

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139709.19 + phase: 0
(353 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
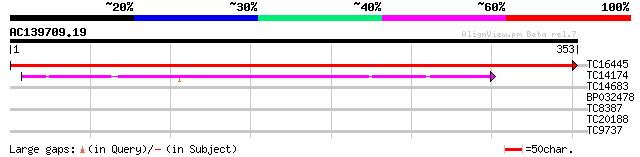
TC16445 UP|AAQ67339 (AAQ67339) Photosystem II thylakoid membrane... 717 0.0
TC14174 UP|PSBC_LOTJA (Q9BBT1) Photosystem II 44 kDa reaction ce... 163 4e-41
TC14683 homologue to GB|AAA84544.1|343028|PEACPHD psbA peptide {... 38 0.003
BP032478 30 0.47
TC8387 similar to UP|BCA2_ARATH (Q9M439) Branched-chain amino ac... 29 1.0
TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 27 4.0
TC9737 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, pa... 26 8.8
>TC16445 UP|AAQ67339 (AAQ67339) Photosystem II thylakoid membrane protein,
complete
Length = 1162
Score = 717 bits (1852), Expect = 0.0
Identities = 351/353 (99%), Positives = 352/353 (99%)
Frame = +2
Query: 1 MTAILERRDSENLWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI 60
MTAILERR+SENLW RFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI
Sbjct: 56 MTAILERRESENLWGRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI 235
Query: 61 DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL 120
DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL
Sbjct: 236 DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL 415
Query: 121 LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF 180
LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF
Sbjct: 416 LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF 595
Query: 181 NFMIVFQAEHNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFG 240
NFMIVFQAEHNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFG
Sbjct: 596 NFMIVFQAEHNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFG 775
Query: 241 QEEETYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGF 300
QEEETYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGF
Sbjct: 776 QEEETYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGF 955
Query: 301 NFNQSVVDSQGRVINTWADIINRANLGMEVMHERNAHNFPLDLAAVEAPSING 353
NFNQSVVDSQGRVINTWADIINRANLGMEVMHERNAHNFPLDLAAVEAPSING
Sbjct: 956 NFNQSVVDSQGRVINTWADIINRANLGMEVMHERNAHNFPLDLAAVEAPSING 1114
>TC14174 UP|PSBC_LOTJA (Q9BBT1) Photosystem II 44 kDa reaction center
protein (P6 protein) (CP43), complete
Length = 3406
Score = 163 bits (413), Expect = 4e-41
Identities = 90/298 (30%), Positives = 160/298 (53%), Gaps = 3/298 (1%)
Frame = +3
Query: 8 RDSENLWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDIDGIREPV 67
+D +L+ +W+ + +++GW G+L+ P A + G+
Sbjct: 531 KDENDLFDIMDDWLRR-DRFVFVGWSGLLLFPCAYFALGGWFTGTTFVTSWYTHGL---- 695
Query: 68 SGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDE---WLYNGGPYELIVLHFLLGVA 124
+ S L G N ++ A+ + ++ +W + + W GG + + LH G+
Sbjct: 696 ASSYLEGCNFLTAAVSTPANSLAHSLLLLWGPEAQGDFTRWCQLGGLWTFVALHGAFGLI 875
Query: 125 CYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTFNFMI 184
+M R++EL+ + +RP+ A+A+S P+A +VFLIYP+GQ + G++ F F++
Sbjct: 876 GFMLRQFELARSVQLRPYNAIAFSGPIAVFVSVFLIYPLGQSGWFFAPSFGVAAIFRFIL 1055
Query: 185 VFQAEHNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFGQEEE 244
FQ HN ++PFHM+GVAGV G +L A+HG+ V ++L E + + + Q EE
Sbjct: 1056FFQGFHNWTLNPFHMMGVAGVLGAALLCAIHGATVENTLF-EDGDGANTFRAFNPTQAEE 1232
Query: 245 TYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGFNF 302
TY++V A+ ++ ++ +F+N R LHFF+ PV G+W +ALG+ +A NL ++F
Sbjct: 1233TYSMVTANRFWSQIF--GVAFSNKRWLHFFMLFVPVTGLWMSALGVVGLALNLRAYDF 1400
>TC14683 homologue to GB|AAA84544.1|343028|PEACPHD psbA peptide {Pisum
sativum;} , partial (67%)
Length = 619
Score = 37.7 bits (86), Expect = 0.003
Identities = 17/19 (89%), Positives = 18/19 (94%)
Frame = +2
Query: 335 NAHNFPLDLAAVEAPSING 353
+AHN PLDLAAVEAPSING
Sbjct: 212 DAHNSPLDLAAVEAPSING 268
>BP032478
Length = 509
Score = 30.4 bits (67), Expect = 0.47
Identities = 17/53 (32%), Positives = 28/53 (52%)
Frame = -2
Query: 225 RETTENESANEGYRFGQEEETYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAA 277
RETT+ ES + + +E + +N +GY L Y +SR LH F+++
Sbjct: 433 RETTQEES*QKWIQEEEEIKGFNWTTVYGYVRSLCGCYNDLLSSRCLHNFVSS 275
>TC8387 similar to UP|BCA2_ARATH (Q9M439) Branched-chain amino acid
aminotransferase 2, chloroplast precursor (Atbcat-2) ,
partial (84%)
Length = 1883
Score = 29.3 bits (64), Expect = 1.0
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = +3
Query: 32 WFGVLMIPTLLTATSVFIIAF 52
WF LMI TLLTATSV+ +
Sbjct: 1704 WFDGLMISTLLTATSVYFCIY 1766
>TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (11%)
Length = 494
Score = 27.3 bits (59), Expect = 4.0
Identities = 13/48 (27%), Positives = 22/48 (45%)
Frame = -2
Query: 246 YNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTM 293
Y ++A + +L +A F N +F + AW V WF L + +
Sbjct: 175 YALLARRPFRTKLTSIFALFQNPFQNYFHVQAWHVHNFWFDCLHLXVI 32
>TC9737 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, partial
(11%)
Length = 563
Score = 26.2 bits (56), Expect = 8.8
Identities = 11/44 (25%), Positives = 20/44 (45%)
Frame = +3
Query: 13 LWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAP 56
LW ++C + S++++ GW + L +FI I P
Sbjct: 279 LWQKWCKFFRSSKHQFLKGWMRHASLFN*LQVQKIFITVEIVIP 410
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,596,265
Number of Sequences: 28460
Number of extensions: 93304
Number of successful extensions: 581
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 578
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 580
length of query: 353
length of database: 4,897,600
effective HSP length: 91
effective length of query: 262
effective length of database: 2,307,740
effective search space: 604627880
effective search space used: 604627880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC139709.19


