

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144514.8 - phase: 0 /pseudo
(555 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
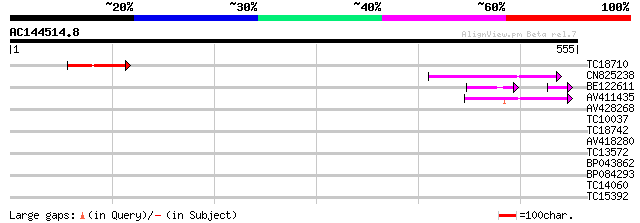
Score E
Sequences producing significant alignments: (bits) Value
TC18710 54 7e-08
CN825238 54 7e-08
BE122611 44 1e-06
AV411435 45 2e-05
AV428268 35 0.024
TC10037 33 0.092
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 32 0.21
AV418280 28 3.0
TC13572 28 3.9
BP043862 27 6.6
BP084293 27 8.7
TC14060 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {... 27 8.7
TC15392 27 8.7
>TC18710
Length = 843
Score = 53.9 bits (128), Expect = 7e-08
Identities = 27/63 (42%), Positives = 41/63 (64%), Gaps = 1/63 (1%)
Frame = +2
Query: 57 GFNNRWIY*MSMCVETVDFSVLVNNDAVGP-FIPGRGLRQGDPLSPYMFILCVEGLSSLI 115
GF+ W+ + + V V ++ +N VGP +P RGLRQGDP SPY+F+ +E LS LI
Sbjct: 638 GFSTNWVKMIMILVSGVSYNYKING-VVGPKLLPQRGLRQGDPFSPYLFLFTMEVLSLLI 814
Query: 116 RDA 118
+++
Sbjct: 815 QNS 823
>CN825238
Length = 692
Score = 53.9 bits (128), Expect = 7e-08
Identities = 36/133 (27%), Positives = 63/133 (47%), Gaps = 3/133 (2%)
Frame = +3
Query: 411 LIDDETKCWDVPFIQ*VFSKDIAYDVLNTSLVNQVTNDKLIWKAEKNGIYSVKST-FRLC 469
L+ + W+V I +F K+ A + + L + T + +W+ +NG +SV+S + +
Sbjct: 297 LLIPNARRWNVQLIDQLFPKETAAVIKSIPLGWRTTRNSCLWRWSRNGAFSVRSAYYNIQ 476
Query: 470 VEDLMDTTHLRCPEF-WMGIWRLKVPPKIKKSHVDDMLDEILPTRMQLKDKGV*CPTYCV 528
V M T+ F W +WR VP K+ K + I+P R L+ +G+ C
Sbjct: 477 VAKEMQTSSSAASSFPWKKLWRTPVPNKM-KHFAYRVARNIIPCRENLQRRGIAVQNECP 653
Query: 529 SYEHQ-EEDLEHI 540
+ HQ E + H+
Sbjct: 654 TCNHQIPESMSHL 692
>BE122611
Length = 357
Score = 44.3 bits (103), Expect(2) = 1e-06
Identities = 18/51 (35%), Positives = 29/51 (56%)
Frame = -3
Query: 448 DKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHLRCPEFWMGIWRLKVPPKIK 498
D+++W E NG YS KST+ L +ED + + FW +W+ P K++
Sbjct: 331 DRILWTEESNGCYSAKSTYNLLIEDGGNES-----RFWSLVWKANAPEKVR 194
Score = 25.0 bits (53), Expect(2) = 1e-06
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = -2
Query: 527 CVSYEHQEEDLEHIFFQCPFSVQVW 551
C+ ED H+F CP S +W
Sbjct: 113 CIRCGAPSEDANHVFRWCPGSAWMW 39
>AV411435
Length = 426
Score = 45.4 bits (106), Expect = 2e-05
Identities = 28/112 (25%), Positives = 48/112 (42%), Gaps = 6/112 (5%)
Frame = +3
Query: 446 TNDKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHLRCPE------FWMGIWRLKVPPKIKK 499
+ D+L W +G YSVKS +R + ++L W IW +VP K++
Sbjct: 33 SEDRLFWPFIPSGEYSVKSGYRAVKATQFNLSNLASSSSHPSSFLWHTIWGAQVPKKVR- 209
Query: 500 SHVDDMLDEILPTRMQLKDKGV*CPTYCVSYEHQEEDLEHIFFQCPFSVQVW 551
S + + +P + LK + + +C ED+ H C ++ VW
Sbjct: 210 SFLWRAANNAIPVKRNLKRRNMGRDDFCPICNKGPEDINHALLACEWTRAVW 365
>AV428268
Length = 429
Score = 35.4 bits (80), Expect = 0.024
Identities = 22/92 (23%), Positives = 40/92 (42%)
Frame = +3
Query: 460 YSVKSTFRLCVEDLMDTTHLRCPEFWMGIWRLKVPPKIKKSHVDDMLDEILPTRMQLKDK 519
YSVKS + + L + + + +W +KVP +++ + T++ L +
Sbjct: 6 YSVKSAY----DFLHGCANAQWDNIFKLLWSVKVPSNAIALSWKVLINRV-QTKVNLNRR 170
Query: 520 GV*CPTYCVSYEHQEEDLEHIFFQCPFSVQVW 551
GV C EE +H+ F CP ++W
Sbjct: 171 GVVISNVCPLCSLDEESTDHLLFSCPIV*RIW 266
>TC10037
Length = 560
Score = 33.5 bits (75), Expect = 0.092
Identities = 17/47 (36%), Positives = 27/47 (57%), Gaps = 1/47 (2%)
Frame = -3
Query: 506 LDEILPTRMQLKDKGV*CPT-YCVSYEHQEEDLEHIFFQCPFSVQVW 551
L I+ TR +L +KG PT +C+ + EE ++H CP + +VW
Sbjct: 453 LRSIVLTRKRLWNKGAVVPTIFCLRCDRLEETMDHALRDCP*ARRVW 313
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 32.3 bits (72), Expect = 0.21
Identities = 20/68 (29%), Positives = 35/68 (51%)
Frame = -3
Query: 136 ISHLLFADDCFLFFKADEGQTQVMKNILITYKAASGQAISLPISEIYCNQNVFDALKASI 195
ISHL FAD+ +F ++ N+L ++ SG + SE++ V + K I
Sbjct: 373 ISHLCFADNLMVFSNGYLESIAIINNVLHIFQHLSGLTPNPAKSEVFI--LVKETTKNQI 200
Query: 196 TNIIGVQD 203
TN++G ++
Sbjct: 199 TNMLGYKE 176
>AV418280
Length = 398
Score = 28.5 bits (62), Expect = 3.0
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = -2
Query: 523 CPTYCVSYEHQEEDLEHIFFQCPFSVQVWQ 552
CP +C + E E H+FF C FS+ VWQ
Sbjct: 388 CP-FCTTVE---ECSGHLFFTCVFSMGVWQ 311
>TC13572
Length = 534
Score = 28.1 bits (61), Expect = 3.9
Identities = 13/35 (37%), Positives = 21/35 (59%)
Frame = -3
Query: 432 IAYDVLNTSLVNQVTNDKLIWKAEKNGIYSVKSTF 466
++ D++ L+ V DK +WK ++G YSV S F
Sbjct: 145 LSLDIVRVPLLAGVP-DKWVWKPSEDGSYSVNSAF 44
>BP043862
Length = 558
Score = 27.3 bits (59), Expect = 6.6
Identities = 20/58 (34%), Positives = 26/58 (44%), Gaps = 5/58 (8%)
Frame = +3
Query: 448 DKLIWKAEKNGIYSVKSTFRLC---VEDLMDTTHLRCPE--FWMGIWRLKVPPKIKKS 500
D L +N I +++S F + V L+ H R W GIWRL P K KS
Sbjct: 189 DVLFVLGRRNKITNIRSCFNMLQG*VRVLLTEVHNRRQTTWHWPGIWRLT*PVKQSKS 362
>BP084293
Length = 344
Score = 26.9 bits (58), Expect = 8.7
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 3/49 (6%)
Frame = +1
Query: 3 LQFFLQISTSYLSDCMNLGLTWFFDFILLLLL---LAKS*NYTTLKAYH 48
L +S L D N+ +T DFI +LL L ++ +TT+K H
Sbjct: 145 LGLLCHVSNRNLEDLKNINITCI*DFISKILLSSFLTRTIRFTTVKKIH 291
>TC14060 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {Nicotiana
sylvestris;} , partial (49%)
Length = 567
Score = 26.9 bits (58), Expect = 8.7
Identities = 16/71 (22%), Positives = 34/71 (47%), Gaps = 2/71 (2%)
Frame = -2
Query: 418 CWD--VPFIQ*VFSKDIAYDVLNTSLVNQVTNDKLIWKAEKNGIYSVKSTFRLCVEDLMD 475
CW +P + *+ + L S+V++++ K++ ++ V+ ++C+ L
Sbjct: 500 CWSGGIPSLS*ILA-------LTLSIVSELSTSKVM-------VFPVRVLTKICMPPLNR 363
Query: 476 TTHLRCPEFWM 486
+T R FWM
Sbjct: 362 STKCRVDSFWM 330
>TC15392
Length = 1297
Score = 26.9 bits (58), Expect = 8.7
Identities = 10/25 (40%), Positives = 17/25 (68%)
Frame = +3
Query: 485 WMGIWRLKVPPKIKKSHVDDMLDEI 509
W G+ R + P + ++S VDD L+E+
Sbjct: 384 WGGVERTRCPSR*QRSKVDDSLEEV 458
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.343 0.152 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,020,258
Number of Sequences: 28460
Number of extensions: 166178
Number of successful extensions: 1271
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1251
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1267
length of query: 555
length of database: 4,897,600
effective HSP length: 95
effective length of query: 460
effective length of database: 2,193,900
effective search space: 1009194000
effective search space used: 1009194000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144514.8


