

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144483.8 + phase: 0
(253 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
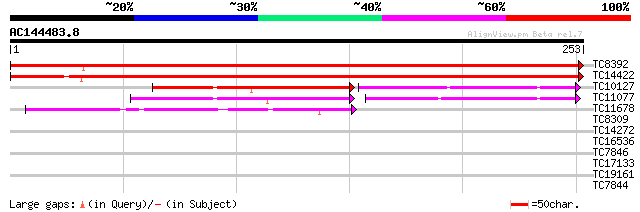
Score E
Sequences producing significant alignments: (bits) Value
TC8392 similar to UP|Q8LK52 (Q8LK52) Cp10-like protein, partial ... 397 e-111
TC14422 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor (1... 390 e-109
TC10127 weakly similar to GB|BAC16475.1|23237901|AP004671 10 kDa... 65 1e-11
TC11077 similar to UP|CH10_ARATH (P34893) 10 kDa chaperonin (Pro... 60 3e-10
TC11678 similar to GB|AAO64777.1|29028796|BT005842 At3g60210 {Ar... 54 3e-08
TC8309 UP|O22306 (O22306) Nlj21, complete 27 2.6
TC14272 homologue to UP|O49154 (O49154) Ferredoxin-dependent glu... 26 5.8
TC16536 similar to UP|O22477 (O22477) Betaine aldehyde dehydroge... 26 5.8
TC7846 homologue to UP|PSBO_PEA (P14226) Oxygen-evolving enhance... 26 7.5
TC17133 25 9.8
TC19161 similar to UP|Q9LTX3 (Q9LTX3) Similarity to pyridoxamine... 25 9.8
TC7844 homologue to UP|PSBO_PEA (P14226) Oxygen-evolving enhance... 25 9.8
>TC8392 similar to UP|Q8LK52 (Q8LK52) Cp10-like protein, partial (81%)
Length = 1014
Score = 397 bits (1019), Expect = e-111
Identities = 205/254 (80%), Positives = 226/254 (88%), Gaps = 1/254 (0%)
Frame = +1
Query: 1 MATTQLTASSISTRNLSSFERLRTSSIQFPR-NNARIAILTQWSLPSMVVKAATVVAPKH 59
MAT+Q TASSIS RNL S E LR S++QF RIA TQ PS+VVKAATVVAPKH
Sbjct: 109 MATSQFTASSISARNLPSLEGLRPSALQFHHVGRVRIATPTQKWFPSLVVKAATVVAPKH 288
Query: 60 TTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKA 119
T +KPLGDRVLVKIK+AEEKT GGILLPSTAQSKPQGGEVVA+G+GK++G NKVDISVK
Sbjct: 289 TAIKPLGDRVLVKIKEAEEKTVGGILLPSTAQSKPQGGEVVAVGEGKSIGNNKVDISVKT 468
Query: 120 GAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTA 179
GA+VVYSKYAGTEVEF+GSKHLILKD+DIVGILET+EV+DLKPLNDRVLIKVA AEEKTA
Sbjct: 469 GAKVVYSKYAGTEVEFNGSKHLILKDDDIVGILETDEVQDLKPLNDRVLIKVAAAEEKTA 648
Query: 180 GGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSD 239
GGLLLTEATK+KPSIGTVIA GPG +D+EGNRKPLS+ PG+TVLYSKYAGNDFKGKDGS+
Sbjct: 649 GGLLLTEATKEKPSIGTVIAAGPGTLDEEGNRKPLSVTPGSTVLYSKYAGNDFKGKDGSE 828
Query: 240 YIALRASDVMAILS 253
YI LR SDVMA LS
Sbjct: 829 YIVLRVSDVMATLS 870
>TC14422 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor (10 kDa
chaperonin) (Protein Cpn10) (groES protein), partial
(93%)
Length = 1077
Score = 390 bits (1001), Expect = e-109
Identities = 203/254 (79%), Positives = 230/254 (89%), Gaps = 1/254 (0%)
Frame = +2
Query: 1 MATTQLTASSISTRNLSSFERLRTSSIQFP-RNNARIAILTQWSLPSMVVKAATVVAPKH 59
MATTQLTASSISTRN++SFE LR IQFP RIA T+ S ++VKAATVVAPK
Sbjct: 128 MATTQLTASSISTRNVASFEGLRP--IQFPCAGLVRIANPTRRSFRGLLVKAATVVAPKF 301
Query: 60 TTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKA 119
T +KPLGDRVLVKIK++EEK+QGGILLP+TAQ+KPQGGEVVA+G+GKT+GK++V+ISV
Sbjct: 302 TAIKPLGDRVLVKIKESEEKSQGGILLPTTAQTKPQGGEVVAVGEGKTIGKSQVEISVTT 481
Query: 120 GAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTA 179
GAQVVYSKYAGTEVEF+G+KHLILKD+DIVGILETE++KDLKPLNDRVLIKVA AEEKTA
Sbjct: 482 GAQVVYSKYAGTEVEFNGTKHLILKDDDIVGILETEDIKDLKPLNDRVLIKVAVAEEKTA 661
Query: 180 GGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSD 239
GGLLLTEATK+KPSIGTVIAVGPGP+ +EG RKPL+I PGNTVLYSKYAGNDFKGKDGSD
Sbjct: 662 GGLLLTEATKEKPSIGTVIAVGPGPLGEEGKRKPLAIAPGNTVLYSKYAGNDFKGKDGSD 841
Query: 240 YIALRASDVMAILS 253
YI LR+SDVMAILS
Sbjct: 842 YITLRSSDVMAILS 883
>TC10127 weakly similar to GB|BAC16475.1|23237901|AP004671 10 kDa chaperonin
{Oryza sativa (japonica cultivar-group);} , complete
Length = 618
Score = 64.7 bits (156), Expect = 1e-11
Identities = 36/90 (40%), Positives = 57/90 (63%), Gaps = 1/90 (1%)
Frame = +2
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDG-KTVGKNKVDISVKAGAQ 122
P +RVL++ KT GGILLP + S+ G+V+A+G G + N + +SVK G Q
Sbjct: 89 PSLNRVLIEKILPPTKTSGGILLPEKS-SQLNSGKVIAVGPGSRDRAGNLIPVSVKEGDQ 265
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V+ +Y G++++ D + L+ +DEDI+GIL
Sbjct: 266 VLLPEYGGSQIKLDDKEFLLFRDEDILGIL 355
Score = 62.4 bits (150), Expect = 7e-11
Identities = 39/98 (39%), Positives = 54/98 (54%)
Frame = +2
Query: 155 EEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPL 214
E K P +RVLI+ KT+GG+LL E + S G VIAVGPG D GN P+
Sbjct: 68 EMAKRFLPSLNRVLIEKILPPTKTSGGILLPEKSSQLNS-GKVIAVGPGSRDRAGNLIPV 244
Query: 215 SILPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
S+ G+ VL +Y G+ K D +++ R D++ IL
Sbjct: 245 SVKEGDQVLLPEYGGSQIK-LDDKEFLLFRDEDILGIL 355
>TC11077 similar to UP|CH10_ARATH (P34893) 10 kDa chaperonin (Protein CPN10)
(Protein groES), partial (98%)
Length = 583
Score = 60.5 bits (145), Expect = 3e-10
Identities = 38/100 (38%), Positives = 58/100 (58%), Gaps = 1/100 (1%)
Frame = +3
Query: 54 VVAPKHTTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNK- 112
++ P + PL +RVLV+ KT GILLP + SK G+V+A+G G K
Sbjct: 63 LLLPMAKRLIPLFNRVLVEKIIPPSKTTTGILLPEKS-SKLNSGKVLAVGPGIHSNDGKL 239
Query: 113 VDISVKAGAQVVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V ++VK G V+ +Y GTEV+ D ++ + +D+DI+G L
Sbjct: 240 VPVTVKEGDTVLLPEYGGTEVKLDNKEYHLYRDDDILGTL 359
Score = 57.0 bits (136), Expect = 3e-09
Identities = 35/95 (36%), Positives = 53/95 (54%)
Frame = +3
Query: 158 KDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSIL 217
K L PL +RVL++ KT G+LL E + K + G V+AVGPG ++G P+++
Sbjct: 81 KRLIPLFNRVLVEKIIPPSKTTTGILLPEKSS-KLNSGKVLAVGPGIHSNDGKLVPVTVK 257
Query: 218 PGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
G+TVL +Y G + K D +Y R D++ L
Sbjct: 258 EGDTVLLPEYGGTEVK-LDNKEYHLYRDDDILGTL 359
>TC11678 similar to GB|AAO64777.1|29028796|BT005842 At3g60210 {Arabidopsis
thaliana;}, partial (75%)
Length = 637
Score = 53.5 bits (127), Expect = 3e-08
Identities = 47/147 (31%), Positives = 72/147 (48%), Gaps = 1/147 (0%)
Frame = +2
Query: 8 ASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHTTVKPLGD 67
AS+ T + TS +N R+ L + SL V AT P V P D
Sbjct: 5 ASTFVTLPTPFLHKTSTSVPSTSFSNKRLPFLKRNSLKVNAV--ATKWEP--AKVVPQAD 172
Query: 68 RVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAGAQVVYSK 127
RVL+++++ +KT GGILLP +A E +G+ TVG + VKAGA+V+++
Sbjct: 173 RVLIRLEELAQKTTGGILLPKSAVK----FERYLVGEVLTVGAEAGE--VKAGAKVLFTD 334
Query: 128 YAGTEVEF-DGSKHLILKDEDIVGILE 153
EV+ +KH K D++ ++E
Sbjct: 335 VNAYEVDLGTDAKHCFCKASDLLAVVE 415
Score = 35.0 bits (79), Expect = 0.012
Identities = 27/93 (29%), Positives = 47/93 (50%), Gaps = 2/93 (2%)
Frame = +2
Query: 162 PLNDRVLIKVAEAEEKTAGGLLLTEATK--DKPSIGTVIAVGPGPVDDEGNRKPLSILPG 219
P DRVLI++ E +KT GG+LL ++ ++ +G V+ VG + + K +L
Sbjct: 161 PQADRVLIRLEELAQKTTGGILLPKSAVKFERYLVGEVLTVGAEAGEVKAGAK---VLFT 331
Query: 220 NTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
+ Y G D K + +ASD++A++
Sbjct: 332 DVNAYEVDLGTDAK------HCFCKASDLLAVV 412
>TC8309 UP|O22306 (O22306) Nlj21, complete
Length = 1063
Score = 27.3 bits (59), Expect = 2.6
Identities = 37/191 (19%), Positives = 78/191 (40%), Gaps = 4/191 (2%)
Frame = +3
Query: 8 ASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHTTVKPLGD 67
+++I T ++ F + S + + N +++L ++ + A VA ++ + D
Sbjct: 66 STAI*TNLIAHFFSSKASIVSYTTLN--LSLLESFAYTLCISMATAQVASANSALHD--D 233
Query: 68 RVLVKIKDAEEKTQGGILLPSTAQSKPQG----GEVVAIGDGKTVGKNKVDISVKAGAQV 123
+KD EEKT ++ T + +P E A+ + + + S++A
Sbjct: 234 TTKASLKDHEEKTTEQVVETQTPEPEPVSEKTKEEDSAVTEASEPEPTEDNPSIEAETTE 413
Query: 124 VYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAGGLL 183
V + V + + + D E E K+ K D V ++ EA E+ L
Sbjct: 414 VVEEVVTVTVT---DEPKVEEKTDGEAKKEATETKETKESADPVEVQAPEAGEEPETVTL 584
Query: 184 LTEATKDKPSI 194
+A+K++ +
Sbjct: 585 KEDASKEEEKV 617
>TC14272 homologue to UP|O49154 (O49154) Ferredoxin-dependent glutamate
synthase (Fragment) , partial (40%)
Length = 1846
Score = 26.2 bits (56), Expect = 5.8
Identities = 18/50 (36%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Frame = +2
Query: 204 PVDDEGNRKPLSILPGNTVLYSKYAGNDF-KGKDGSDYIALRASDVMAIL 252
PVD+ G + + + GNT LY G F +GK G + A+R S A++
Sbjct: 713 PVDNTGFQPEDAAIVGNTCLYGATGGQVFIRGKAGERF-AVRNSLAEAVV 859
>TC16536 similar to UP|O22477 (O22477) Betaine aldehyde dehydrogenase ,
partial (71%)
Length = 1097
Score = 26.2 bits (56), Expect = 5.8
Identities = 16/46 (34%), Positives = 21/46 (44%)
Frame = +2
Query: 84 ILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAGAQVVYSKYA 129
+LLP KPQ + AI GK + + DI AG Y+ A
Sbjct: 263 LLLPRLLSKKPQLAKFEAIDSGKPLDEAAWDIDDVAGCFEYYADLA 400
>TC7846 homologue to UP|PSBO_PEA (P14226) Oxygen-evolving enhancer protein
1, chloroplast precursor (OEE1) (33 kDa subunit of
oxygen evolving system of photosystem II) (OEC 33 kDa
subunit) (33 kDa thylakoid membrane protein), partial
(50%)
Length = 689
Score = 25.8 bits (55), Expect = 7.5
Identities = 16/54 (29%), Positives = 26/54 (47%)
Frame = +3
Query: 70 LVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAGAQV 123
L K + + G + S QSKP+ GE+ IG +++ + D+ KA V
Sbjct: 315 LAKENNKSASSSKGKITLSVTQSKPETGEI--IGVFESIQPSDTDLGAKAPKDV 470
>TC17133
Length = 556
Score = 25.4 bits (54), Expect = 9.8
Identities = 17/63 (26%), Positives = 32/63 (49%), Gaps = 1/63 (1%)
Frame = -2
Query: 7 TASSISTRNLSSFERLRTSSIQFPRNNARIAILTQW-SLPSMVVKAATVVAPKHTTVKPL 65
T ++IS + + R R S + P +R + +LPS+ +++ + P T+V P
Sbjct: 339 TTAAISRNSAPALGRRRR*SAKPPHVPSRTSRARSMPALPSLRSRSSPLAKPSTTSVAPF 160
Query: 66 GDR 68
G+R
Sbjct: 159 GNR 151
>TC19161 similar to UP|Q9LTX3 (Q9LTX3) Similarity to pyridoxamine
5-phosphate oxidase, partial (15%)
Length = 507
Score = 25.4 bits (54), Expect = 9.8
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = +3
Query: 212 KPLSILPGNTVLYSKYAGNDFKGKDGS 238
K S++PG VLY +Y + K DGS
Sbjct: 99 KQSSVVPGRHVLYEEYEELERKFADGS 179
>TC7844 homologue to UP|PSBO_PEA (P14226) Oxygen-evolving enhancer protein
1, chloroplast precursor (OEE1) (33 kDa subunit of
oxygen evolving system of photosystem II) (OEC 33 kDa
subunit) (33 kDa thylakoid membrane protein), complete
Length = 1240
Score = 25.4 bits (54), Expect = 9.8
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = +2
Query: 80 TQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAGAQV 123
+QG I L S QSKP GEV IG +++ + D+ KA V
Sbjct: 920 SQGKITL-SVTQSKPDTGEV--IGVFESIQPSDTDLGAKAPKDV 1042
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.131 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,028,219
Number of Sequences: 28460
Number of extensions: 33080
Number of successful extensions: 144
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 127
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 132
length of query: 253
length of database: 4,897,600
effective HSP length: 88
effective length of query: 165
effective length of database: 2,393,120
effective search space: 394864800
effective search space used: 394864800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144483.8


