

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
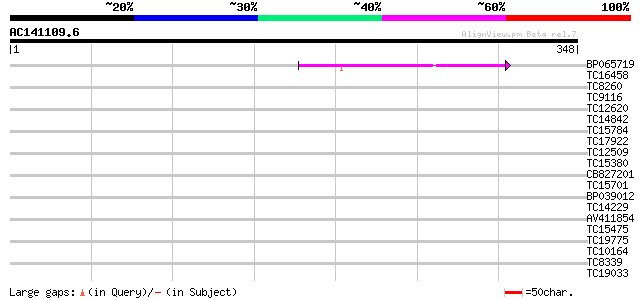
Query= AC141109.6 - phase: 0
(348 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP065719 62 1e-10
TC16458 similar to UP|O48679 (O48679) F3I6.5 protein, partial (20%) 37 0.006
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 35 0.014
TC9116 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydrata... 34 0.042
TC12620 weakly similar to UP|Q9FK05 (Q9FK05) Pectinesterase, par... 33 0.054
TC14842 similar to UP|RK4_TOBAC (O80361) 50S ribosomal protein L... 32 0.16
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 32 0.16
TC17922 31 0.27
TC12509 similar to UP|ACRO_HUMAN (P10323) Acrosin precursor , p... 31 0.35
TC15380 similar to UP|Q8H5P5 (Q8H5P5) OJ1003_C06.27 protein, par... 30 0.46
CB827201 30 0.46
TC15701 similar to UP|Q9FMM3 (Q9FMM3) Gb|AAC80623.1, partial (10%) 30 0.60
BP039012 30 0.60
TC14229 similar to UP|P93166 (P93166) SCOF-1, partial (38%) 30 0.60
AV411854 30 0.78
TC15475 similar to UP|Q93VZ2 (Q93VZ2) AT4g01050/F2N1_31, partial... 30 0.78
TC19775 similar to GB|AAF75234.1|8453100|AF233752 short-root pro... 30 0.78
TC10164 weakly similar to UP|O24099 (O24099) MtN12 protein (Frag... 30 0.78
TC8339 homologue to UP|Q84L89 (Q84L89) L-asparaginase, complete 30 0.78
TC19033 similar to UP|Q8H6X8 (Q8H6X8) GSK-3-like protein MsK4, p... 29 1.0
>BP065719
Length = 567
Score = 62.4 bits (150), Expect = 1e-10
Identities = 36/132 (27%), Positives = 64/132 (48%), Gaps = 2/132 (1%)
Frame = +3
Query: 178 VLNVQIPHKFKVPYFEKYKGNSCPE--EHLKMYVRRMSTYARNDQVFIYYFQESLASPAS 235
VL ++P +KVP F K+ G+S EH+ Y A N+ + + YF SL A
Sbjct: 147 VLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNAF 326
Query: 236 KWYTNLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTA 295
W+T L + T+ L +F EQ+ + + DL ++ ++ E+ +Y +R+R
Sbjct: 327 TWFTTLAPRSVHTWAQLERIFHEQF-FRGECKVSXKDLASVKRKPAESIDDYLNRFRMLK 503
Query: 296 TQVSPRIEEKEM 307
++ + E E+
Sbjct: 504 SRCFTHVSEHEL 539
>TC16458 similar to UP|O48679 (O48679) F3I6.5 protein, partial (20%)
Length = 539
Score = 36.6 bits (83), Expect = 0.006
Identities = 28/106 (26%), Positives = 41/106 (38%), Gaps = 8/106 (7%)
Frame = -3
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWF-------PPFTTGEILCPIAC 91
PPPPP T A + P S P H P PP T C A
Sbjct: 366 PPPPPPNTSPAADDLPTPPHMHNSSQPLSSTPSHYYPPNH*TPNPPPPPNTISPSCAPAK 187
Query: 92 EAQMPTHQYVAHVPPPPVRAPLVVMTYSAPM-IHTVPQIEEPIFHS 136
P+ + H P P ++ ++ + P +H+VP+ +PI+ S
Sbjct: 186 HPPPPSLHHHHHRPSPTPKSTCHMLLNTTPTHLHSVPKHSQPIYAS 49
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 35.4 bits (80), Expect = 0.014
Identities = 31/109 (28%), Positives = 40/109 (36%), Gaps = 5/109 (4%)
Frame = +3
Query: 33 HAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQ-----HSMPWFPPFTTGEILC 87
H PPPPP + + SS P H+ P HS P PP T
Sbjct: 345 HPVYHSPPPPPPKKHYKYSSPP---------PPVHTYPHPHPVYHSPP--PPVHTYPHPH 491
Query: 88 PIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHS 136
P+ Y + PPPP P + P +HT P P++HS
Sbjct: 492 PV----------YHSPPPPPPHEKPYKYASPPPPPVHTYP---HPVYHS 599
Score = 31.6 bits (70), Expect = 0.21
Identities = 16/44 (36%), Positives = 21/44 (47%), Gaps = 3/44 (6%)
Frame = +3
Query: 96 PTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEE---PIFHS 136
P Y H PPPPV +P P+ H+ P E P++HS
Sbjct: 132 PPKPYSYHSPPPPVHSPPPPYEKPHPVYHSPPPPYEKPHPVYHS 263
>TC9116 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydratase, small
subunit, partial (25%)
Length = 426
Score = 33.9 bits (76), Expect = 0.042
Identities = 20/78 (25%), Positives = 34/78 (42%)
Frame = +1
Query: 31 QAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIA 90
++HA + PPPP Q S+++ + + TPT S+P P + + P +
Sbjct: 175 KSHAPTLLLPPPPPNPQPHPSTASATS-SATTSTPTRSSP-------PSISPSSLPTPTS 330
Query: 91 CEAQMPTHQYVAHVPPPP 108
+ PT + P PP
Sbjct: 331 TRSSAPTPSSASPPPTPP 384
>TC12620 weakly similar to UP|Q9FK05 (Q9FK05) Pectinesterase, partial (10%)
Length = 430
Score = 33.5 bits (75), Expect = 0.054
Identities = 24/85 (28%), Positives = 32/85 (37%), Gaps = 6/85 (7%)
Frame = +1
Query: 34 AQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPP------FTTGEILC 87
A A+ PPP +T E+ E D+P ++ S PP +T
Sbjct: 172 APAEAPPPAKTQTSAESRRRQSPEPAAGHDSPLSASTLSSTSLAPPQPPSRTSSTSPST* 351
Query: 88 PIACEAQMPTHQYVAHVPPPPVRAP 112
P A T +H PPPP AP
Sbjct: 352 PTTTSAGRSTPPPASHSPPPPEPAP 426
>TC14842 similar to UP|RK4_TOBAC (O80361) 50S ribosomal protein L4,
chloroplast precursor (R-protein L4), partial (51%)
Length = 618
Score = 32.0 bits (71), Expect = 0.16
Identities = 25/109 (22%), Positives = 39/109 (34%)
Frame = +1
Query: 19 QAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFP 78
Q+ + L +++ ++ + PP P + S + S A P+ S P H P P
Sbjct: 118 QSSSLTLQNSKSPQNSLILIQPPSPSPASSQPSRFSPSPARKSASPPSTSKPPHQTPHVP 297
Query: 79 PFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVP 127
T PT + A PPPP A +P + P
Sbjct: 298 SSTA----------PSSPTSKTSAAAPPPPRPARRSAAVARSPTLRRRP 414
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 32.0 bits (71), Expect = 0.16
Identities = 26/87 (29%), Positives = 35/87 (39%), Gaps = 13/87 (14%)
Frame = +1
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQ-------------HSMPWFPPFTTGEI 85
PPPPP + +SS+T S T A TP S+PQ P P ++ E+
Sbjct: 187 PPPPPAPPRSASSSTTAS--TPTAATPPSSSPQLPSTKPNATPTSTSPSPATPSTSSSEL 360
Query: 86 LCPIACEAQMPTHQYVAHVPPPPVRAP 112
P A P+ + PPP P
Sbjct: 361 KPPSNSLAPTPSPAPTSSPPPPATS*P 441
>TC17922
Length = 470
Score = 31.2 bits (69), Expect = 0.27
Identities = 21/68 (30%), Positives = 30/68 (43%)
Frame = +1
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTH 98
PPPPP+ ++V +S++ S + P S P S P PP + PT
Sbjct: 61 PPPPPITSKVPTTSTSRSTT*L---PPQASPPTSSPPSSPP-----------SRSTTPTS 198
Query: 99 QYVAHVPP 106
Q A+ PP
Sbjct: 199 QATANAPP 222
>TC12509 similar to UP|ACRO_HUMAN (P10323) Acrosin precursor , partial (5%)
Length = 710
Score = 30.8 bits (68), Expect = 0.35
Identities = 14/42 (33%), Positives = 21/42 (49%)
Frame = +2
Query: 82 TGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMI 123
TG + P++ Q TH PPPP+ +P V + S M+
Sbjct: 158 TGLVQSPLSLSNQTTTHDQPPPPPPPPIHSPPDVQSASPSML 283
Score = 27.7 bits (60), Expect = 3.0
Identities = 14/50 (28%), Positives = 23/50 (46%)
Frame = +2
Query: 21 QAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAP 70
Q+ + Q H Q PPPPP+ + + S++ S + T T + P
Sbjct: 170 QSPLSLSNQTTTHDQPPPPPPPPIHSPPDVQSASPSMLVATSFTITITPP 319
>TC15380 similar to UP|Q8H5P5 (Q8H5P5) OJ1003_C06.27 protein, partial (28%)
Length = 1306
Score = 30.4 bits (67), Expect = 0.46
Identities = 22/86 (25%), Positives = 32/86 (36%), Gaps = 5/86 (5%)
Frame = +2
Query: 32 AHAQAQVPPPP-----PVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEIL 86
A + PP P P ASS++ + P+ ++P + PW PP T+G
Sbjct: 116 ASPSSSSPPSPTPLAAPTAPSTAASSTSSTARFPQTPNPSTASPPPTSPWTPPATSGS-- 289
Query: 87 CPIACEAQMPTHQYVAHVPPPPVRAP 112
A +PPPP P
Sbjct: 290 ---------------ASLPPPPPPPP 322
>CB827201
Length = 374
Score = 30.4 bits (67), Expect = 0.46
Identities = 27/109 (24%), Positives = 42/109 (37%), Gaps = 9/109 (8%)
Frame = +3
Query: 40 PPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMP----WFPPFTTGEIL-----CPIA 90
PPPP T V +SSST + T + + +A P + PP T P +
Sbjct: 21 PPPPCPTTVSSSSSTQHKAT*TQHSNSQNASSVPAPNTSLYAPPSTPTVAFPTNPPSPTS 200
Query: 91 CEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHSGNM 139
+ PT A PP ++ SA +++ P P+ N+
Sbjct: 201 PSSPSPTASTTASNSPPTPTTRSTPLSSSAAALNSSPTSSSPLHRRVNL 347
>TC15701 similar to UP|Q9FMM3 (Q9FMM3) Gb|AAC80623.1, partial (10%)
Length = 1030
Score = 30.0 bits (66), Expect = 0.60
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = -2
Query: 74 MPWFPPFTTGEILCPIACEAQMP 96
+P+FPPF T +CPIA ++ P
Sbjct: 699 LPFFPPFDTDTSVCPIARKSAFP 631
>BP039012
Length = 582
Score = 30.0 bits (66), Expect = 0.60
Identities = 25/103 (24%), Positives = 42/103 (40%)
Frame = +3
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTH 98
PPPP ++ + SST S + T + + + S+P P TT T
Sbjct: 144 PPPPLIKPSDQCESSTSSS----SSTTSFTQWRFSLPTTPNTTT--------------TQ 269
Query: 99 QYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHSGNMEV 141
PPPP +P + T+P ++E +FH +++
Sbjct: 270 NDSTTPPPPPSPSPSTITI-------TIPNLQE-LFHVSELQL 374
>TC14229 similar to UP|P93166 (P93166) SCOF-1, partial (38%)
Length = 1175
Score = 30.0 bits (66), Expect = 0.60
Identities = 28/99 (28%), Positives = 37/99 (37%)
Frame = +2
Query: 27 QAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEIL 86
QA A H + ++PPPPP T + + T T S + S P T
Sbjct: 437 QATANPHRKIKLPPPPPSTTTLPCRHPPV---TAERCTSVQSVTRASQPAKHSAVTSGAT 607
Query: 87 CPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHT 125
A A +PT +P P R P +AP HT
Sbjct: 608 TKAARAATVPT-ATSTQIPAAPSRLP----PTAAPPRHT 709
>AV411854
Length = 427
Score = 29.6 bits (65), Expect = 0.78
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = -3
Query: 172 AYDLCLVLNVQIPHKFKVPY 191
++DLC VLN+ + H F +PY
Sbjct: 356 SWDLCCVLNIIVKHLFPIPY 297
>TC15475 similar to UP|Q93VZ2 (Q93VZ2) AT4g01050/F2N1_31, partial (60%)
Length = 1271
Score = 29.6 bits (65), Expect = 0.78
Identities = 31/102 (30%), Positives = 47/102 (45%), Gaps = 3/102 (2%)
Frame = -2
Query: 14 TMAEAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTP--THSAPQ 71
T+ Q+ Q+ Q Q +QVP T +ASS T++ C P + S PQ
Sbjct: 799 TLQPVQSSHQI---CQEQTMWYSQVP*-----TSKKASSKTLACHHPCKLWPPVSSSTPQ 644
Query: 72 HSMPWFPPFTTGEILCPIACEAQMPTHQYVA-HVPPPPVRAP 112
H P +PP + ++L C+A +HQ+ A H P + P
Sbjct: 643 H--PGYPPA*SPQVL---LCQATEHSHQHQASHTHSSPTQHP 533
>TC19775 similar to GB|AAF75234.1|8453100|AF233752 short-root protein
{Arabidopsis thaliana;} , partial (7%)
Length = 581
Score = 29.6 bits (65), Expect = 0.78
Identities = 19/70 (27%), Positives = 27/70 (38%), Gaps = 1/70 (1%)
Frame = -1
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPW-FPPFTTGEILCPIACEAQMPT 97
PPP P + + ST+S T+ T + H P PP+++ P
Sbjct: 179 PPPAPSTSTISLHRSTVSSTTLPYTFSTTPSHHHKPPH*TPPYSS-------------PL 39
Query: 98 HQYVAHVPPP 107
H Y A PP
Sbjct: 38 HAYAASTSPP 9
>TC10164 weakly similar to UP|O24099 (O24099) MtN12 protein (Fragment),
partial (66%)
Length = 566
Score = 29.6 bits (65), Expect = 0.78
Identities = 27/100 (27%), Positives = 39/100 (39%)
Frame = +3
Query: 37 QVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMP 96
Q PPPP++T + IS HS P +PP+ I P
Sbjct: 15 QHSPPPPIQTYPPQIPNPIS----------HSPPPPGQT-YPPYIPNPIFL------SPP 143
Query: 97 THQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHS 136
++H PPPPV+ P+ H+ P + PI H+
Sbjct: 144 PPFPISHSPPPPVQT--YPPNIPIPIDHSPPPV--PINHA 251
>TC8339 homologue to UP|Q84L89 (Q84L89) L-asparaginase, complete
Length = 1467
Score = 29.6 bits (65), Expect = 0.78
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +1
Query: 315 LSYTTRRWSEVRPRTLLRWLVWVYSWKKES 344
LS++TR W+ R TL +VW+ +W+ S
Sbjct: 865 LSWSTRAWASSRLWTLSSSIVWMKAWRASS 954
>TC19033 similar to UP|Q8H6X8 (Q8H6X8) GSK-3-like protein MsK4, partial
(18%)
Length = 665
Score = 29.3 bits (64), Expect = 1.0
Identities = 14/19 (73%), Positives = 15/19 (78%)
Frame = -1
Query: 19 QAQAQVLAQAQAQAHAQAQ 37
QAQAQ AQAQAQAH+ Q
Sbjct: 530 QAQAQAQAQAQAQAHSLCQ 474
Score = 27.3 bits (59), Expect = 3.9
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -1
Query: 13 KTMAEAQAQAQVLAQAQAQAHAQAQVP 39
+ A+AQAQAQ A + Q H Q + P
Sbjct: 530 QAQAQAQAQAQAQAHSLCQCHHQGRAP 450
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.130 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,765,646
Number of Sequences: 28460
Number of extensions: 110287
Number of successful extensions: 1230
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 1129
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1195
length of query: 348
length of database: 4,897,600
effective HSP length: 91
effective length of query: 257
effective length of database: 2,307,740
effective search space: 593089180
effective search space used: 593089180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC141109.6


