

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140022.1 + phase: 0 /pseudo
(583 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
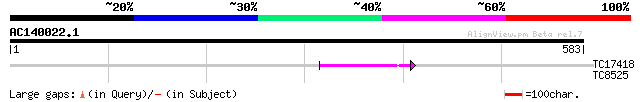
TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partia... 54 5e-08
TC8525 similar to UP|MDHP_PEA (P21528) Malate dehydrogenase [NAD... 32 0.37
>TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partial (20%)
Length = 739
Score = 54.3 bits (129), Expect = 5e-08
Identities = 44/99 (44%), Positives = 59/99 (59%), Gaps = 1/99 (1%)
Frame = -2
Query: 316 RRSMQVIF*RNLRWSIQSQFPRRLK-KS*S*QEKAMVKG*TQLITKV*LEV*DI*LQQGQ 374
R+ M +IF*RN R IQS FP+ L K + +G QLITKV*L V*DI*+Q Q
Sbjct: 738 RKYMLMIF*RNSR*QIQSTFPQLLXGKKLKLAGEMEARGLIQLITKV*LGV*DI*IQ*DQ 559
Query: 375 I*YMELVYLANTWRIRVLVTCKEPRGFFVILKVL*PKEF 413
I*++E ++W ++++ KE R I KVL* E+
Sbjct: 558 I*FVE*DCALDSWSHEIVIS-KEHRDLSDISKVL*RMEY 445
>TC8525 similar to UP|MDHP_PEA (P21528) Malate dehydrogenase [NADP],
chloroplast precursor (NADP-MDH) , partial (38%)
Length = 963
Score = 31.6 bits (70), Expect = 0.37
Identities = 15/32 (46%), Positives = 22/32 (67%)
Frame = -3
Query: 230 LILILFKTVFRDVHSSTHSTSNSLILEMFLLC 261
L+L+L KT H++ +STS S+IL +LLC
Sbjct: 634 LVLLLSKTFVALFHNNHYSTSISVILSQYLLC 539
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.369 0.164 0.579
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,646,895
Number of Sequences: 28460
Number of extensions: 130203
Number of successful extensions: 1752
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 982
Number of HSP's successfully gapped in prelim test: 78
Number of HSP's that attempted gapping in prelim test: 753
Number of HSP's gapped (non-prelim): 1091
length of query: 583
length of database: 4,897,600
effective HSP length: 95
effective length of query: 488
effective length of database: 2,193,900
effective search space: 1070623200
effective search space used: 1070623200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140022.1


