

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139709.6 + phase: 0
(138 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
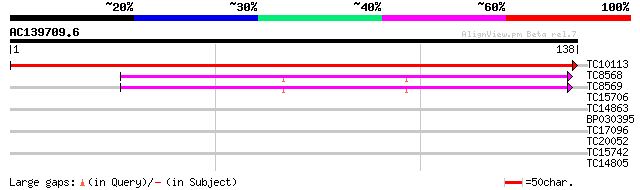
Score E
Sequences producing significant alignments: (bits) Value
TC10113 UP|RR11_LOTJA (Q9BBQ3) Chloroplast 30S ribosomal protein... 248 3e-67
TC8568 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein... 64 7e-12
TC8569 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein... 64 7e-12
TC15706 weakly similar to UP|P93408 (P93408) Mitochondrial ribos... 35 0.006
TC14863 30 0.20
BP030395 27 1.7
TC17096 similar to PIR|C53394|C53394 ribosomal protein L12.C pre... 27 1.7
TC20052 similar to UP|Q94AF2 (Q94AF2) AT5g19150/T24G5_50, partia... 26 2.8
TC15742 25 6.3
TC14805 similar to UP|RP3A_ARATH (Q39211) DNA-directed RNA polym... 25 6.3
>TC10113 UP|RR11_LOTJA (Q9BBQ3) Chloroplast 30S ribosomal protein S11,
complete
Length = 879
Score = 248 bits (633), Expect = 3e-67
Identities = 123/138 (89%), Positives = 129/138 (93%)
Frame = +1
Query: 1 MAKSIPKIGSRKNGRIGSRKHPRKIPKGVIYVQASFNNTIVTVTDVRGRVISWSSAGSCG 60
MAK IPKIGSRKN R GSRKH RKIPKG+I+VQASFNNTIVTVTDVRGRVISWSSAG+CG
Sbjct: 463 MAKPIPKIGSRKNARSGSRKHLRKIPKGIIHVQASFNNTIVTVTDVRGRVISWSSAGTCG 642
Query: 61 FKGTRRGTPFAAQTAAANAIRTVVDQGMQRAVVIIKGPGLGRDAALRAIARSGILLRFIR 120
FKGTRRGTPFAAQTAA NAIRTV DQGMQRA V+IKGPGLGRDAALRAI RSGILL FIR
Sbjct: 643 FKGTRRGTPFAAQTAAGNAIRTVADQGMQRAEVMIKGPGLGRDAALRAIRRSGILLNFIR 822
Query: 121 DVTPIPHNGCRAPKKRRV 138
DVTP+PHNGCR+PKKRRV
Sbjct: 823 DVTPMPHNGCRSPKKRRV 876
>TC8568 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein S14,
complete
Length = 771
Score = 64.3 bits (155), Expect = 7e-12
Identities = 44/119 (36%), Positives = 59/119 (48%), Gaps = 9/119 (7%)
Frame = +2
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 149 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVATRCKEL 328
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G AALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 329 GITALHIRLRATGGNKTKTPGPGAQAALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 505
>TC8569 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein S14,
complete
Length = 633
Score = 64.3 bits (155), Expect = 7e-12
Identities = 44/119 (36%), Positives = 59/119 (48%), Gaps = 9/119 (7%)
Frame = +1
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 148 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVATRCKEL 327
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G AALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 328 GITALHIKLRATGGNKTKTPGPGAQAALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 504
>TC15706 weakly similar to UP|P93408 (P93408) Mitochondrial ribosomal
protein S11 (Nuclear encoded), partial (31%)
Length = 600
Score = 34.7 bits (78), Expect = 0.006
Identities = 29/102 (28%), Positives = 45/102 (43%), Gaps = 10/102 (9%)
Frame = +2
Query: 47 RGRVISWSSAGSC-GFKGTRRGTPFAAQTAAANAIRTVVDQGMQRAVVIIKGPGLGRDAA 105
+G + SAGS K ++ + +AA+ A R G++ V+ + G R
Sbjct: 2 KGNIKLSGSAGSLKDMKSGQKLSRYAAEATAEVVGRRARGLGLKSVVMKVNGYTHFRRKR 181
Query: 106 LRAIA-RSGIL--------LRFIRDVTPIPHNGCRAPKKRRV 138
++ R G + F+ D T HNGCR PKKRR+
Sbjct: 182 QAILSWREGFTDSKADKNPIVFVEDTTRRAHNGCRLPKKRRI 307
>TC14863
Length = 1803
Score = 29.6 bits (65), Expect = 0.20
Identities = 18/61 (29%), Positives = 29/61 (47%)
Frame = +3
Query: 12 KNGRIGSRKHPRKIPKGVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTRRGTPFA 71
++ R+G + P ++ +F+NT + DVRG V + SC + RG FA
Sbjct: 183 RDHRVGGLNNAMLQPSSCTLLRKNFHNTHL---DVRGGVSLHENGRSCAIRSELRGPVFA 353
Query: 72 A 72
A
Sbjct: 354 A 356
>BP030395
Length = 461
Score = 26.6 bits (57), Expect = 1.7
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = -1
Query: 118 FIRDVTPIPHNGCRAPKKRRV 138
F+ D T HNGCR P++ V
Sbjct: 377 FVEDTTRRAHNGCRLPQETDV 315
>TC17096 similar to PIR|C53394|C53394 ribosomal protein L12.C precursor,
chloroplast - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (74%)
Length = 677
Score = 26.6 bits (57), Expect = 1.7
Identities = 16/39 (41%), Positives = 24/39 (61%)
Frame = +2
Query: 71 AAQTAAANAIRTVVDQGMQRAVVIIKGPGLGRDAALRAI 109
AA AAA+A VV++ + VVI + P R AA++A+
Sbjct: 266 AAAPAAADAAVAVVEEQTEFDVVIQEVPSNARIAAIKAV 382
>TC20052 similar to UP|Q94AF2 (Q94AF2) AT5g19150/T24G5_50, partial (35%)
Length = 556
Score = 25.8 bits (55), Expect = 2.8
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = +1
Query: 84 VDQGMQRAVVIIKGPGLGRDAAL 106
VD+ ++R ++ GPGLGRD L
Sbjct: 469 VDKWLERFDCLVIGPGLGRDPFL 537
>TC15742
Length = 1208
Score = 24.6 bits (52), Expect = 6.3
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = +3
Query: 74 TAAANAIRTVVDQGMQRAVVI 94
TA NAIR DQG++R V I
Sbjct: 528 TANINAIRAASDQGVKRFVYI 590
>TC14805 similar to UP|RP3A_ARATH (Q39211) DNA-directed RNA polymerase II 36
kDa polypeptide A (RNA polymerase II subunit 3) ,
complete
Length = 1273
Score = 24.6 bits (52), Expect = 6.3
Identities = 13/33 (39%), Positives = 20/33 (60%)
Frame = +2
Query: 82 TVVDQGMQRAVVIIKGPGLGRDAALRAIARSGI 114
T D + + ++I+K G++ LRAIAR GI
Sbjct: 461 TSSDSAVNKGIIIVK-LRRGQELRLRAIARKGI 556
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.138 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,136,595
Number of Sequences: 28460
Number of extensions: 22368
Number of successful extensions: 129
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 126
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 127
length of query: 138
length of database: 4,897,600
effective HSP length: 81
effective length of query: 57
effective length of database: 2,592,340
effective search space: 147763380
effective search space used: 147763380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 50 (23.9 bits)
Medicago: description of AC139709.6


