

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.3 + phase: 0
(85 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
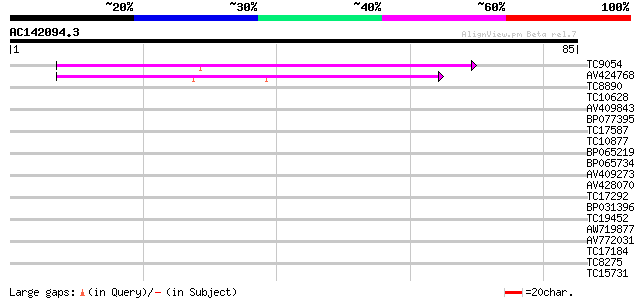
Score E
Sequences producing significant alignments: (bits) Value
TC9054 53 1e-08
AV424768 44 5e-06
TC8890 36 0.001
TC10628 similar to UP|Q8RX87 (Q8RX87) AT5g20250/F5O24_140, parti... 27 0.85
AV409843 26 1.1
BP077395 26 1.1
TC17587 similar to UP|HIS5_ARATH (Q9SZ30) Imidazole glycerol pho... 26 1.1
TC10877 similar to UP|O22840 (O22840) At2g43630 protein, partial... 26 1.4
BP065219 26 1.4
BP065734 25 1.9
AV409273 25 1.9
AV428070 25 2.5
TC17292 25 2.5
BP031396 24 4.2
TC19452 homologue to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, parti... 24 4.2
AW719877 24 5.5
AV772031 24 5.5
TC17184 similar to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, partial... 24 5.5
TC8275 homologue to UP|Q94C02 (Q94C02) Myo-inositol-1-phosphate ... 24 5.5
TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Pr... 24 5.5
>TC9054
Length = 697
Score = 52.8 bits (125), Expect = 1e-08
Identities = 27/64 (42%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Frame = +3
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H HR QI R SY DV R SE+QK D
Sbjct: 264 CKFHADSHKSECNMYCLDCMNGALCSLCLGHHKDHRAIQIRRSSYHDVIRVSEIQKVLDI 443
Query: 67 SNIQ 70
+ +Q
Sbjct: 444 TGVQ 455
>AV424768
Length = 417
Score = 43.9 bits (102), Expect = 5e-06
Identities = 24/60 (40%), Positives = 31/60 (51%), Gaps = 2/60 (3%)
Frame = +1
Query: 8 CDDHRDLRSNEKNTFCVDC-AVRFCRHCKEA-HSIHRRFQIYRYSYQDVFRHSELQKHFD 65
C H D +E N FC+DC FC +C+ + H H QI R SY +V R E+ K D
Sbjct: 235 CRIHGDAARSECNMFCLDCNGDAFCFYCRSSRHKDHHVIQIRRSSYHNVVRVGEIDKVLD 414
>TC8890
Length = 410
Score = 35.8 bits (81), Expect = 0.001
Identities = 20/50 (40%), Positives = 23/50 (46%), Gaps = 1/50 (2%)
Frame = +2
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFR 56
C H D +E N +C+DC C C H HR QI SY DV R
Sbjct: 242 CKMHADSHKSECNMYCLDCMNGALCSLCLAYHKDHRAIQI-GVSYHDVIR 388
>TC10628 similar to UP|Q8RX87 (Q8RX87) AT5g20250/F5O24_140, partial (35%)
Length = 1085
Score = 26.6 bits (57), Expect = 0.85
Identities = 16/37 (43%), Positives = 20/37 (53%)
Frame = -2
Query: 27 AVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKH 63
AV F + C E H++HR + R S V RHS Q H
Sbjct: 283 AVLFFKFCSEKHTLHRATR-RR*SLHHVLRHSR-QNH 179
>AV409843
Length = 414
Score = 26.2 bits (56), Expect = 1.1
Identities = 16/51 (31%), Positives = 23/51 (44%)
Frame = +1
Query: 22 FCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCSNIQVT 72
FC D A +C C E +QI R S Q R + +HF+C ++
Sbjct: 160 FCCDVA*LWCWQCSECQEC---YQIPRLSNQ---RCANPTRHFECKQTNLS 294
>BP077395
Length = 397
Score = 26.2 bits (56), Expect = 1.1
Identities = 19/71 (26%), Positives = 35/71 (48%), Gaps = 2/71 (2%)
Frame = +2
Query: 17 NEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDV--FRHSELQKHFDCSNIQVTLN 74
NE + F DC + + K++ +IH+ +YR + RH + K+ C N+ +
Sbjct: 2 NEISFFT*DCKLVYIS*SKQSTTIHKPE*VYRLQTKQACNLRH-*VAKYKSCPNMNFLVI 178
Query: 75 ALLQVIDSFSL 85
++L+ SF L
Sbjct: 179SMLKQQSSFCL 211
>TC17587 similar to UP|HIS5_ARATH (Q9SZ30) Imidazole glycerol phosphate
synthase hisHF, chloroplast precursor (IGP synthase)
(ImGP synthase) (IGPS) [Includes: Glutamine
amidotransferase ; Cyclase ] , partial (12%)
Length = 431
Score = 26.2 bits (56), Expect = 1.1
Identities = 16/51 (31%), Positives = 23/51 (44%)
Frame = +2
Query: 22 FCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCSNIQVT 72
FC D A +C C E +QI R S Q R + +HF+C ++
Sbjct: 152 FCCDVA*LWCWQCSECQEC---YQIPRLSNQ---RCANPTRHFECKQTNLS 286
>TC10877 similar to UP|O22840 (O22840) At2g43630 protein, partial (17%)
Length = 629
Score = 25.8 bits (55), Expect = 1.4
Identities = 9/28 (32%), Positives = 13/28 (46%)
Frame = -1
Query: 11 HRDLRSNEKNTFCVDCAVRFCRHCKEAH 38
H+D + + C A CRHC+ H
Sbjct: 524 HQDCHHHHRRRHCHPVAWLHCRHCQNIH 441
>BP065219
Length = 547
Score = 25.8 bits (55), Expect = 1.4
Identities = 13/26 (50%), Positives = 15/26 (57%), Gaps = 2/26 (7%)
Frame = +1
Query: 20 NTFCVDC--AVRFCRHCKEAHSIHRR 43
+T VDC RFC H +HS HRR
Sbjct: 190 STKLVDC*KQHRFCLHSYLSHSFHRR 267
>BP065734
Length = 406
Score = 25.4 bits (54), Expect = 1.9
Identities = 7/14 (50%), Positives = 11/14 (78%)
Frame = -3
Query: 26 CAVRFCRHCKEAHS 39
C++ +C HCKE +S
Sbjct: 317 CSISYCSHCKEHYS 276
>AV409273
Length = 417
Score = 25.4 bits (54), Expect = 1.9
Identities = 12/29 (41%), Positives = 14/29 (47%)
Frame = +3
Query: 6 GYCDDHRDLRSNEKNTFCVDCAVRFCRHC 34
G C+DHR +R NE F R R C
Sbjct: 174 GGCEDHRRIRRNEDRQFGGGGKARQRRRC 260
>AV428070
Length = 356
Score = 25.0 bits (53), Expect = 2.5
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +1
Query: 12 RDLRSNEKNTFCVDCA 27
RDL+S N CVDC+
Sbjct: 166 RDLQSEPSNKICVDCS 213
>TC17292
Length = 644
Score = 25.0 bits (53), Expect = 2.5
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = -2
Query: 28 VRFCRHCKEAHSIHRRFQIYRY 49
VR+CR E +IH FQ+Y Y
Sbjct: 337 VRYCRIYIEKLNIHIHFQLYFY 272
>BP031396
Length = 429
Score = 24.3 bits (51), Expect = 4.2
Identities = 8/20 (40%), Positives = 13/20 (65%)
Frame = -2
Query: 38 HSIHRRFQIYRYSYQDVFRH 57
HS+H FQI+ Y +V ++
Sbjct: 86 HSVHELFQIFTKLYSEVMKY 27
>TC19452 homologue to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, partial (41%)
Length = 562
Score = 24.3 bits (51), Expect = 4.2
Identities = 10/32 (31%), Positives = 17/32 (52%)
Frame = +3
Query: 47 YRYSYQDVFRHSELQKHFDCSNIQVTLNALLQ 78
YRY +VF+H + K D + + L +L+
Sbjct: 414 YRYMAPEVFKHRKYDKKVDVYSFAMILYEMLE 509
>AW719877
Length = 520
Score = 23.9 bits (50), Expect = 5.5
Identities = 8/13 (61%), Positives = 9/13 (68%)
Frame = +1
Query: 31 CRHCKEAHSIHRR 43
CR C+E H HRR
Sbjct: 292 CRGCRECHPDHRR 330
>AV772031
Length = 599
Score = 23.9 bits (50), Expect = 5.5
Identities = 7/12 (58%), Positives = 10/12 (83%)
Frame = -3
Query: 28 VRFCRHCKEAHS 39
+ +CRHCKE +S
Sbjct: 507 ISYCRHCKEHYS 472
>TC17184 similar to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, partial (33%)
Length = 598
Score = 23.9 bits (50), Expect = 5.5
Identities = 10/32 (31%), Positives = 16/32 (49%)
Frame = +3
Query: 47 YRYSYQDVFRHSELQKHFDCSNIQVTLNALLQ 78
YRY +VF+H K D + + L +L+
Sbjct: 111 YRYMAPEVFKHRRYDKKVDVFSFAMILYEMLE 206
>TC8275 homologue to UP|Q94C02 (Q94C02) Myo-inositol-1-phosphate synthase
, complete
Length = 1889
Score = 23.9 bits (50), Expect = 5.5
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = -1
Query: 15 RSNEKNTFCVDCAVRFCRHCKEAHSIHR 42
+S +KN FC VR+ R+C S R
Sbjct: 242 QSTDKNHFCSHERVRWFRNCTRTESQSR 159
>TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Probable
serine/threonine-protein kinase PK5) , partial (17%)
Length = 940
Score = 23.9 bits (50), Expect = 5.5
Identities = 16/57 (28%), Positives = 25/57 (43%), Gaps = 9/57 (15%)
Frame = -3
Query: 38 HSIHR---RFQIYRYSY------QDVFRHSELQKHFDCSNIQVTLNALLQVIDSFSL 85
H+IH+ Q+Y + +Q +CS IQ+T + LL +I S L
Sbjct: 440 HTIHKLQ*NMQLYNQALIAELD*PQALATKRVQNSINCSRIQITHSVLLNLI*SHPL 270
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.329 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,873,236
Number of Sequences: 28460
Number of extensions: 26436
Number of successful extensions: 179
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 179
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 179
length of query: 85
length of database: 4,897,600
effective HSP length: 61
effective length of query: 24
effective length of database: 3,161,540
effective search space: 75876960
effective search space used: 75876960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 48 (23.1 bits)
Medicago: description of AC142094.3


