

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.8 - phase: 0 /pseudo
(82 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
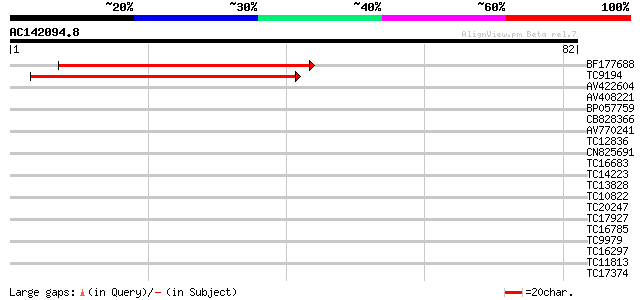
Score E
Sequences producing significant alignments: (bits) Value
BF177688 45 3e-06
TC9194 similar to UP|Q9LYP1 (Q9LYP1) Membrane protein (AT5g07250... 44 7e-06
AV422604 27 0.87
AV408221 26 1.5
BP057759 25 1.9
CB828366 25 2.5
AV770241 25 2.5
TC12836 GB|CAA71302.1|2292921|LJPANC pantoate--beta-alanine liga... 24 4.3
CN825691 24 4.3
TC16683 24 5.7
TC14223 similar to UP|Q8GZV0 (Q8GZV0) Obtusifoliol-14-demethylas... 24 5.7
TC13828 homologue to UP|Q9AXD9 (Q9AXD9) Mago nashi-like protein,... 24 5.7
TC10822 24 5.7
TC20247 similar to AAQ56809 (AAQ56809) At4g12700, partial (19%) 23 7.4
TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete 23 7.4
TC16785 23 7.4
TC9979 similar to UP|HIR5_MOUSE (Q9QZ23) HIRA-interacting protei... 23 7.4
TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type pro... 23 7.4
TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-lik... 23 7.4
TC17374 23 7.4
>BF177688
Length = 509
Score = 44.7 bits (104), Expect = 3e-06
Identities = 20/37 (54%), Positives = 27/37 (72%)
Frame = +1
Query: 8 MVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
++ V NL +GI+P V+NF IGG + GFLLGFV L +
Sbjct: 34 LIIVVNLALGILPHVDNFAHIGGFLSGFLLGFVFLIR 144
>TC9194 similar to UP|Q9LYP1 (Q9LYP1) Membrane protein
(AT5g07250/T28J14_190), partial (78%)
Length = 1467
Score = 43.5 bits (101), Expect = 7e-06
Identities = 19/39 (48%), Positives = 28/39 (71%)
Frame = +2
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
L++ ++ NL IG++P V+NF IGG + GFLLGF+ L
Sbjct: 872 LTLVLIIAVNLAIGMLPHVDNFAHIGGFLAGFLLGFIFL 988
>AV422604
Length = 460
Score = 26.6 bits (57), Expect = 0.87
Identities = 13/26 (50%), Positives = 15/26 (57%)
Frame = -2
Query: 42 LCKKDPFVLPDQKLHKRCLPIICFIL 67
L K DP + P K K CL IIC +L
Sbjct: 420 LIKCDPVLHPVPKFLKACLSIICEVL 343
>AV408221
Length = 354
Score = 25.8 bits (55), Expect = 1.5
Identities = 17/45 (37%), Positives = 25/45 (54%), Gaps = 3/45 (6%)
Frame = -2
Query: 31 LIPGFLLGFVLLCKKDPFVLP-DQKLHKRCLPII--CFILLSTGW 72
+ F +GF+L+ + VLP + +R L +I CF LLS GW
Sbjct: 185 IFQSF*VGFLLVYSINTKVLP*TSSIMQRFLVVIFLCFWLLSFGW 51
>BP057759
Length = 563
Score = 25.4 bits (54), Expect = 1.9
Identities = 17/42 (40%), Positives = 26/42 (61%), Gaps = 2/42 (4%)
Frame = +3
Query: 12 FNLTIGIVPIVNNFGLIGGLIPGFLLGFVL--LCKKDPFVLP 51
F+L IG P+++ GLIG LIP +G +L + + F+LP
Sbjct: 378 FSLRIG--PLLSMKGLIGTLIPFSRIGLLLS*VVEMGGFLLP 497
>CB828366
Length = 568
Score = 25.0 bits (53), Expect = 2.5
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = -3
Query: 43 CKKDPFVLPDQKLHKRCLPIICFIL 67
C KDP P +K R +P++C +L
Sbjct: 101 CCKDPPNNPQRKAWNRTMPVMCCML 27
>AV770241
Length = 447
Score = 25.0 bits (53), Expect = 2.5
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +3
Query: 35 FLLGFVLLCKKDPFVLPDQKLHKRCL 60
FL +L+C+K F +KL K CL
Sbjct: 237 FLTALLLMCQKQLFFAK*RKLKKTCL 314
>TC12836 GB|CAA71302.1|2292921|LJPANC pantoate--beta-alanine ligase {Lotus
corniculatus var. japonicus;} , complete
Length = 1310
Score = 24.3 bits (51), Expect = 4.3
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = +3
Query: 27 LIGGLIPGFLLGFVLLCKKDPFV 49
LIGG++ G LG L ++PFV
Sbjct: 387 LIGGVVLGMKLGLELRSWRNPFV 455
>CN825691
Length = 590
Score = 24.3 bits (51), Expect = 4.3
Identities = 8/19 (42%), Positives = 13/19 (68%)
Frame = +3
Query: 52 DQKLHKRCLPIICFILLST 70
DQK+ + C+P +CF+ T
Sbjct: 42 DQKVMELCVPSLCFLCTQT 98
>TC16683
Length = 476
Score = 23.9 bits (50), Expect = 5.7
Identities = 9/24 (37%), Positives = 15/24 (62%)
Frame = -1
Query: 56 HKRCLPIICFILLSTGWTGVLTKG 79
HK C P+ ILL++ W + ++G
Sbjct: 443 HKNCRPLPYRILLASHWVQLCSQG 372
>TC14223 similar to UP|Q8GZV0 (Q8GZV0) Obtusifoliol-14-demethylase, complete
Length = 1917
Score = 23.9 bits (50), Expect = 5.7
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = +3
Query: 20 PIVNNFGLIGGLIPGFLLGFVLLCKKD 46
PI+ F LIGGLI FL G +++ +++
Sbjct: 213 PILGGFPLIGGLI-RFLKGPIVMLREE 290
>TC13828 homologue to UP|Q9AXD9 (Q9AXD9) Mago nashi-like protein, partial
(89%)
Length = 599
Score = 23.9 bits (50), Expect = 5.7
Identities = 12/31 (38%), Positives = 15/31 (47%)
Frame = +1
Query: 38 GFVLLCKKDPFVLPDQKLHKRCLPIICFILL 68
GF +LC F + + RCL FILL
Sbjct: 460 GFEVLCLLSYFTSLQDQAYIRCLQYYAFILL 552
>TC10822
Length = 619
Score = 23.9 bits (50), Expect = 5.7
Identities = 13/46 (28%), Positives = 22/46 (47%), Gaps = 4/46 (8%)
Frame = +3
Query: 1 MFNLSIGMVPVFNLTIGIVPIVNNF----GLIGGLIPGFLLGFVLL 42
M N + + + IG+ +V NF L+ G + GF+ G +L
Sbjct: 282 MGNHGLTFAAIGGVYIGVEQLVQNFRGKRDLVNGAVGGFVAGAAIL 419
>TC20247 similar to AAQ56809 (AAQ56809) At4g12700, partial (19%)
Length = 540
Score = 23.5 bits (49), Expect = 7.4
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = -3
Query: 35 FLLGFVLLCKKDPFVLPDQKLHKRCLPIIC 64
FL ++L +K VL Q++ +RC+ +IC
Sbjct: 139 FLPKILVLVEKMSGVLIFQRIEERCVRLIC 50
>TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete
Length = 1793
Score = 23.5 bits (49), Expect = 7.4
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +3
Query: 53 QKLHKRCLPIICFILLST 70
Q+ + RCLP +CF LS+
Sbjct: 603 QRRYLRCLP*VCFPTLSS 656
>TC16785
Length = 771
Score = 23.5 bits (49), Expect = 7.4
Identities = 7/16 (43%), Positives = 11/16 (68%)
Frame = -1
Query: 47 PFVLPDQKLHKRCLPI 62
P+ +P Q+ HK LP+
Sbjct: 615 PYAMPSQQKHKHTLPV 568
>TC9979 similar to UP|HIR5_MOUSE (Q9QZ23) HIRA-interacting protein 5
(mHIRIP5), partial (36%)
Length = 1304
Score = 23.5 bits (49), Expect = 7.4
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +2
Query: 49 VLPDQKLHKRCLPIIC 64
VL K +K CLPI+C
Sbjct: 1010 VLGAPKKYKHCLPIVC 1057
>TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease,
partial (10%)
Length = 574
Score = 23.5 bits (49), Expect = 7.4
Identities = 13/45 (28%), Positives = 22/45 (48%), Gaps = 12/45 (26%)
Frame = -3
Query: 36 LLGFVLLCKKDPFVLPDQKL------------HKRCLPIICFILL 68
+ +LL ++PF L QK+ H C+P++C +LL
Sbjct: 551 VFSILLLITQEPFPL-HQKIELHCPQLLILFFHLECIPLLCLLLL 420
>TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-like protein,
partial (36%)
Length = 917
Score = 23.5 bits (49), Expect = 7.4
Identities = 7/14 (50%), Positives = 10/14 (71%)
Frame = -2
Query: 63 ICFILLSTGWTGVL 76
+CF+L+ GW G L
Sbjct: 493 LCFLLVERGWIGTL 452
>TC17374
Length = 593
Score = 23.5 bits (49), Expect = 7.4
Identities = 8/23 (34%), Positives = 14/23 (60%)
Frame = +3
Query: 55 LHKRCLPIICFILLSTGWTGVLT 77
+ K+CLPI C + + W ++T
Sbjct: 30 IKKQCLPISCHLWILIHWHLIIT 98
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.154 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,811,072
Number of Sequences: 28460
Number of extensions: 28265
Number of successful extensions: 156
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 156
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 156
length of query: 82
length of database: 4,897,600
effective HSP length: 58
effective length of query: 24
effective length of database: 3,246,920
effective search space: 77926080
effective search space used: 77926080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 48 (23.1 bits)
Medicago: description of AC142094.8


