

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139882.5 - phase: 0
(379 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
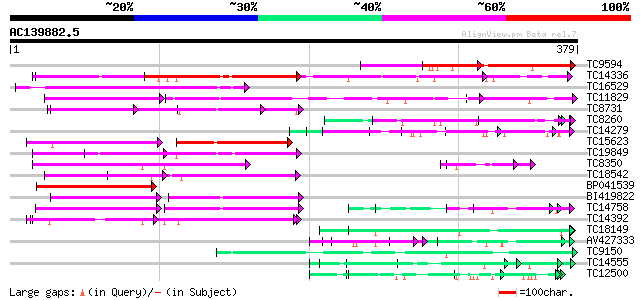
Score E
Sequences producing significant alignments: (bits) Value
TC9594 weakly similar to GB|AAM19966.1|20466089|AY098956 At2g326... 142 7e-35
TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 105 1e-23
TC16529 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 96 1e-20
TC11829 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, p... 91 3e-19
TC8731 homologue to UP|Q9LNS1 (Q9LNS1) F1L3.2, partial (34%) 90 7e-19
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 79 1e-15
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 79 1e-15
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 79 2e-15
TC19849 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, p... 77 5e-15
TC8350 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, par... 74 3e-14
TC18542 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding prote... 74 3e-14
BP041539 71 3e-13
BI419822 69 1e-12
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 69 2e-12
TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 68 2e-12
TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structura... 65 2e-11
AV427333 64 5e-11
TC9150 similar to UP|Q9M6E4 (Q9M6E4) Poly(A)-binding protein (Fr... 64 5e-11
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 63 7e-11
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 63 7e-11
>TC9594 weakly similar to GB|AAM19966.1|20466089|AY098956
At2g32600/T26B15.16 {Arabidopsis thaliana;}, partial
(9%)
Length = 502
Score = 142 bits (359), Expect = 7e-35
Identities = 73/107 (68%), Positives = 78/107 (72%), Gaps = 5/107 (4%)
Frame = +3
Query: 277 PPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHS-MQMPPPPPPQGMQGPPRPLPPPSVM 335
P PGQ WHQQQ GQ PP QQFRPP S MQMPPPP QG+ GPPR LPPP+VM
Sbjct: 18 PMPGQPTWHQQQXGQLPPPTMPPPQYQQFRPPPSGMQMPPPP--QGVPGPPRHLPPPAVM 191
Query: 336 AGQQ-QVWRPPPPP--QQQGGHPMGYPQNSMPPPPP-PNHHGMRPPS 378
GQ QVWRPPPPP QQQ G PM YPQ+SMPPPPP P++HGM PPS
Sbjct: 192 GGQPPQVWRPPPPPQYQQQVGRPMAYPQSSMPPPPPLPHNHGMPPPS 332
Score = 62.0 bits (149), Expect = 2e-10
Identities = 40/97 (41%), Positives = 46/97 (47%), Gaps = 15/97 (15%)
Frame = +3
Query: 235 QANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPP----GQ--QVW---- 284
Q G +P P P P PPP + PPPPQ P +PPP GQ QVW
Sbjct: 48 QQXGQLPPPTMPPPQYQQFRPPPSGMQMPPPPQGVPGPPRHLPPPAVMGGQPPQVWRPPP 227
Query: 285 ---HQQQQGQPMM--QQGMPPPMQQFRPPHSMQMPPP 316
+QQQ G+PM Q MPPP PH+ MPPP
Sbjct: 228 PPQYQQQVGRPMAYPQSSMPPPPPL---PHNHGMPPP 329
>TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(85%)
Length = 1917
Score = 105 bits (262), Expect = 1e-23
Identities = 92/332 (27%), Positives = 154/332 (45%), Gaps = 28/332 (8%)
Frame = +2
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
+ S ++ + +V NL +++ L +F + G + + V +D + + +GFV F +
Sbjct: 338 ESSTDKAKFNNVFVKNLSESTTDDELKNVFGEFGTITSAVVMRDG-DGKSKCFGFVNFEN 514
Query: 76 EEDADYAIKVLNMIKLYGKPIRVNKASQD---------------KKSLDV--GANLFIGN 118
+DA A++ LN K+ K V KA + K++ D GANL++ N
Sbjct: 515 ADDAARAVESLNGKKVDDKEWYVGKAQKKSEREQELKNKFEQSMKEAADKYQGANLYVKN 694
Query: 119 LDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQY 178
LD + ++ L + FS++G I T+ K+MRDP+ G SRG GF+++ + E + A+ MNG+
Sbjct: 695 LDDSIADEKLKELFSSYGTI-TSCKVMRDPN-GISRGSGFVAFSTPEEASRALLEMNGKM 868
Query: 179 LCNRQITVSYAYKKDTKGERHGTPAERVLAASNPTAQKSRPHT-LFASGPPSLPNAPQAN 237
+ ++ + V+ A +K+ + R + A P A P ++ G P +
Sbjct: 869 VVSKPLYVTLAQRKEDRRAR----LQAQFAQMRPVAMPPAPRVPMYPPGGPGIGQQIFYG 1036
Query: 238 GTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQ----------VWHQQ 287
PA +P +P G + ++P P F M M P GQQ Q
Sbjct: 1037QGPPAIIPSQP-GFGYQQQLVPGMRPGGAPVPNFF-MPMVPQGQQGQRPGGRRAGAVQQS 1210
Query: 288 QQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPP 319
QQ P+M Q M P + +R P MP P P
Sbjct: 1211QQPVPLMPQQMLPRGRVYRYPPGRGMPEVPMP 1306
Score = 92.0 bits (227), Expect = 1e-19
Identities = 103/392 (26%), Positives = 163/392 (41%), Gaps = 33/392 (8%)
Frame = +2
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S ++ ++ NLD I + L + F G +++ V D + Q +GYGFV+F +EE
Sbjct: 71 SIRKSGAGNIFIKNLDKAIDHKALHDTFSTFGNILSCKVATDS-SGQSKGYGFVQFETEE 247
Query: 78 DADYAIKVLNMIKLYGKPIRV--------NKASQDKKSLDVGANLFIGNLDPDVDEKLLY 129
A AI+ LN + L K + V ++S DK + N+F+ NL + L
Sbjct: 248 AAQNAIEKLNGMLLNDKQVYVGPFLRKQERESSTDKAKFN---NVFVKNLSESTTDDELK 418
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYA 189
+ F FG I T+ +MRD D G S+ FGF+++++ + + A+E++NG+ + +++ V A
Sbjct: 419 NVFGEFGTI-TSAVVMRDGD-GKSKCFGFVNFENADDAARAVESLNGKKVDDKEWYVGKA 592
Query: 190 YKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFAS------GPPSLPNAPQANGTIPAP 243
K K ER + + A K + L+ L + GTI +
Sbjct: 593 QK---KSEREQELKNKFEQSMKEAADKYQGANLYVKNLDDSIADEKLKELFSSYGTITSC 763
Query: 244 VPPRPFANGVAPPPIHVIQPPPPQAPAF-----QPMQMPPPGQQVWHQQQQGQPMMQQGM 298
R NG++ V P +A M + P Q+++ + Q
Sbjct: 764 KVMRD-PNGISRGSGFVAFSTPEEASRALLEMNGKMVVSKPLYVTLAQRKEDRRARLQA- 937
Query: 299 PPPMQQFRPPHSMQMPPPP-----PPQGM---------QGPPRPLPPPSVMAGQQQVWRP 344
QF + MPP P PP G QGPP +P QQQ+
Sbjct: 938 -----QFAQMRPVAMPPAPRVPMYPPGGPGIGQQIFYGQGPPAIIPSQPGFGYQQQL--- 1093
Query: 345 PPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRP 376
P + GG P+ N P P G RP
Sbjct: 1094-VPGMRPGGAPV---PNFFMPMVPQGQQGQRP 1177
Score = 72.8 bits (177), Expect = 9e-14
Identities = 41/106 (38%), Positives = 66/106 (61%), Gaps = 1/106 (0%)
Frame = +2
Query: 91 LYGKPIRVNKASQDKKSLDVGA-NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPD 149
L +PIR+ + +D GA N+FI NLD +D K L+DTFS FG I++ K+ D
Sbjct: 26 LNNRPIRIMYSHRDPSIRKSGAGNIFIKNLDKAIDHKALHDTFSTFGNILSC-KVATD-S 199
Query: 150 TGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTK 195
+G S+G+GF+ +++ EA+ +AIE +NG L ++Q+ V +K +
Sbjct: 200 SGQSKGYGFVQFETEEAAQNAIEKLNGMLLNDKQVYVGPFLRKQER 337
>TC16529 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(22%)
Length = 761
Score = 95.9 bits (237), Expect = 1e-20
Identities = 61/157 (38%), Positives = 87/157 (54%), Gaps = 1/157 (0%)
Frame = +1
Query: 5 IAPGVGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQ 64
+AP AN + + YVG+LD +++ L++LF Q G VV+V V +D T +
Sbjct: 298 VAPAPNANFV---------TTSLYVGDLDQNVNDSQLYDLFNQVGQVVSVRVCRDLTTRR 450
Query: 65 HQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDV 123
GYG+V F + DA A+ VLN L +PIRV + +D G AN+FI NLD +
Sbjct: 451 SLGYGYVNFSNPPDAARALDVLNFTPLNNRPIRVMYSHRDPSIRKSGTANIFIKNLDKAI 630
Query: 124 DEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFIS 160
D K L+DTFS+FG I+ + KI D +G S+ S
Sbjct: 631 DHKALHDTFSSFGSIL-SCKIATDA-SGQSKAMALFS 735
>TC11829 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, partial (4%)
Length = 738
Score = 90.9 bits (224), Expect = 3e-19
Identities = 73/220 (33%), Positives = 100/220 (45%), Gaps = 7/220 (3%)
Frame = +1
Query: 105 KKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSF 164
KK +D NL+IG L P +++ L F FG IV K+++D TG S+G+GF+ Y
Sbjct: 199 KKEID-DTNLYIGYLPPTLEDDGLIQLFQQFGEIVM-AKVIKDRMTGLSKGYGFVKYADI 372
Query: 165 EASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFA 224
+++AI AMNG L R I V A G P + V+ P A
Sbjct: 373 TMANNAILAMNGYRLEGRTIAVRVA----------GKPPQPVVPPGPP-----------A 489
Query: 225 SGPPSLPNAPQANGTIPAPVPPRPFAN----GVAPPPIHVIQP---PPPQAPAFQPMQMP 277
SG P+ P Q +G P+ + +A GVAPP + P PP P + P P
Sbjct: 490 SGMPTYPVPSQPHGAYPS---QQQYAAGGPLGVAPPTNYGGTPVPWAPPVPPPYAPYAPP 660
Query: 278 PPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPP 317
PPG + + QGQPM PP+ + PPPP
Sbjct: 661 PPGSTM-YPPMQGQPM-------------PPYGVHYPPPP 738
Score = 55.5 bits (132), Expect = 1e-08
Identities = 30/80 (37%), Positives = 45/80 (55%)
Frame = +1
Query: 24 DATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAI 83
D Y+G L P + ++ L +LF Q G +V V KDR+T +GYGFV++ A+ AI
Sbjct: 214 DTNLYIGYLPPTLEDDGLIQLFQQFGEIVMAKVIKDRMTGLSKGYGFVKYADITMANNAI 393
Query: 84 KVLNMIKLYGKPIRVNKASQ 103
+N +L G+ I V A +
Sbjct: 394 LAMNGYRLEGRTIAVRVAGK 453
Score = 42.4 bits (98), Expect = 1e-04
Identities = 28/78 (35%), Positives = 33/78 (41%), Gaps = 4/78 (5%)
Frame = +1
Query: 306 RPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPP----PPQQQGGHPMGYPQN 361
+PP + +PP PP GM P P P QQQ P PP GG P+ +
Sbjct: 451 KPPQPV-VPPGPPASGMPTYPVPSQPHGAYPSQQQYAAGGPLGVAPPTNYGGTPVPWA-- 621
Query: 362 SMPPPPPPNHHGMRPPSG 379
PP PPP PP G
Sbjct: 622 --PPVPPPYAPYAPPPPG 669
>TC8731 homologue to UP|Q9LNS1 (Q9LNS1) F1L3.2, partial (34%)
Length = 721
Score = 89.7 bits (221), Expect = 7e-19
Identities = 53/148 (35%), Positives = 82/148 (54%), Gaps = 2/148 (1%)
Frame = +2
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
+ YVGN+ Q++E LL E+F GPV + + + YGF+ + A AI
Sbjct: 284 SVYVGNIHTQVTELLLQEVFAGTGPVEGCKLFR----KEKSSYGFIHYFDRRSAALAILT 451
Query: 86 LNMIKLYGKPIRVNKASQDKKSLDVGA--NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
LN L+G+PI+VN A + D N+F+G+L P+V + L+ FS + ++ +
Sbjct: 452 LNGRHLFGQPIKVNWAYASGQREDTSGHYNIFVGDLSPEVTDATLFACFSVYPSC-SDAR 628
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAI 171
+M D TG SRGFGF+S+ S + + SAI
Sbjct: 629 VMWDQKTGRSRGFGFVSFRSQQDAQSAI 712
Score = 42.4 bits (98), Expect = 1e-04
Identities = 21/59 (35%), Positives = 35/59 (58%)
Frame = +2
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVL 86
+VG+L P++++ L+ F + V D+ T + +G+GFV FRS++DA AI L
Sbjct: 545 FVGDLSPEVTDATLFACFSVYPSCSDARVMWDQKTGRSRGFGFVSFRSQQDAQSAINDL 721
Score = 40.8 bits (94), Expect = 4e-04
Identities = 27/88 (30%), Positives = 47/88 (52%), Gaps = 4/88 (4%)
Frame = +2
Query: 113 NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIE 172
++++GN+ V E LL + F+ G V K+ R + +GFI Y ++ AI
Sbjct: 284 SVYVGNIHTQVTELLLQEVFAGTGP-VEGCKLFRKEKSS----YGFIHYFDRRSAALAIL 448
Query: 173 AMNGQYLCNRQITVSYAY----KKDTKG 196
+NG++L + I V++AY ++DT G
Sbjct: 449 TLNGRHLFGQPIKVNWAYASGQREDTSG 532
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 79.3 bits (194), Expect = 1e-15
Identities = 46/138 (33%), Positives = 59/138 (42%), Gaps = 10/138 (7%)
Frame = +3
Query: 243 PVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQ----VWHQ---QQQGQPMMQ 295
P PP+P++ PPP+H PPPP PPP + V+H +P
Sbjct: 126 PPPPKPYSYHSPPPPVH--SPPPPYEKPHPVYHSPPPPYEKPHPVYHSPPPPPPHKPYKY 299
Query: 296 QGMPPPMQQFRPPHS---MQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQG 352
PPP ++ PH PPPPPP+ P PP V+ PPPP
Sbjct: 300 PSPPPPPHKYPHPHPHPVYHSPPPPPPKKHYKYSSPPPPVHTYPHPHPVYHSPPPPVHTY 479
Query: 353 GHPMGYPQNSMPPPPPPN 370
HP +P PPPPPP+
Sbjct: 480 PHP--HPVYHSPPPPPPH 527
Score = 77.8 bits (190), Expect = 3e-15
Identities = 50/174 (28%), Positives = 66/174 (37%), Gaps = 12/174 (6%)
Frame = +3
Query: 211 NPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVI--------- 261
+P +PH ++ S PP + P P P +P+ PPP H
Sbjct: 177 SPPPPYEKPHPVYHSPPPPYEKPHPVYHSPPPPPPHKPYKYPSPPPPPHKYPHPHPHPVY 356
Query: 262 --QPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSM-QMPPPPP 318
PPPP ++ PPP H + PPP+ + PH + PPPPP
Sbjct: 357 HSPPPPPPKKHYKYSSPPPPVHTYPHPHP-----VYHSPPPPVHTYPHPHPVYHSPPPPP 521
Query: 319 PQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHH 372
P PPP V V+ PPPP P Y ++ PPPP HH
Sbjct: 522 PHEKPYKYASPPPPPVHTYPHPVYHSPPPPVHSPPPPHYYYKS----PPPPYHH 671
Score = 75.9 bits (185), Expect = 1e-14
Identities = 48/168 (28%), Positives = 66/168 (38%), Gaps = 1/168 (0%)
Frame = +3
Query: 211 NPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPA 270
+P +PH ++ S PP P+ P P+P PP P P++ PPPP
Sbjct: 219 SPPPPYEKPHPVYHSPPPPPPHKPYK---YPSPPPPPHKYPHPHPHPVYHSPPPPPPKKH 389
Query: 271 FQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSM-QMPPPPPPQGMQGPPRPL 329
++ PPP H + PPP+ + PH + PPPPPP
Sbjct: 390 YKYSSPPPPVHTYPHPHP-----VYHSPPPPVHTYPHPHPVYHSPPPPPPHEKPYKYASP 554
Query: 330 PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPP 377
PPP V V+ PPPP PPPP+++ PP
Sbjct: 555 PPPPVHTYPHPVYHSPPPPVHS--------------PPPPHYYYKSPP 656
Score = 47.0 bits (110), Expect = 5e-06
Identities = 25/66 (37%), Positives = 32/66 (47%), Gaps = 1/66 (1%)
Frame = +3
Query: 314 PPPPPPQGMQGPPRPL-PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHH 372
PPPP P PP P+ PP V+ PPPP ++ HP+ + PPPPP H
Sbjct: 126 PPPPKPYSYHSPPPPVHSPPPPYEKPHPVYHSPPPPYEK-PHPVYH-----SPPPPPPHK 287
Query: 373 GMRPPS 378
+ PS
Sbjct: 288 PYKYPS 305
Score = 35.4 bits (80), Expect = 0.016
Identities = 18/50 (36%), Positives = 22/50 (44%), Gaps = 3/50 (6%)
Frame = +3
Query: 331 PPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPP---PPPNHHGMRPP 377
P + A PPPPP+ H P +S PPP P P +H PP
Sbjct: 84 PSEISANHYAYSSPPPPPKPYSYHSPPPPVHSPPPPYEKPHPVYHSPPPP 233
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 79.0 bits (193), Expect = 1e-15
Identities = 49/149 (32%), Positives = 59/149 (38%), Gaps = 12/149 (8%)
Frame = +1
Query: 241 PAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPP 300
P P P+P+ PPP + + PPP +P+ PPP Q P PP
Sbjct: 28 PPPPVPKPYYYKSPPPPPYYYKSPPPPSPS------PPPPYYY-----QSPPPPSHSPPP 174
Query: 301 PMQQFRPPHSMQMPPPP-----PPQGMQGPPRPL-----PPPSVMAGQQQVWRPPPPPQQ 350
P PP PPPP PP PP P PPPS + ++ PPPP
Sbjct: 175 PYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPTS 354
Query: 351 QGGHPMGY--PQNSMPPPPPPNHHGMRPP 377
P Y P P PPPP H+ PP
Sbjct: 355 SPPPPYHYVSPPPPSPSPPPPYHYASPPP 441
Score = 78.6 bits (192), Expect = 2e-15
Identities = 55/179 (30%), Positives = 73/179 (40%), Gaps = 11/179 (6%)
Frame = +1
Query: 210 SNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAP 269
S P S P + PP ++P +P PP P +PPP + + PPP +P
Sbjct: 94 SPPPPSPSPPPPYYYQSPPPPSHSPPPPYYYKSPPPPSP-----SPPPPYYYKSPPPPSP 258
Query: 270 AFQP---MQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPP 326
+ P + PPP + H P + PPP PP+ PPPP P PP
Sbjct: 259 SPPPPYYYKSPPPPSPIPHP-----PYYYKSPPPPTSSPPPPYHYVSPPPPSPS----PP 411
Query: 327 RPL-----PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPP---PPPNHHGMRPP 377
P PPPS +++ PPPP + P Y S PPP P P + PP
Sbjct: 412 PPYHYASPPPPSPSPAPTYIYKSPPPPVKLPPPPYHY--TSPPPPSPSPAPTYIYKSPP 582
Score = 67.0 bits (162), Expect = 5e-12
Identities = 47/172 (27%), Positives = 68/172 (39%), Gaps = 4/172 (2%)
Frame = +1
Query: 199 HGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPI 258
H P + P + P + S PP P+ P +P PP P + PP
Sbjct: 160 HSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPY-YYKSPPPPSPIPH----PPY 324
Query: 259 HVIQPPPPQA---PAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFR-PPHSMQMP 314
+ PPPP + P + + PPP P P P ++ PP +++P
Sbjct: 325 YYKSPPPPTSSPPPPYHYVSPPPPSPSPPPPYHYASPPPPSPSPAPTYIYKSPPPPVKLP 504
Query: 315 PPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPP 366
PPP PP P P P+ +++ PPPP + P+ Y S PPP
Sbjct: 505 PPPYHYTSPPPPSPSPAPT------YIYKSPPPPTKSPPPPV-YIYASPPPP 639
Score = 64.7 bits (156), Expect = 2e-11
Identities = 44/135 (32%), Positives = 53/135 (38%), Gaps = 20/135 (14%)
Frame = +1
Query: 263 PPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPP-----PP 317
PPPP P + PPP P + PPP PP+ Q PP PP
Sbjct: 25 PPPPPVPKPYYYKSPPP-----------PPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPP 171
Query: 318 PPQGMQGPPRPL------------PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPP 365
PP + PP P PPPS ++ PPPP HP Y ++ PP
Sbjct: 172 PPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPI-PHPPYYYKSPPPP 348
Query: 366 ---PPPPNHHGMRPP 377
PPPP H+ PP
Sbjct: 349 TSSPPPPYHYVSPPP 393
Score = 50.8 bits (120), Expect = 4e-07
Identities = 29/88 (32%), Positives = 36/88 (39%), Gaps = 2/88 (2%)
Frame = +1
Query: 292 PMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQ 351
P + PPP +P + PPPP PP P PPP ++ PPPP
Sbjct: 4 PYWYKSPPPPPPVPKPYYYKSPPPPPYYYKSPPPPSPSPPP------PYYYQSPPPPSHS 165
Query: 352 GGHPMGY--PQNSMPPPPPPNHHGMRPP 377
P Y P P PPPP ++ PP
Sbjct: 166 PPPPYYYKSPPPPSPSPPPPYYYKSPPP 249
Score = 49.7 bits (117), Expect = 8e-07
Identities = 37/145 (25%), Positives = 48/145 (32%), Gaps = 3/145 (2%)
Frame = +1
Query: 188 YAYKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPR 247
Y YK P + P+ P+ + PP+ P + P P P
Sbjct: 226 YYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPTSSPPPPYHYVSPPPPSPS 405
Query: 248 PFANGVAPPPIHVIQPPPPQ---APAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQ 304
P PPP H PPPP AP + PPP + P P P
Sbjct: 406 P------PPPYHYASPPPPSPSPAPTYIYKSPPPPVKLPPPPYHYTSPPPPSPSPAPTYI 567
Query: 305 FRPPHSMQMPPPPPPQGMQGPPRPL 329
++ P PPPP PP P+
Sbjct: 568 YKSPPPPTKSPPPPVYIYASPPPPI 642
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 78.6 bits (192), Expect = 2e-15
Identities = 41/78 (52%), Positives = 54/78 (68%)
Frame = +1
Query: 112 ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAI 171
+ LFIG L VD+K L D FS+FG V +++ D DTG SRGFGF+S+DS E++ SA+
Sbjct: 184 SKLFIGGLSYGVDDKSLEDAFSSFGT-VAEARVIVDRDTGRSRGFGFVSFDSEESASSAL 360
Query: 172 EAMNGQYLCNRQITVSYA 189
+M+GQ L R I VSYA
Sbjct: 361 SSMDGQDLNGRNIRVSYA 414
Score = 48.9 bits (115), Expect = 1e-06
Identities = 33/105 (31%), Positives = 52/105 (49%), Gaps = 14/105 (13%)
Frame = +1
Query: 12 NLLGQHSAERNQDATA--------------YVGNLDPQISEELLWELFVQAGPVVNVYVP 57
N+L Q +A+R+Q A ++G L + ++ L + F G V V
Sbjct: 103 NVLRQGAAQRSQAPVASMLNYIRCMSSSKLFIGGLSYGVDDKSLEDAFSSFGTVAEARVI 282
Query: 58 KDRVTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKAS 102
DR T + +G+GFV F SEE A A+ ++ L G+ IRV+ A+
Sbjct: 283 VDRDTGRSRGFGFVSFDSEESASSALSSMDGQDLNGRNIRVSYAN 417
>TC19849 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, partial
(30%)
Length = 603
Score = 77.0 bits (188), Expect = 5e-15
Identities = 54/162 (33%), Positives = 89/162 (54%), Gaps = 17/162 (10%)
Frame = +2
Query: 51 VVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQ-DKKSLD 109
+ + V +D V + + +GFV F + ++A A+ LN K K V KA + ++ L+
Sbjct: 5 ITSAVVMRD-VDGKSKCFGFVNFENADEAAKAVDALNGKKFDDKEWYVGKAKKKSERELE 181
Query: 110 V----------------GANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNS 153
+ GANL++ NLD +VD++ L + FS FG I T+ KIMRDP G S
Sbjct: 182 LKEQHEQSMQETVDKYQGANLYLKNLDDNVDDEKLRELFSEFGTI-TSCKIMRDPH-GVS 355
Query: 154 RGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTK 195
RG GF+++ + E + A+ MNG+ + + + V+ A KK+ +
Sbjct: 356 RGSGFVAFSTPEEATRALGEMNGKMVDGKPLYVALAQKKEER 481
Score = 47.8 bits (112), Expect = 3e-06
Identities = 28/91 (30%), Positives = 49/91 (53%)
Frame = +2
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
Q + ++ Q A Y+ NLD + +E L ELF + G + + + +D +G GFV F +
Sbjct: 209 QETVDKYQGANLYLKNLDDNVDDEKLRELFSEFGTITSCKIMRD-PHGVSRGSGFVAFST 385
Query: 76 EEDADYAIKVLNMIKLYGKPIRVNKASQDKK 106
E+A A+ +N + GKP+ V A + ++
Sbjct: 386 PEEATRALGEMNGKMVDGKPLYVALAQKKEE 478
>TC8350 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (38%)
Length = 735
Score = 74.3 bits (181), Expect = 3e-14
Identities = 47/152 (30%), Positives = 81/152 (52%), Gaps = 6/152 (3%)
Frame = +1
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
Q A + T ++G+L + E L+ F G V V V +++ T+Q +GYGF+EF S
Sbjct: 271 QQPATAEEVRTLWIGDLQYWMDENYLYTCFAHTGEVTTVKVIRNKQTSQSEGYGFIEFNS 450
Query: 76 EEDADYAIKVLN--MIKLYGKPIRVNKAS---QDKKSLDV-GANLFIGNLDPDVDEKLLY 129
A+ ++ N ++ G+ R+N AS +K+ D +F+G+L DV + LL
Sbjct: 451 RAAAERVLQTFNGAIMPNGGQNFRLNWASLSAGEKRHDDTPDYTIFVGDLAADVTDYLLQ 630
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISY 161
+TF A V K++ D +G ++G+GF+ +
Sbjct: 631 ETFRARYNSVKGAKVVIDRLSGRTKGYGFVRF 726
Score = 41.6 bits (96), Expect = 2e-04
Identities = 26/63 (41%), Positives = 29/63 (45%), Gaps = 4/63 (6%)
Frame = +1
Query: 293 MMQQG---MPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRP-PPPP 348
MMQ G PP M Q +P Q PPP P Q PP + PP A +W P P
Sbjct: 97 MMQPGHGMAPPTMVQQQPAQQYQQPPPLQP---QQPPYAMMPPQAQAPPPHMWAPNVQTP 267
Query: 349 QQQ 351
QQQ
Sbjct: 268 QQQ 276
Score = 39.7 bits (91), Expect = 8e-04
Identities = 22/52 (42%), Positives = 28/52 (53%)
Frame = +1
Query: 289 QGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQ 340
Q QP Q PPP+Q +PP++M PPQ Q PP + P+V QQQ
Sbjct: 139 QQQPAQQYQQPPPLQPQQPPYAMM-----PPQA-QAPPPHMWAPNVQTPQQQ 276
Score = 35.4 bits (80), Expect = 0.016
Identities = 20/63 (31%), Positives = 33/63 (51%), Gaps = 1/63 (1%)
Frame = +1
Query: 15 GQHSAERNQDATAYVGNLDPQISEELLWELF-VQAGPVVNVYVPKDRVTNQHQGYGFVEF 73
G+ + D T +VG+L +++ LL E F + V V DR++ + +GYGFV F
Sbjct: 547 GEKRHDDTPDYTIFVGDLAADVTDYLLQETFRARYNSVKGAKVVIDRLSGRTKGYGFVRF 726
Query: 74 RSE 76
+
Sbjct: 727 ADD 735
Score = 32.7 bits (73), Expect = 0.10
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 2/47 (4%)
Frame = +1
Query: 322 MQGPPRPLPPPSVMAGQ--QQVWRPPPPPQQQGGHPMGYPQNSMPPP 366
M P + PP+++ Q QQ +PPP QQ + M PQ PPP
Sbjct: 97 MMQPGHGMAPPTMVQQQPAQQYQQPPPLQPQQPPYAMMPPQAQAPPP 237
Score = 28.5 bits (62), Expect = 1.9
Identities = 20/51 (39%), Positives = 22/51 (42%), Gaps = 4/51 (7%)
Frame = +1
Query: 241 PAPVPPRPFANGVAPPPIHVIQPP----PPQAPAFQPMQMPPPGQQVWHQQ 287
P V +P PPP+ QPP PPQA A P M P Q QQ
Sbjct: 127 PTMVQQQPAQQYQQPPPLQPQQPPYAMMPPQAQA-PPPHMWAPNVQTPQQQ 276
>TC18542 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding protein, partial
(29%)
Length = 568
Score = 74.3 bits (181), Expect = 3e-14
Identities = 47/133 (35%), Positives = 70/133 (52%), Gaps = 4/133 (3%)
Frame = +2
Query: 66 QGYGFVEFRSEEDADYAIKVLNMIKLYG--KPIRVNKASQDKKSLDVGAN--LFIGNLDP 121
+GYGFVEF S A+ + N + G +P R+ AS D G + +F+G+L P
Sbjct: 17 EGYGFVEFVSHAAAERVLHTYNGTLMPGTEQPFRLIWASSGGGG-DAGPDHSIFVGDLAP 193
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
DV + LL +TF V K++ DP TG S+G+GF+ + + A+ MNG Y +
Sbjct: 194 DVTDFLLQETFRTHYPSVRGAKVVTDPVTGRSKGYGFVKFSDEAQRNRAMTEMNGVYCSS 373
Query: 182 RQITVSYAYKKDT 194
R + +S A K T
Sbjct: 374 RPMRISAATPKKT 412
Score = 48.9 bits (115), Expect = 1e-06
Identities = 28/83 (33%), Positives = 46/83 (54%), Gaps = 1/83 (1%)
Frame = +2
Query: 24 DATAYVGNLDPQISEELLWELFVQAGPVVN-VYVPKDRVTNQHQGYGFVEFRSEEDADYA 82
D + +VG+L P +++ LL E F P V V D VT + +GYGFV+F E + A
Sbjct: 161 DHSIFVGDLAPDVTDFLLQETFRTHYPSVRGAKVVTDPVTGRSKGYGFVKFSDEAQRNRA 340
Query: 83 IKVLNMIKLYGKPIRVNKASQDK 105
+ +N + +P+R++ A+ K
Sbjct: 341 MTEMNGVYCSSRPMRISAATPKK 409
>BP041539
Length = 521
Score = 70.9 bits (172), Expect = 3e-13
Identities = 36/80 (45%), Positives = 51/80 (63%)
Frame = +2
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + + YVG+LD +I++ L++LF Q G VV+V V +D T Q GYG+V F + +D
Sbjct: 269 ANQPMPTSLYVGDLDTEINDSQLYDLFNQIGQVVSVRVCRDLATQQSLGYGYVNFTNPKD 448
Query: 79 ADYAIKVLNMIKLYGKPIRV 98
A A+ VLN L GKPIR+
Sbjct: 449 AATALDVLNFTPLNGKPIRI 508
>BI419822
Length = 380
Score = 69.3 bits (168), Expect = 1e-12
Identities = 38/90 (42%), Positives = 54/90 (59%)
Frame = +2
Query: 107 SLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEA 166
S ++ F+G L D + L FS +G IV + K++ D +TG SRGFGF+++ S +A
Sbjct: 62 SAEIEFRCFVGGLAWATDNEALEKAFSPYGEIVES-KVINDRETGRSRGFGFVTFASEQA 238
Query: 167 SDSAIEAMNGQYLCNRQITVSYAYKKDTKG 196
AIEAMNGQ L R ITV+ A + + G
Sbjct: 239 MKDAIEAMNGQNLDGRNITVNEAQSRGSSG 328
Score = 46.2 bits (108), Expect = 9e-06
Identities = 28/74 (37%), Positives = 39/74 (51%)
Frame = +2
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
+VG L E L + F G +V V DR T + +G+GFV F SE+ AI+ +N
Sbjct: 86 FVGGLAWATDNEALEKAFSPYGEIVESKVINDRETGRSRGFGFVTFASEQAMKDAIEAMN 265
Query: 88 MIKLYGKPIRVNKA 101
L G+ I VN+A
Sbjct: 266 GQNLDGRNITVNEA 307
Score = 26.2 bits (56), Expect = 9.6
Identities = 16/33 (48%), Positives = 17/33 (51%)
Frame = -3
Query: 299 PPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPP 331
PPP PP+ PPPPPP PP PL P
Sbjct: 378 PPP-----PPY----PPPPPP---YLPPPPLLP 316
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 68.6 bits (166), Expect = 2e-12
Identities = 39/93 (41%), Positives = 52/93 (54%)
Frame = +1
Query: 104 DKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDS 163
D DV F+G L D L FS++G I+ + K++ D +TG SRGFGF+++ S
Sbjct: 58 DSAMADVEYRCFVGGLAWTTDSHTLEQAFSSYGEIIDS-KVVNDRETGRSRGFGFVTFTS 234
Query: 164 FEASDSAIEAMNGQYLCNRQITVSYAYKKDTKG 196
EA SAIE MNG L R ITV+ A + G
Sbjct: 235 EEAMRSAIEGMNGNELDGRNITVNEAQARGGGG 333
Score = 58.5 bits (140), Expect = 2e-09
Identities = 43/125 (34%), Positives = 49/125 (38%)
Frame = -1
Query: 245 PPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQ 304
PP P PPP+ +PPP +P + P + PP PPP
Sbjct: 545 PPPP------PPPLP*REPPP--SPPYPPSRRPP*PP-----------------PPP--- 450
Query: 305 FRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMP 364
PP+ PPPPPP PP P PP RPPPPP YP P
Sbjct: 449 --PPY----PPPPPP*PRDPPPPPYPPSR---------RPPPPP---------YPPPPRP 342
Query: 365 PPPPP 369
PPPPP
Sbjct: 341 PPPPP 327
Score = 47.0 bits (110), Expect = 5e-06
Identities = 29/84 (34%), Positives = 44/84 (51%)
Frame = +1
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
SA + + +VG L L + F G +++ V DR T + +G+GFV F SEE
Sbjct: 61 SAMADVEYRCFVGGLAWTTDSHTLEQAFSSYGEIIDSKVVNDRETGRSRGFGFVTFTSEE 240
Query: 78 DADYAIKVLNMIKLYGKPIRVNKA 101
AI+ +N +L G+ I VN+A
Sbjct: 241 AMRSAIEGMNGNELDGRNITVNEA 312
Score = 47.0 bits (110), Expect = 5e-06
Identities = 27/69 (39%), Positives = 34/69 (49%), Gaps = 2/69 (2%)
Frame = -1
Query: 311 MQMPPPPPPQG--MQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPP 368
+Q PPPPPP + PP P PPS ++ PPPPP P P++ PPP P
Sbjct: 554 LQFPPPPPPPLP*REPPPSPPYPPS-----RRPP*PPPPPPPYPPPPPP*PRDPPPPPYP 390
Query: 369 PNHHGMRPP 377
P+ PP
Sbjct: 389 PSRRPPPPP 363
Score = 45.8 bits (107), Expect = 1e-05
Identities = 32/88 (36%), Positives = 39/88 (43%), Gaps = 3/88 (3%)
Frame = -1
Query: 293 MMQQGMPPPMQQFRPPHSMQMPPPPPPQG-MQGPPRPLPPPSVMAGQQQVWRPPPPPQQQ 351
++Q PPP PP + PPP PP + PP P PPP + PPPPP +
Sbjct: 557 LLQFPPPPP-----PPLP*REPPPSPPYPPSRRPP*PPPPPPP-------YPPPPPP*PR 414
Query: 352 GGHPMGYPQNSMP--PPPPPNHHGMRPP 377
P YP + P PP PP PP
Sbjct: 413 DPPPPPYPPSRRPPPPPYPPPPRPPPPP 330
Score = 43.1 bits (100), Expect = 8e-05
Identities = 43/140 (30%), Positives = 49/140 (34%), Gaps = 2/140 (1%)
Frame = -1
Query: 227 PPSLPNAPQANGTIPAPVPPRPFANGVA-PPPIHVIQPPPPQAPAFQPMQMPPPGQQVWH 285
PP P P P P PP P + PPP PPPP PP +
Sbjct: 542 PPPPPPLP*RE---PPPSPPYPPSRRPP*PPPPPPPYPPPP----------PP*PRD--- 411
Query: 286 QQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPP 345
PPP PP+ PPPPP PP P PPP PP
Sbjct: 410 -------------PPP-----PPYPPSRRPPPPPY----PPPPRPPP-----------PP 330
Query: 346 PPPQQQGGHP-MGYPQNSMP 364
PPP+ M P +S+P
Sbjct: 329 PPPRAWASFTVMLRPSSSLP 270
>TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (77%)
Length = 1576
Score = 68.2 bits (165), Expect = 2e-12
Identities = 48/187 (25%), Positives = 87/187 (45%), Gaps = 9/187 (4%)
Frame = +3
Query: 16 QHSAERNQDA--TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEF 73
QH+ D T ++G+L + E L++ F G + V V +++ TNQ +GYGF+EF
Sbjct: 300 QHAQPSTPDEVRTLWIGDLQYWMDENYLYQCFAHTGELATVKVIRNKNTNQSEGYGFLEF 479
Query: 74 RSEEDADYAIKVLN--MIKLYGKPIRVNKAS-----QDKKSLDVGANLFIGNLDPDVDEK 126
S A+ ++ N ++ G+ R+N A+ + + +F+G+L DV +
Sbjct: 480 TSRAGAERILQTYNGTIMPNGGQNFRLNWATFSAGGERRHDDSPDYTIFVGDLAADVTDY 659
Query: 127 LLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITV 186
L + F V K++ D T ++G+GF+ + A+ M G R + +
Sbjct: 660 HLTEVFRTRYNSVKGAKVVIDRLTSRTKGYGFVRFADESEQVRAMTEMQGVVCSTRPMRI 839
Query: 187 SYAYKKD 193
A K+
Sbjct: 840 GPAANKN 860
Score = 63.5 bits (153), Expect = 5e-11
Identities = 47/203 (23%), Positives = 88/203 (43%), Gaps = 22/203 (10%)
Frame = +3
Query: 15 GQHSAERNQDATAYVGNLDPQISEELLWELF-VQAGPVVNVYVPKDRVTNQHQGYGFVEF 73
G+ + + D T +VG+L +++ L E+F + V V DR+T++ +GYGFV F
Sbjct: 585 GERRHDDSPDYTIFVGDLAADVTDYHLTEVFRTRYNSVKGAKVVIDRLTSRTKGYGFVRF 764
Query: 74 RSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVG---------------------A 112
E + A+ + + +P+R+ A+ G
Sbjct: 765 ADESEQVRAMTEMQGVVCSTRPMRIGPAANKNLGTGTGTQPKASYQNSQGPQNENDPNNT 944
Query: 113 NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIE 172
+F+GNLDP+V + L F +G +V + KI + + GF+ + ++ A+
Sbjct: 945 TIFVGNLDPNVTDDHLRQVFGLYGDLV-HVKIPQ------GKRCGFVQFADRSCAEEALR 1103
Query: 173 AMNGQYLCNRQITVSYAYKKDTK 195
+NG L + + +S+ K
Sbjct: 1104VLNGTLLGGQNVRLSWGRSPSNK 1172
Score = 48.5 bits (114), Expect = 2e-06
Identities = 28/88 (31%), Positives = 49/88 (54%)
Frame = +3
Query: 12 NLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFV 71
N G + + T +VGNLDP ++++ L ++F G +V+V +P Q + GFV
Sbjct: 903 NSQGPQNENDPNNTTIFVGNLDPNVTDDHLRQVFGLYGDLVHVKIP------QGKRCGFV 1064
Query: 72 EFRSEEDADYAIKVLNMIKLYGKPIRVN 99
+F A+ A++VLN L G+ +R++
Sbjct: 1065QFADRSCAEEALRVLNGTLLGGQNVRLS 1148
Score = 37.7 bits (86), Expect = 0.003
Identities = 27/76 (35%), Positives = 29/76 (37%), Gaps = 9/76 (11%)
Frame = +3
Query: 276 MPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPP------RPL 329
M PP Q QQQ QP M MPP Q +PP PPQ PP +P
Sbjct: 144 MAPPQQPNQQYQQQQQPYMM--MPPQQHQPQPPTMWAPSAAQPPQQQHAPPQQHQHAQPS 317
Query: 330 PPPSVMA---GQQQVW 342
P V G Q W
Sbjct: 318 TPDEVRTLWIGDLQYW 365
Score = 30.8 bits (68), Expect = 0.39
Identities = 21/61 (34%), Positives = 23/61 (37%), Gaps = 2/61 (3%)
Frame = +3
Query: 297 GMPPPMQQFRPPHSMQMP--PPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGH 354
GM PP Q + Q P PP Q PP P + QQQ PP Q Q
Sbjct: 141 GMAPPQQPNQQYQQQQQPYMMMPPQQHQPQPPTMWAPSAAQPPQQQ---HAPPQQHQHAQ 311
Query: 355 P 355
P
Sbjct: 312 P 314
Score = 28.1 bits (61), Expect = 2.5
Identities = 19/62 (30%), Positives = 23/62 (36%), Gaps = 4/62 (6%)
Frame = +3
Query: 321 GMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPP----PPPNHHGMRP 376
G G P P QQQ + PP Q Q P + ++ PP PP H
Sbjct: 132 GGPGMAPPQQPNQQYQQQQQPYMMMPPQQHQPQPPTMWAPSAAQPPQQQHAPPQQHQHAQ 311
Query: 377 PS 378
PS
Sbjct: 312 PS 317
>TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structural protein
pp78/81, partial (5%)
Length = 710
Score = 64.7 bits (156), Expect = 2e-11
Identities = 54/180 (30%), Positives = 65/180 (36%), Gaps = 9/180 (5%)
Frame = +3
Query: 208 AASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAP-----VPPRPFANGVAPPPIHVIQ 262
A ++P Q + PS N PQ+ +P P +PP+ PPP Q
Sbjct: 18 AGTDPVQQAASAPLQSQQQFPSPVNLPQSIPVVPPPNAPPQLPPQQGLPPSFPPPNQFSQ 197
Query: 263 PPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMP----PPMQQFRPPHSMQMPPPPP 318
P P P PP Q QQ QP+ Q P PP QQ++ Q P P
Sbjct: 198 TQIPAVPQRDPYFPPPVQSQETPNQQYQQPLAPQPHPQPGAPPHQQYQQTPHPQYSQPAP 377
Query: 319 PQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPPS 378
PQ P PP + V PP P Q YP N PP P PPS
Sbjct: 378 PQHQPPIPSINPPQLQSSLGHHVEEPPYVPSQT------YPPNLRQPPSQPPSGPPAPPS 539
Score = 58.2 bits (139), Expect = 2e-09
Identities = 43/155 (27%), Positives = 57/155 (36%), Gaps = 4/155 (2%)
Frame = +3
Query: 227 PPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQ 286
PP + + N P+ P+P APP Q P PQ P Q PP +
Sbjct: 237 PPPVQSQETPNQQYQQPLAPQPHPQPGAPPHQQYQQTPHPQYSQPAPPQHQPPIPSINPP 416
Query: 287 QQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPP 346
Q Q PP + P +++ PP PP G P PP G PP
Sbjct: 417 QLQSSLGHHVEEPPYVPSQTYPPNLRQPPSQPPSG-----PPAPPSQQFYGTAPHAYEPP 581
Query: 347 PPQQQGGHPMGYPQNSMPPPP----PPNHHGMRPP 377
P + G+ GY +S P P +G +PP
Sbjct: 582 PSRSGSGYSSGYGSSSGPAEQYRYGGPPQYGGKPP 686
>AV427333
Length = 387
Score = 63.5 bits (153), Expect = 5e-11
Identities = 52/163 (31%), Positives = 63/163 (37%), Gaps = 6/163 (3%)
Frame = +3
Query: 216 KSRPHTLFASGPPSL--PNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQP 273
KSR + AS P+ P+ P P P P+ PP H PPP + F
Sbjct: 21 KSRSKSKMASTQPNFYFPHFP--------PPPSHPY--NPPTPPSHPFNPPPTPSHYF-- 164
Query: 274 MQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRP----L 329
+ PPP + +G P PPP PP P PPP ++ PP P
Sbjct: 165 -KSPPPPSHSFAPPPRGHP-----SPPPSS---PPPPSSHPFTPPPPHVRPPPSPHHPIT 317
Query: 330 PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHH 372
PPP V RPPPPP +PP P PNHH
Sbjct: 318 PPPHV--------RPPPPP--------------LPPSPSPNHH 380
Score = 52.4 bits (124), Expect = 1e-07
Identities = 39/125 (31%), Positives = 43/125 (34%), Gaps = 2/125 (1%)
Frame = +3
Query: 255 PPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMP 314
PPP H PP P + F P P P F+ P
Sbjct: 84 PPPSHPYNPPTPPSHPFNPP------------------------PTPSHYFKSP------ 173
Query: 315 PPPPPQGMQGPPR--PLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHH 372
PPP PPR P PPPS PPPP HP P + PPP P HH
Sbjct: 174 -PPPSHSFAPPPRGHPSPPPS----------SPPPPSS---HPFTPPPPHVRPPPSP-HH 308
Query: 373 GMRPP 377
+ PP
Sbjct: 309 PITPP 323
Score = 47.8 bits (112), Expect = 3e-06
Identities = 31/96 (32%), Positives = 36/96 (37%), Gaps = 5/96 (5%)
Frame = +3
Query: 287 QQQGQPMMQQGMPPPMQQFRPPHSMQMPPPP-----PPQGMQGPPRPLPPPSVMAGQQQV 341
Q+Q + + M F PH PPPP PP P P P PS
Sbjct: 9 QEQDKSRSKSKMASTQPNFYFPH---FPPPPSHPYNPPTPPSHPFNPPPTPS-----HYF 164
Query: 342 WRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPP 377
PPPP P G+P PPPP+ H PP
Sbjct: 165 KSPPPPSHSFAPPPRGHPSPPPSSPPPPSSHPFTPP 272
Score = 44.3 bits (103), Expect = 3e-05
Identities = 25/73 (34%), Positives = 32/73 (43%), Gaps = 3/73 (4%)
Frame = +3
Query: 210 SNPTAQKSRPHTLFASGPP---SLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPP 266
S+P P F S PP S P+ + + P PP P ++ PPP HV PP P
Sbjct: 123 SHPFNPPPTPSHYFKSPPPPSHSFAPPPRGHPSPPPSSPPPPSSHPFTPPPPHVRPPPSP 302
Query: 267 QAPAFQPMQMPPP 279
P P + PP
Sbjct: 303 HHPITPPPHVRPP 341
Score = 40.0 bits (92), Expect = 6e-04
Identities = 25/75 (33%), Positives = 33/75 (43%), Gaps = 2/75 (2%)
Frame = +3
Query: 201 TPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAP--VPPRPFANGVAPPPI 258
TP+ + P+ + P S PPS P P ++ P P V P P + PP
Sbjct: 147 TPSHYFKSPPPPSHSFAPPPRGHPSPPPSSPPPPSSHPFTPPPPHVRPPPSPHHPITPPP 326
Query: 259 HVIQPPPPQAPAFQP 273
HV PPPP P+ P
Sbjct: 327 HVRPPPPPLPPSPSP 371
>TC9150 similar to UP|Q9M6E4 (Q9M6E4) Poly(A)-binding protein (Fragment),
partial (65%)
Length = 1191
Score = 63.5 bits (153), Expect = 5e-11
Identities = 65/243 (26%), Positives = 94/243 (37%), Gaps = 2/243 (0%)
Frame = +3
Query: 139 VTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGER 198
+T+ KIMRDP TG SRG GF+++ + E + A+ MNG+ + + + V+ A +K+ + R
Sbjct: 33 ITSFKIMRDP-TGISRGSGFVAFSTPEEASRALGEMNGKMISGKPLYVALAQRKEDRRAR 209
Query: 199 HGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPI 258
+ RP + S P +P P AP + + G PP +
Sbjct: 210 -----------LQAQFSQMRPVAISPSVAPRMPIYPPG-----APGLGQQYLYGQGPPAM 341
Query: 259 HVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQG--QPMMQQGMPPPMQQFRPPHSMQMPPP 316
PQ F Q PG + Q PM+QQG R +Q P
Sbjct: 342 ------IPQQAGFGYQQQLMPGMRPGGAPMQSFFVPMVQQGQQGQRPGGRRAGHVQQPQQ 503
Query: 317 PPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRP 376
P P Q M + +V+R PP Q G G M P +R
Sbjct: 504 PMPMMQQ----------QMLPRGRVYRYPPGRNMQDGPMQGVAGGMMSVPYDMGGLPIRD 653
Query: 377 PSG 379
P G
Sbjct: 654 PVG 662
Score = 28.5 bits (62), Expect = 1.9
Identities = 17/55 (30%), Positives = 29/55 (51%)
Frame = +3
Query: 49 GPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQ 103
G + + + +D T +G GFV F + E+A A+ +N + GKP+ V A +
Sbjct: 27 GTITSFKIMRDP-TGISRGSGFVAFSTPEEASRALGEMNGKMISGKPLYVALAQR 188
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 63.2 bits (152), Expect = 7e-11
Identities = 45/149 (30%), Positives = 53/149 (35%), Gaps = 5/149 (3%)
Frame = +2
Query: 226 GPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWH 285
G S N+P + P P P+P A +PPP PP +P P PPP Q
Sbjct: 179 GAQSPSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPP--PASTPPPAQS--- 343
Query: 286 QQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPP 345
P + PPP+Q PP PPP P PP PPP+ PP
Sbjct: 344 ----SPPPVSS--PPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAA--------TPP 481
Query: 346 P-----PPQQQGGHPMGYPQNSMPPPPPP 369
P P P S P P P
Sbjct: 482 PALTPVPATSPAPAPAKVKSKSPAPAPAP 568
Score = 52.4 bits (124), Expect = 1e-07
Identities = 39/121 (32%), Positives = 42/121 (34%), Gaps = 2/121 (1%)
Frame = +2
Query: 260 VIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPP--P 317
VI Q+P+ P P P QP Q PPP Q PP PP P
Sbjct: 164 VIASVGAQSPSNSPTTSPAPPTPT-----TPQPPAAQSSPPPAQSSPPPVQSSPPPASTP 328
Query: 318 PPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPP 377
PP PP PPP V PPP P P S PP PP PP
Sbjct: 329 PPAQSSPPPVSSPPP--------VQSTPPPAPASTPPPASPPPFSPPPATPPPPAATPPP 484
Query: 378 S 378
+
Sbjct: 485 A 487
Score = 48.1 bits (113), Expect = 2e-06
Identities = 39/134 (29%), Positives = 45/134 (33%)
Frame = +2
Query: 201 TPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHV 260
TP +S P AQ S P S PP P A + P P P + PPP
Sbjct: 236 TPQPPAAQSSPPPAQSSPPPV--QSSPPPASTPPPAQSSPPPVSSPPPVQS--TPPPAPA 403
Query: 261 IQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQ 320
PPP P F P PP PPP P S P P P +
Sbjct: 404 STPPPASPPPFSPPPATPP--------------PPAATPPPALTPVPATS---PAPAPAK 532
Query: 321 GMQGPPRPLPPPSV 334
P P P P++
Sbjct: 533 VKSKSPAPAPAPAL 574
Score = 45.4 bits (106), Expect = 2e-05
Identities = 37/135 (27%), Positives = 48/135 (35%)
Frame = +2
Query: 208 AASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQ 267
A++ P AQ S P +S PP P A + P P P PF+ A PP PPP
Sbjct: 317 ASTPPPAQSSPPPV--SSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATPPPAL 490
Query: 268 APAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPR 327
P P P P + + P P P S+ P P P
Sbjct: 491 TPV--PATSPAPA--------PAKVKSKSPAPAPAPALAPVVSLSPSEAPGPSLSSLSPA 640
Query: 328 PLPPPSVMAGQQQVW 342
P S +G ++ W
Sbjct: 641 LSPAASDDSGVEKSW 685
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 63.2 bits (152), Expect = 7e-11
Identities = 54/183 (29%), Positives = 67/183 (36%), Gaps = 14/183 (7%)
Frame = +1
Query: 201 TPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHV 260
TP + + A P S P F +G + ++ G + P P P PPP
Sbjct: 58 TPPKSLKALPPPPPPPSAP---FIAGSAIISKVSKSVGDVSLPCSPSP------PPP--- 201
Query: 261 IQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPP--HSMQMPPPPP 318
QPPPP P P PP G+ +P P Q PP + + PP P
Sbjct: 202 -QPPPPSLPPTTP---PPASVIAGSAIMSGEASFPCSLPSPPQPPPPPPKYGVTTIPPSP 369
Query: 319 PQGMQGPPR-PLPPPSVMAGQQQVWRPP-----------PPPQQQGGHPMGYPQNSMPPP 366
P M R PLPPPS PP P P Q P Y ++PPP
Sbjct: 370 PLLMSATDRAPLPPPSTHLTSLTTPSPPSSRTTITQTPLPSPPQPPSPPPKYGVATIPPP 549
Query: 367 PPP 369
P P
Sbjct: 550 PLP 558
Score = 49.3 bits (116), Expect = 1e-06
Identities = 55/187 (29%), Positives = 62/187 (32%), Gaps = 43/187 (22%)
Frame = +1
Query: 226 GPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQ---------------PPPPQAPA 270
G SLP +P P P PP P PPP VI P PPQ P
Sbjct: 160 GDVSLPCSPSP----PPPQPPPPSLPPTTPPPASVIAGSAIMSGEASFPCSLPSPPQPP- 324
Query: 271 FQPMQMPPPGQQVWHQQQQGQPMMQQG-----MPPPMQQFR------PPHSM-------- 311
PPP + P++ +PPP PP S
Sbjct: 325 ------PPPPKYGVTTIPPSPPLLMSATDRAPLPPPSTHLTSLTTPSPPSSRTTITQTPL 486
Query: 312 ----QMPPPPPPQGMQG-PPRPLPPPSVMAGQQQVWRPPPP----PQQQGGHPMGYPQNS 362
Q P PPP G+ PP PLP S+ R P P P P P S
Sbjct: 487 PSPPQPPSPPPKYGVATIPPPPLPMSSID-------RTPLPLLSTPLTSLDAPPPTPTLS 645
Query: 363 MPPPPPP 369
PPPPPP
Sbjct: 646 PPPPPPP 666
Score = 47.0 bits (110), Expect = 5e-06
Identities = 44/155 (28%), Positives = 58/155 (37%), Gaps = 13/155 (8%)
Frame = +1
Query: 227 PPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQ 286
PPS P + +P P P P + PPP PPPP AP G + +
Sbjct: 4 PPSTPMSSADRAPLPPPSTP-PKSLKALPPP-----PPPPSAPFIA-------GSAIISK 144
Query: 287 QQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVW---- 342
+ + P PP PP PPP + PP PP SV+AG +
Sbjct: 145 VSKSVGDVSLPCSPS-----PP-----PPQPPPPSL--PPTTPPPASVIAGSAIMSGEAS 288
Query: 343 ---------RPPPPPQQQGGHPMGYPQNSMPPPPP 368
+PPPPP + G ++PP PP
Sbjct: 289 FPCSLPSPPQPPPPPPKYG-------VTTIPPSPP 372
Score = 41.2 bits (95), Expect = 3e-04
Identities = 33/112 (29%), Positives = 36/112 (31%), Gaps = 7/112 (6%)
Frame = +1
Query: 227 PPSLPNAPQANGTIPAPVPPRPFANGVAPPP-------IHVIQPPPPQAPAFQPMQMPPP 279
PPS P A P P P + P P P PPQ P+ PPP
Sbjct: 358 PPSPPLLMSATDRAPLPPPSTHLTSLTTPSPPSSRTTITQTPLPSPPQPPS------PPP 519
Query: 280 GQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPP 331
V PM P P S+ PPP P PP P PP
Sbjct: 520 KYGVATIPPPPLPMSSIDRTPLPLLSTPLTSLDAPPPTPTLSPPPPPPPTPP 675
Score = 40.4 bits (93), Expect = 5e-04
Identities = 31/111 (27%), Positives = 38/111 (33%), Gaps = 37/111 (33%)
Frame = +1
Query: 298 MPPPMQQFRPPHSMQ-MPPPPPPQG------------------------MQGPPRPLPPP 332
+PPP PP S++ +PPPPPP PP P PPP
Sbjct: 43 LPPPST---PPKSLKALPPPPPPPSAPFIAGSAIISKVSKSVGDVSLPCSPSPPPPQPPP 213
Query: 333 SVMAGQQQVWRPP--PPPQQ----------QGGHPMGYPQNSMPPPPPPNH 371
+ PP PPP + P P PPPPPP +
Sbjct: 214 PSL--------PPTTPPPASVIAGSAIMSGEASFPCSLPSPPQPPPPPPKY 342
Score = 35.4 bits (80), Expect = 0.016
Identities = 37/136 (27%), Positives = 46/136 (33%), Gaps = 27/136 (19%)
Frame = +1
Query: 224 ASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQ-----PPPPQAPAFQPMQMP- 277
AS P SLP+ PQ P P PP+ + P P ++ P PP + + P
Sbjct: 283 ASFPCSLPSPPQ-----PPPPPPKYGVTTIPPSPPLLMSATDRAPLPPPSTHLTSLTTPS 447
Query: 278 PPGQQVWHQQQQGQPMMQQGMPPPMQQFR--PPHSMQMP-------------------PP 316
PP + Q Q PPP PP + M PP
Sbjct: 448 PPSSRTTITQTPLPSPPQPPSPPPKYGVATIPPPPLPMSSIDRTPLPLLSTPLTSLDAPP 627
Query: 317 PPPQGMQGPPRPLPPP 332
P P PP P PP
Sbjct: 628 PTPTLSPPPPPPPTPP 675
Score = 33.5 bits (75), Expect = 0.060
Identities = 22/75 (29%), Positives = 30/75 (39%), Gaps = 15/75 (20%)
Frame = +1
Query: 210 SNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGV---------------A 254
S P+++ + T S PP P+ P G P PP P ++ A
Sbjct: 445 SPPSSRTTITQTPLPS-PPQPPSPPPKYGVATIPPPPLPMSSIDRTPLPLLSTPLTSLDA 621
Query: 255 PPPIHVIQPPPPQAP 269
PPP + PPPP P
Sbjct: 622 PPPTPTLSPPPPPPP 666
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,229,510
Number of Sequences: 28460
Number of extensions: 334011
Number of successful extensions: 13573
Number of sequences better than 10.0: 1170
Number of HSP's better than 10.0 without gapping: 3667
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8076
length of query: 379
length of database: 4,897,600
effective HSP length: 92
effective length of query: 287
effective length of database: 2,279,280
effective search space: 654153360
effective search space used: 654153360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC139882.5


