

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144723.3 + phase: 0
(127 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
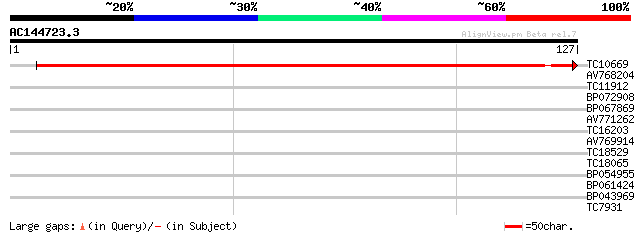
Score E
Sequences producing significant alignments: (bits) Value
TC10669 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, part... 224 3e-60
AV768204 34 0.007
TC11912 27 0.81
BP072908 27 1.4
BP067869 26 2.4
AV771262 25 3.1
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 25 3.1
AV769914 25 3.1
TC18529 25 5.2
TC18065 25 5.2
BP054955 24 6.8
BP061424 24 8.9
BP043969 24 8.9
TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial... 24 8.9
>TC10669 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, partial (87%)
Length = 1031
Score = 224 bits (571), Expect = 3e-60
Identities = 104/121 (85%), Positives = 111/121 (90%)
Frame = +1
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A GCSVNFEFLNYTIITSKCKGP YPPK+CCG+FKEFACPY DV+NDLTNDCASTMFSY
Sbjct: 418 AKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSY 597
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILL 126
INLYG+YPPGLFA ECREGKEGLAC ALPPS ADDT++Q+VH PSLVLVLTAC FLILL
Sbjct: 598 INLYGKYPPGLFAHECREGKEGLACPALPPSALADDTSSQVVHFPSLVLVLTAC-FLILL 774
Query: 127 F 127
F
Sbjct: 775 F 777
>AV768204
Length = 268
Score = 34.3 bits (77), Expect = 0.007
Identities = 16/30 (53%), Positives = 21/30 (69%)
Frame = -3
Query: 94 LPPSVSADDTANQIVHTPSLVLVLTACIFL 123
LPPS AD+T++Q+ H PSL L LTA +
Sbjct: 266 LPPSALADNTSSQVGHFPSLELGLTASFLI 177
>TC11912
Length = 505
Score = 27.3 bits (59), Expect = 0.81
Identities = 17/37 (45%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Frame = +2
Query: 90 ACDALPPSVSADDTANQIVHTP-SLVLVLTACIFLIL 125
A +LPPS ++ + VHT S VLVLT + LIL
Sbjct: 122 AISSLPPSSLPSSSSQEKVHTSISHVLVLTFLLLLIL 232
>BP072908
Length = 405
Score = 26.6 bits (57), Expect = 1.4
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = -1
Query: 84 EGKEGLACDALPPSVSADDT 103
+G EGL C A PP S D+
Sbjct: 348 QGSEGLTCSAAPPDQSTGDS 289
>BP067869
Length = 495
Score = 25.8 bits (55), Expect = 2.4
Identities = 11/17 (64%), Positives = 12/17 (69%)
Frame = -3
Query: 27 CKGPKYPPKECCGSFKE 43
C G + PPK CCGS KE
Sbjct: 346 CAG-QLPPKSCCGS*KE 299
>AV771262
Length = 360
Score = 25.4 bits (54), Expect = 3.1
Identities = 11/23 (47%), Positives = 12/23 (51%)
Frame = +1
Query: 66 YINLYGRYPPGLFASECREGKEG 88
Y+NLY RYP EC G G
Sbjct: 202 YVNLYRRYPST*VTVECTLGLHG 270
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 25.4 bits (54), Expect = 3.1
Identities = 14/43 (32%), Positives = 23/43 (52%)
Frame = -3
Query: 1 MTLFHIAPAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKE 43
M +F++ S + + ++TSK K +PP+ CGS KE
Sbjct: 1290 MNVFNLPLLHRSGGINPVRWFLVTSKYKNLPFPPR-LCGSTKE 1165
>AV769914
Length = 287
Score = 25.4 bits (54), Expect = 3.1
Identities = 15/37 (40%), Positives = 20/37 (53%), Gaps = 1/37 (2%)
Frame = +2
Query: 85 GKEGLA-CDALPPSVSADDTANQIVHTPSLVLVLTAC 120
GK G C+ LP +V+A D A+ + H P L T C
Sbjct: 107 GKTGAEDCNCLPFAVAAADAADLVGHPPILHNRHTCC 217
>TC18529
Length = 470
Score = 24.6 bits (52), Expect = 5.2
Identities = 12/36 (33%), Positives = 16/36 (44%), Gaps = 6/36 (16%)
Frame = +2
Query: 17 FLNYTIITSKCK------GPKYPPKECCGSFKEFAC 46
F++Y + SK GPK CGSF + C
Sbjct: 20 FISYNQLKSKTSSNVVLIGPKKAGSNKCGSFSSYTC 127
>TC18065
Length = 522
Score = 24.6 bits (52), Expect = 5.2
Identities = 11/35 (31%), Positives = 18/35 (51%)
Frame = +3
Query: 36 ECCGSFKEFACPYADVINDLTNDCASTMFSYINLY 70
ECCG F+C + +ND +C +++ LY
Sbjct: 180 ECCGLSMNFSCYWPIAMND--ENCGVYCKNFLYLY 278
>BP054955
Length = 525
Score = 24.3 bits (51), Expect = 6.8
Identities = 11/33 (33%), Positives = 17/33 (51%)
Frame = +2
Query: 34 PKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
P E C SF YADV+ LT + +++ +
Sbjct: 239 PSESCNSFGRTNLSYADVLIRLTPNAFNSILCF 337
>BP061424
Length = 362
Score = 23.9 bits (50), Expect = 8.9
Identities = 16/55 (29%), Positives = 25/55 (45%), Gaps = 10/55 (18%)
Frame = -2
Query: 26 KCKGPKYPPKECCGSFKEFAC-------PYADVINDLTNDCA---STMFSYINLY 70
+C P + CCG F AC P+ D + ++T C+ STM N++
Sbjct: 190 RC*EPMF*S*SCCGDFFCLACSLRLASKPF*DSLQEITFFCSFMYSTMAYDANVF 26
>BP043969
Length = 445
Score = 23.9 bits (50), Expect = 8.9
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = -3
Query: 78 FASECREGKEGLACDA 93
FAS C + ++G ACDA
Sbjct: 338 FASICTDTRKGCACDA 291
>TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial (29%)
Length = 1477
Score = 23.9 bits (50), Expect = 8.9
Identities = 11/36 (30%), Positives = 18/36 (49%)
Frame = +1
Query: 31 KYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
+YP C S + F+C + ++ + C S FSY
Sbjct: 1273 RYP----CSSLRSFSCLFCFLLYLCLSFCLSLQFSY 1368
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.141 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,901,586
Number of Sequences: 28460
Number of extensions: 43887
Number of successful extensions: 262
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 260
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 261
length of query: 127
length of database: 4,897,600
effective HSP length: 80
effective length of query: 47
effective length of database: 2,620,800
effective search space: 123177600
effective search space used: 123177600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 50 (23.9 bits)
Medicago: description of AC144723.3


