... 39 0.002
BP064239 33 0.057
TC13687 similar to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determin... 32 0.16
TC10769 homologue to UP|Q9FI52 (Q9FI52) Nucleotide-binding prote... 31 0.28
BI418217 28 1.8
BP030985 27 5.3
TC8985 similar to UP|CAD27794 (CAD27794) Phosphatidylinositol-4-... 27 6.9
TC19965 homologue to UP|Q8VZF3 (Q8VZF3) At2g47390/T8I13.23, part... 26 9.0
>TC11159 similar to UP|Q9ST64 (Q9ST64) ATPase (Fragment), partial (31%)
Length = 581
Score = 224 bits (572), Expect = 1e-59
Identities = 109/128 (85%), Positives = 119/128 (92%)
Frame = +1
Query: 8 LSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVS 67
L V+SVAAP E+++ FDDMV+GTERKYY+LGGKGGVGKTSCA+SLAVKFANNGHPTLVVS
Sbjct: 196 LQVRSVAAPAEAVAGFDDMVSGTERKYYLLGGKGGVGKTSCASSLAVKFANNGHPTLVVS 375
Query: 68 TDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDF 127
TDPAHSLSDSFAQDLTGG LV V+GPD PLFALEINPEKARE+FR+ AK NGGS GVKDF
Sbjct: 376 TDPAHSLSDSFAQDLTGGLLVPVEGPDSPLFALEINPEKAREEFRNAAKTNGGSGGVKDF 555
Query: 128 MDGMGLGM 135
MDGMGLGM
Sbjct: 556 MDGMGLGM 579
>TC16006 similar to UP|Q9FF47 (Q9FF47) Arsenite translocating ATPase-like
protein, partial (18%)
Length = 609
Score = 107 bits (267), Expect = 3e-24
Identities = 58/75 (77%), Positives = 61/75 (81%)
Frame = +2
Query: 285 EERKCSCEETHCQSTSSTICI*LQVLCNEKKGSNTCS*YDSE*SRTLQPINDPSTTC*C* 344
EE KCSCEET+ QS SSTICI*LQVL EKKGS+ CS*YDSE* RTL I+D ST C*C
Sbjct: 2 EEGKCSCEETYRQSDSSTICI*LQVLFYEKKGSDACS*YDSE*PRTLLLIDDSSTPC*CR 181
Query: 345 DKRCSCSQISG*HYL 359
DKRCS SQISG*H L
Sbjct: 182 DKRCSSSQISG*HNL 226
>TC18703 weakly similar to UP|Q9FF47 (Q9FF47) Arsenite translocating
ATPase-like protein, partial (12%)
Length = 496
Score = 38.5 bits (88), Expect = 0.002
Identities = 29/65 (44%), Positives = 34/65 (51%)
Frame = +2
Query: 291 CEETHCQSTSSTICI*LQVLCNEKKGSNTCS*YDSE*SRTLQPINDPSTTC*C*DKRCSC 350
C ET C + CI LQVL NEK+GSN C+* +* RT T +K CS
Sbjct: 2 C*ET-CH*SGLASCIRLQVLFNEKEGSNACN*NYPQ*FRTFWFEIMRGTLSGYGNKGCSS 178
Query: 351 SQISG 355
SQI G
Sbjct: 179 SQIHG 193
>BP064239
Length = 534
Score = 33.5 bits (75), Expect = 0.057
Identities = 17/48 (35%), Positives = 25/48 (51%)
Frame = -1
Query: 40 KGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGAL 87
KGG GKT+ A LA +FA + + ++ DP S A GG++
Sbjct: 189 KGGSGKTTVAIILAGEFAKHDYSVAIIDADPQGSSYQWHASSSRGGSV 46
>TC13687 similar to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determining MinD)
(At5g24020), partial (45%)
Length = 470
Score = 32.0 bits (71), Expect = 0.16
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = +2
Query: 30 TERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
T R + GKGGVGKT+ A++ + A G + + D
Sbjct: 65 TPRVVVVTSGKGGVGKTTTTANMGLSLARLGFSVVAIDAD 184
>TC10769 homologue to UP|Q9FI52 (Q9FI52) Nucleotide-binding protein, partial
(43%)
Length = 478
Score = 31.2 bits (69), Expect = 0.28
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = +2
Query: 16 PTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFA 57
P + + +A + K +L GKGGVGK++ +A LA A
Sbjct: 200 PDPDLVAIAERMATVKHKILVLSGKGGVGKSTFSAQLAFALA 325
>BI418217
Length = 602
Score = 28.5 bits (62), Expect = 1.8
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = +1
Query: 27 VAGTERKYYMLGGKGGVGKTSCAASLAVKFAN 58
+ G + + GKGGVGK++ A +LAV A+
Sbjct: 229 IDGVKDTIAVASGKGGVGKSTTAVNLAVALAS 324
>BP030985
Length = 402
Score = 26.9 bits (58), Expect = 5.3
Identities = 18/52 (34%), Positives = 25/52 (47%), Gaps = 5/52 (9%)
Frame = +1
Query: 27 VAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVV-----STDPAHS 73
+A + K +GG G +GK ASL GHPT ++ +DPA S
Sbjct: 61 MAAADTKILSIGGTGFIGKFIVEASLKA-----GHPTYLLIRESSLSDPARS 201
>TC8985 similar to UP|CAD27794 (CAD27794) Phosphatidylinositol-4-phosphate
5-kinase (Fragment) , partial (13%)
Length = 726
Score = 26.6 bits (57), Expect = 6.9
Identities = 13/33 (39%), Positives = 17/33 (51%)
Frame = +1
Query: 291 CEETHCQSTSSTICI*LQVLCNEKKGSNTCS*Y 323
C + C + +T C L V+ N KGS TC Y
Sbjct: 493 CSQRACPADVTTYCYSLAVVRNP*KGSPTCFRY 591
>TC19965 homologue to UP|Q8VZF3 (Q8VZF3) At2g47390/T8I13.23, partial (17%)
Length = 503
Score = 26.2 bits (56), Expect = 9.0
Identities = 14/50 (28%), Positives = 23/50 (46%)
Frame = +1
Query: 42 GVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVD 91
G+G TS LA +FA PT+ + + +D + + L A V+
Sbjct: 250 GIGPTSALLWLARRFAILSGPTIPIIGEGDEEANDRYVEQLVASAEAAVE 399
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.347 0.152 0.531
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,617,741
Number of Sequences: 28460
Number of extensions: 101168
Number of successful extensions: 737
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 733
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 737
length of query: 359
length of database: 4,897,600
effective HSP length: 91
effective length of query: 268
effective length of database: 2,307,740
effective search space: 618474320
effective search space used: 618474320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144928.8


