

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137839.10 - phase: 0
(502 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
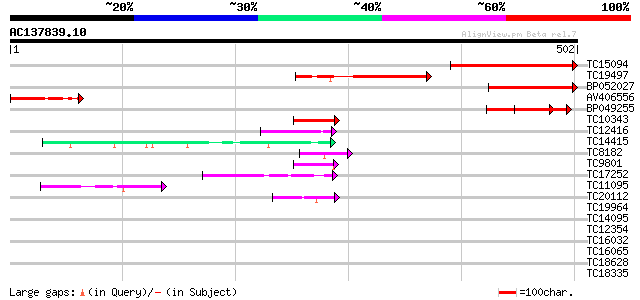
Score E
Sequences producing significant alignments: (bits) Value
TC15094 similar to UP|Q8LA02 (Q8LA02) Contains similarity to RNA... 203 6e-53
TC19497 weakly similar to UP|Q8LA02 (Q8LA02) Contains similarity... 132 2e-31
BP052027 127 4e-30
AV406556 65 2e-11
BP049255 59 2e-09
TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (... 52 2e-07
TC12416 similar to GB|AAH56066.1|33417132|BC056066 sop-prov prot... 49 2e-06
TC14415 similar to UP|O04237 (O04237) Transcription factor, part... 46 2e-05
TC8182 weakly similar to UP|Q93VA8 (Q93VA8) At1g76010/T4O12_22, ... 43 1e-04
TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like... 42 2e-04
TC17252 similar to UP|O04237 (O04237) Transcription factor, part... 42 3e-04
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 41 5e-04
TC20112 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich... 40 7e-04
TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%) 40 0.001
TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein... 40 0.001
TC12354 similar to UP|O23702 (O23702) Dehydrogenase, partial (25%) 40 0.001
TC16032 similar to BAC87886 (BAC87886) PGL-3, partial (4%) 39 0.002
TC16065 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich... 39 0.003
TC18628 weakly similar to PIR|T09527|T09527 glycine-rich protein... 38 0.004
TC18335 similar to UP|Q7XJ13 (Q7XJ13) Chloroplast nucleoid DNA-b... 37 0.010
>TC15094 similar to UP|Q8LA02 (Q8LA02) Contains similarity to RNA-binding
protein from Arabidopsis thaliana gi|2129727 and
contains RNA recognition PF|00076 domain, partial (21%)
Length = 599
Score = 203 bits (516), Expect = 6e-53
Identities = 97/112 (86%), Positives = 105/112 (93%)
Frame = +2
Query: 391 RDALEKMKPFLMTYEGIRSQEEWEEVIEELMQRVPLLKKIVDHYSGPDRVTAKKQQEELE 450
RDALEKMKPFLM YEGI+SQEEWEE +EE M RVPL+KKIVDHY GPDRVTAKKQQ ELE
Sbjct: 2 RDALEKMKPFLMVYEGIQSQEEWEEAMEETMARVPLMKKIVDHYCGPDRVTAKKQQGELE 181
Query: 451 RVAKTLPTSAPSSVKEFTNRAVVSLQSNPGWGFDKKCQFMDKLVFEVSQHHK 502
RVAKTLPTSAPSSVK+FT+RAV+SLQSNPGWGFDKKC FMDKLV+EVSQH+K
Sbjct: 182 RVAKTLPTSAPSSVKQFTDRAVISLQSNPGWGFDKKCHFMDKLVWEVSQHYK 337
>TC19497 weakly similar to UP|Q8LA02 (Q8LA02) Contains similarity to
RNA-binding protein from Arabidopsis thaliana gi|2129727
and contains RNA recognition PF|00076 domain, partial
(11%)
Length = 436
Score = 132 bits (331), Expect = 2e-31
Identities = 74/122 (60%), Positives = 84/122 (68%), Gaps = 2/122 (1%)
Frame = +2
Query: 254 DGDGSGRGRGRGRGRDVYARGRGDRGRGG--RGGRGDGRGGFKRYGDDRTSHQDIARSNA 311
+G G GRGRGRGRGR RGRG RGR G R GR QD S+A
Sbjct: 113 EGGGLGRGRGRGRGR---GRGRGFRGRDGEERSGRS----------------QDTDISHA 235
Query: 312 DGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDINCAIEFEP 371
GLY+GD+ADGEKLAKKLGPEIM+Q+TE +EE+ RVLPSPL+DE VEAMD+N AIE EP
Sbjct: 236 GGLYLGDDADGEKLAKKLGPEIMNQLTEGFEEMASRVLPSPLEDELVEAMDVNYAIECEP 415
Query: 372 EY 373
EY
Sbjct: 416 EY 421
Score = 28.9 bits (63), Expect = 2.0
Identities = 13/20 (65%), Positives = 13/20 (65%)
Frame = +2
Query: 30 GFGGGGGDGRGGGRGRGGSG 49
G G G G GRG GRGRG G
Sbjct: 122 GLGRGRGRGRGRGRGRGFRG 181
Score = 28.1 bits (61), Expect = 3.5
Identities = 11/12 (91%), Positives = 11/12 (91%)
Frame = +2
Query: 115 GTGRGRGRGRGF 126
G GRGRGRGRGF
Sbjct: 140 GRGRGRGRGRGF 175
Score = 28.1 bits (61), Expect = 3.5
Identities = 12/15 (80%), Positives = 12/15 (80%)
Frame = +2
Query: 32 GGGGGDGRGGGRGRG 46
GGG G GRG GRGRG
Sbjct: 116 GGGLGRGRGRGRGRG 160
>BP052027
Length = 472
Score = 127 bits (319), Expect = 4e-30
Identities = 61/78 (78%), Positives = 69/78 (88%)
Frame = -1
Query: 425 PLLKKIVDHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSLQSNPGWGFD 484
PL+KKIVDHY GPDRVT KK ELERVAKTLP SAPSSVK FT+RAV+SLQS+PGWGFD
Sbjct: 472 PLMKKIVDHYCGPDRVTPKKPPGELERVAKTLPPSAPSSVKPFTDRAVISLQSHPGWGFD 293
Query: 485 KKCQFMDKLVFEVSQHHK 502
+KC FMDKLV+EVSQ ++
Sbjct: 292 QKCPFMDKLVWEVSQPYQ 239
>AV406556
Length = 344
Score = 65.5 bits (158), Expect = 2e-11
Identities = 39/65 (60%), Positives = 44/65 (67%)
Frame = +2
Query: 1 MRGTIGVRLQNSTISNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAA 60
+RGTIGVR+Q ++ISNA R+ LVPFSTSS F GGG GRGG G GG FNF A
Sbjct: 86 VRGTIGVRIQGASISNAGRRALVPFSTSSNF--GGGRGRGGRPGGGG----PFNF----A 235
Query: 61 PGNPN 65
PG N
Sbjct: 236 PGKTN 250
Score = 32.0 bits (71), Expect = 0.24
Identities = 20/57 (35%), Positives = 24/57 (42%)
Frame = +3
Query: 37 DGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGHGRG 93
D GGGR G + P +PN D T+ IPPG+G G GRG
Sbjct: 204 DPEGGGRSTSLQG--------RLIPASPNL---------DTTEPSIPPGSGHGDGRG 323
>BP049255
Length = 468
Score = 58.5 bits (140), Expect = 2e-09
Identities = 33/61 (54%), Positives = 38/61 (62%), Gaps = 1/61 (1%)
Frame = -1
Query: 423 RVPLLKKIVDHYSGPDRVTAKKQQEELER-VAKTLPTSAPSSVKEFTNRAVVSLQSNPGW 481
RVPL+KKIVDHY GPDRVTAKKQQ E R + K P + +FT+RAV P
Sbjct: 465 RVPLMKKIVDHYCGPDRVTAKKQQREN*RELLKLFPQALHLP*SQFTDRAVYLSSEQPRL 286
Query: 482 G 482
G
Sbjct: 285 G 283
Score = 47.8 bits (112), Expect = 4e-06
Identities = 26/51 (50%), Positives = 34/51 (65%), Gaps = 1/51 (1%)
Frame = -3
Query: 448 ELERVAKTLPTSAPSSVKE-FTNRAVVSLQSNPGWGFDKKCQFMDKLVFEV 497
ELERVAKTLPTSAPSS+K + R + ++ ++KC FMDKL+ V
Sbjct: 388 ELERVAKTLPTSAPSSLKPIY*PRCLSLFRATQAGELNQKCHFMDKLILGV 236
>TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (Fibrillarin
2) (AtFib2), partial (85%)
Length = 1116
Score = 52.4 bits (124), Expect = 2e-07
Identities = 26/41 (63%), Positives = 26/41 (63%)
Frame = +1
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGG 292
R G G G GRG GRG ARG G G GGRGGRG GRGG
Sbjct: 88 RGRGRGDGGGRGGGRGTPFKARGGGRGGGGGRGGRGGGRGG 210
Score = 38.9 bits (89), Expect = 0.002
Identities = 26/44 (59%), Positives = 26/44 (59%), Gaps = 3/44 (6%)
Frame = +1
Query: 260 RGRGRGRGRDVYARGRGDRGRG--GRGGRGDGRG-GFKRYGDDR 300
RGRG G G RGRGDRGRG GGRG GRG FK G R
Sbjct: 46 RGRGGGGG----FRGRGDRGRGRGDGGGRGGGRGTPFKARGGGR 165
Score = 35.8 bits (81), Expect = 0.017
Identities = 24/41 (58%), Positives = 24/41 (58%), Gaps = 7/41 (17%)
Frame = +1
Query: 259 GRGRGRG-RGRDVYARGRGDRG--RGGRG----GRGDGRGG 292
GRG G G RGR RGRGD G GGRG RG GRGG
Sbjct: 49 GRGGGGGFRGRGDRGRGRGDGGGRGGGRGTPFKARGGGRGG 171
Score = 32.3 bits (72), Expect = 0.18
Identities = 20/47 (42%), Positives = 21/47 (44%), Gaps = 13/47 (27%)
Frame = +1
Query: 252 RDDGDGSGRGRGRGR-------------GRDVYARGRGDRGRGGRGG 285
R GDG GRG GRG GR GRG RG G +GG
Sbjct: 94 RGRGDGGGRGGGRGTPFKARGGGRGGGGGRGGRGGGRGGRGGGMKGG 234
Score = 32.0 bits (71), Expect = 0.24
Identities = 16/28 (57%), Positives = 16/28 (57%), Gaps = 2/28 (7%)
Frame = +1
Query: 24 PFSTSSGFGGGGGD--GRGGGRGRGGSG 49
PF G GGGG GRGGGRG G G
Sbjct: 139 PFKARGGGRGGGGGRGGRGGGRGGRGGG 222
Score = 27.7 bits (60), Expect = 4.6
Identities = 15/29 (51%), Positives = 15/29 (51%), Gaps = 9/29 (31%)
Frame = +1
Query: 30 GFGGGGGDGRGGGR---------GRGGSG 49
G G G G GRGGGR GRGG G
Sbjct: 91 GRGRGDGGGRGGGRGTPFKARGGGRGGGG 177
Score = 27.3 bits (59), Expect = 6.0
Identities = 23/64 (35%), Positives = 23/64 (35%), Gaps = 1/64 (1%)
Frame = +1
Query: 32 GGGGGDGRGG-GRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGH 90
GGGG GRG GRGRG G G G P GRG
Sbjct: 58 GGGGFRGRGDRGRGRGDGG------GRGGGRGTPFKARGGGRGGGGGR-------GGRGG 198
Query: 91 GRGG 94
GRGG
Sbjct: 199 GRGG 210
>TC12416 similar to GB|AAH56066.1|33417132|BC056066 sop-prov protein
{Xenopus laevis;} , partial (32%)
Length = 416
Score = 48.5 bits (114), Expect = 2e-06
Identities = 31/68 (45%), Positives = 36/68 (52%), Gaps = 1/68 (1%)
Frame = +3
Query: 223 VNRHIRVRQTPADAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGR-DVYARGRGDRGRG 281
+NRH+ + T + R GDG GRGRG GRGR + RGRG RGRG
Sbjct: 33 INRHLLCQPTIN*LKMSGDGRGASGEPSAEGFGDGFGRGRGEGRGRGEGRGRGRG-RGRG 209
Query: 282 GRGGRGDG 289
RG RGDG
Sbjct: 210 RRGRRGDG 233
Score = 33.9 bits (76), Expect = 0.064
Identities = 34/115 (29%), Positives = 47/115 (40%), Gaps = 16/115 (13%)
Frame = +3
Query: 261 GRGRGRGRDVYARGRGD---RGRGGRGGRGDGRGGFKRYGDDRTSHQDIARSNADGLYVG 317
G GRG + A G GD RGRG GRG+GRG + G R + G
Sbjct: 84 GDGRGASGEPSAEGFGDGFGRGRGEGRGRGEGRGRGRGRGRGRRGRR------------G 227
Query: 318 DNADGE-----KLAKKLGPEIMDQITEAYE--------EIIERVLPSPLQDEYVE 359
D E KL + + ++ + E Y EII+ L L+DE ++
Sbjct: 228 DGDKKEWVPVTKLGRLVKAGMIKSLEEIYVFSLAIKEFEIIDHFLGDSLKDEVMK 392
Score = 31.2 bits (69), Expect = 0.41
Identities = 13/20 (65%), Positives = 15/20 (75%)
Frame = +3
Query: 27 TSSGFGGGGGDGRGGGRGRG 46
++ GFG G G GRG GRGRG
Sbjct: 114 SAEGFGDGFGRGRGEGRGRG 173
Score = 30.0 bits (66), Expect = 0.92
Identities = 13/22 (59%), Positives = 14/22 (63%)
Frame = +3
Query: 25 FSTSSGFGGGGGDGRGGGRGRG 46
F G G G G+GRG GRGRG
Sbjct: 138 FGRGRGEGRGRGEGRGRGRGRG 203
Score = 30.0 bits (66), Expect = 0.92
Identities = 12/17 (70%), Positives = 13/17 (75%)
Frame = +3
Query: 30 GFGGGGGDGRGGGRGRG 46
GFG G G+GRG G GRG
Sbjct: 135 GFGRGRGEGRGRGEGRG 185
Score = 28.9 bits (63), Expect = 2.0
Identities = 17/41 (41%), Positives = 18/41 (43%)
Frame = +3
Query: 85 GAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRG 125
G GRG GRG + G GRGRGRGRG
Sbjct: 141 GRGRGEGRG------------------RGEGRGRGRGRGRG 209
Score = 28.1 bits (61), Expect = 3.5
Identities = 13/20 (65%), Positives = 13/20 (65%)
Frame = +3
Query: 30 GFGGGGGDGRGGGRGRGGSG 49
G G G G GRG GRGRG G
Sbjct: 159 GRGRGEGRGRGRGRGRGRRG 218
Score = 28.1 bits (61), Expect = 3.5
Identities = 13/22 (59%), Positives = 13/22 (59%)
Frame = +3
Query: 25 FSTSSGFGGGGGDGRGGGRGRG 46
F G G G G GRG GRGRG
Sbjct: 126 FGDGFGRGRGEGRGRGEGRGRG 191
>TC14415 similar to UP|O04237 (O04237) Transcription factor, partial (83%)
Length = 1666
Score = 45.8 bits (107), Expect = 2e-05
Identities = 72/292 (24%), Positives = 105/292 (35%), Gaps = 33/292 (11%)
Frame = +2
Query: 30 GFGGGGGDGRGGGRGRGGSGTVT---FNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGA 86
G GG G GRG GRGRG + + +F AP N P +E + P G
Sbjct: 449 GRGGARGAGRGFGRGRGFNRDFSNDENSFPASGAPDNLGPFEGDSEKASERRGYGAPRGP 628
Query: 87 GRGHG---RGGTVPDFPSFSFSSFMSSIQQPGTGRG-----RGRGRG------------F 126
RG G RGG + + GTGRG G GRG
Sbjct: 629 YRGGGGGRRGGFSNGEADEEGRPRRAFERHSGTGRGSGFKREGAGRGNWGTQSDEIAQVT 808
Query: 127 DPLPPQFENDSVPKKPVFIKREDNVSQTDA--NDFSPPKNPVFTRSEDVRPVEPIDLSGD 184
D + + E + +KP D ++ A N+ P++ T E + +E
Sbjct: 809 DEVANETEKNLSDEKPAVEDVADGNKESSANENEEKEPEDKEMTLEEYEKVLE------- 967
Query: 185 SESDNRFVMTVPKVLPGGGRGRGKPLEEAAQEAP-QAPVVNRHI-------RVRQTPADA 236
+ R + K GR ++E P N I + ++ A
Sbjct: 968 ---EKRKALQAQKT-----EGRKVDIKEFESMQPLSCKKDNNDIFAKLGSDKDKRKEAFE 1123
Query: 237 ESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGD 288
+ + + +N F++ + GRGRG RGRG RG GG RG+
Sbjct: 1124KEEKAKKSVSINEFLKPAEGETYYPSGRGRG-----RGRGSRGGGGGSFRGN 1264
Score = 37.0 bits (84), Expect = 0.008
Identities = 80/343 (23%), Positives = 124/343 (35%), Gaps = 41/343 (11%)
Frame = +2
Query: 170 SEDVRPVEPIDLSGDSESDNRFVMTVPKVLPGGGRGRGKPLEEAA-----QEAPQAPVVN 224
S + + P DL GD D ++ ++ P + A ++A A + +
Sbjct: 212 SRKMATMNPFDLLGDDAEDPSQLIAAEQLKAAAAAATAPPKKAAGAQDQGKQAKPAQMPS 391
Query: 225 RHIRVRQTPADAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRG--RDVY-------ARGR 275
+ + Q DA R +P +R R G+GRG GRGRG RD A G
Sbjct: 392 KPLPPAQAVRDA------RNEP-SRGGRGGARGAGRGFGRGRGFNRDFSNDENSFPASGA 550
Query: 276 GD---------------RGRGG-----RGGRGDGRGGFKRYGDDRTSHQDIARSNADGLY 315
D RG G RGG G RGGF D A G
Sbjct: 551 PDNLGPFEGDSEKASERRGYGAPRGPYRGGGGGRRGGFSNGEADEEGRPRRAFERHSGTG 730
Query: 316 VGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDINCAIEFEPEYAV 375
G E + D+I + +E+ L DE A+E +
Sbjct: 731 RGSGFKREGAGRGNWGTQSDEIAQVTDEVANET-EKNLSDE-------KPAVEDVADGNK 886
Query: 376 EFDNPDIDEKEPIALRDALEKMKPFL-MTYEGIRSQEEWEEVIE----ELMQRVPLLKKI 430
E + +EKEP LE+ + L + +++Q+ ++ E MQ + K
Sbjct: 887 ESSANENEEKEPEDKEMTLEEYEKVLEEKRKALQAQKTEGRKVDIKEFESMQPLSCKKDN 1066
Query: 431 VDHYS--GPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRA 471
D ++ G D+ K+ E+ E+ K++ S+ EF A
Sbjct: 1067NDIFAKLGSDKDKRKEAFEKEEKAKKSV------SINEFLKPA 1177
>TC8182 weakly similar to UP|Q93VA8 (Q93VA8) At1g76010/T4O12_22, partial
(13%)
Length = 611
Score = 43.1 bits (100), Expect = 1e-04
Identities = 30/68 (44%), Positives = 31/68 (45%), Gaps = 21/68 (30%)
Frame = +3
Query: 257 GSGRGRGRGRGRDVYARGRGD--------------------RGRG-GRGGRGDGRGGFKR 295
G GRG GRGRGR RGRG RGRG GRGGRG GRG
Sbjct: 81 GRGRGFGRGRGRGFRGRGRGGYGAQPVGYYDYGEYDAPPAPRGRGRGRGGRGRGRGRNAS 260
Query: 296 YGDDRTSH 303
+G SH
Sbjct: 261 WGVAA*SH 284
Score = 39.3 bits (90), Expect = 0.002
Identities = 35/109 (32%), Positives = 39/109 (35%), Gaps = 9/109 (8%)
Frame = +3
Query: 25 FSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPG---------NPNPTPNVNESKP 75
F + G GGGDG GGRG GG G FG G P + +
Sbjct: 15 FYNNGGMEYGGGDGWDGGRGYGGRGR---GFGRGRGRGFRGRGRGGYGAQPVGYYDYGEY 185
Query: 76 DATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGR 124
DA +P GRG GRG GRGRGRGR
Sbjct: 186 DAPPAP----RGRGRGRG-----------------------GRGRGRGR 251
Score = 35.8 bits (81), Expect = 0.017
Identities = 27/63 (42%), Positives = 30/63 (46%), Gaps = 1/63 (1%)
Frame = +3
Query: 250 FVRDDGDGSGRGRGRGRGRDVYARGRG-DRGRGGRGGRGDGRGGFKRYGDDRTSHQDIAR 308
F + G G G G GR RGRG RGRG RG RG GRGG YG + D
Sbjct: 15 FYNNGGMEYGGGDGWDGGRGYGGRGRGFGRGRG-RGFRGRGRGG---YGAQPVGYYDYGE 182
Query: 309 SNA 311
+A
Sbjct: 183 YDA 191
Score = 28.5 bits (62), Expect = 2.7
Identities = 15/38 (39%), Positives = 18/38 (46%)
Frame = +3
Query: 245 QPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGG 282
QP+ + + D RGRGRGR RGRG G
Sbjct: 153 QPVGYYDYGEYDAPPAPRGRGRGRGGRGRGRGRNASWG 266
>TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like, complete
Length = 981
Score = 42.4 bits (98), Expect = 2e-04
Identities = 23/45 (51%), Positives = 27/45 (59%), Gaps = 5/45 (11%)
Frame = +2
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGG-----RGDGRG 291
R++G G +G GRG D A+G G RGRGG GG RG GRG
Sbjct: 404 REEGAGGRPAKGPGRGFDDGAKGPGGRGRGGLGGKPGGSRGAGRG 538
Score = 31.2 bits (69), Expect = 0.41
Identities = 25/72 (34%), Positives = 26/72 (35%), Gaps = 16/72 (22%)
Frame = +2
Query: 70 VNESKPDATDSPIPPGAGRGHGRGGTVPDF-------PSFSFSSFMSSI---------QQ 113
V E TD PPG GRG GRGG P F +
Sbjct: 332 VQEETKSRTDRK-PPGVGRGRGRGGREEGAGGRPAKGPGRGFDDGAKGPGGRGRGGLGGK 508
Query: 114 PGTGRGRGRGRG 125
PG RG GRGRG
Sbjct: 509 PGGSRGAGRGRG 544
>TC17252 similar to UP|O04237 (O04237) Transcription factor, partial (21%)
Length = 492
Score = 41.6 bits (96), Expect = 3e-04
Identities = 33/120 (27%), Positives = 49/120 (40%)
Frame = +1
Query: 171 EDVRPVEPIDLSGDSESDNRFVMTVPKVLPGGGRGRGKPLEEAAQEAPQAPVVNRHIRVR 230
+++ + P DL GD D ++ + + QE +A + +
Sbjct: 166 DEMATMNPFDLLGDDAEDPSQLIVAEQQKAAAAAAAAAAAPKKGQEHGKAGPAAQ----Q 333
Query: 231 QTPADAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGR 290
PA S +P Q + R +++ GRG GRG GR GRGGRGG G GR
Sbjct: 334 NKPAQLPSKPLPPTQAV-RESKNEASRGGRGGGRGGGRGF--------GRGGRGGSGFGR 486
Score = 32.0 bits (71), Expect = 0.24
Identities = 16/25 (64%), Positives = 18/25 (72%), Gaps = 1/25 (4%)
Frame = +1
Query: 26 STSSGFGGGGGDGRGGGR-GRGGSG 49
++ G GGG G GRG GR GRGGSG
Sbjct: 403 ASRGGRGGGRGGGRGFGRGGRGGSG 477
Score = 28.1 bits (61), Expect = 3.5
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = +1
Query: 275 RGDRGRGGRGGRGDGRGGFKRYG 297
R + RGGRG GRGG + +G
Sbjct: 385 RESKNEASRGGRGGGRGGGRGFG 453
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 40.8 bits (94), Expect = 5e-04
Identities = 37/117 (31%), Positives = 48/117 (40%), Gaps = 5/117 (4%)
Frame = +1
Query: 28 SSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAG 87
S G GGGGG GRGGGR GG G E PG+ A + + G+G
Sbjct: 298 SRGGGGGGGGGRGGGRSGGGGGGSDMKCYECGEPGH------------FARECRMRGGSG 441
Query: 88 RGHGRGGTVPDF---PSFSFSSFMSSIQQPGTGRGRGRGRG-FDPLPPQFE-NDSVP 139
R R + P F PS+ S+ + P GRG + PP + + VP
Sbjct: 442 RRRSR--SPPRFRRSPSYGRRSYSPRGRSPRRRSVSPHGRGSYSRSPPPYRAREEVP 606
>TC20112 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich
glycoprotein DZ-HRGP precursor, partial (11%)
Length = 523
Score = 40.4 bits (93), Expect = 7e-04
Identities = 28/62 (45%), Positives = 30/62 (48%), Gaps = 2/62 (3%)
Frame = +2
Query: 233 PADAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDV--YARGRGDRGRGGRGGRGDGR 290
PA + N P P F G G G GRG R D Y+ G G RGRGG RG GR
Sbjct: 299 PAAHANPNPPFVPPSGGFSVGRGGGFG-GRGGDRKYDTGRYSGGGGGRGRGGGSIRGGGR 475
Query: 291 GG 292
GG
Sbjct: 476 GG 481
Score = 36.6 bits (83), Expect = 0.010
Identities = 23/40 (57%), Positives = 23/40 (57%)
Frame = +2
Query: 253 DDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGG 292
D G SG G GRGRG G RG GGRGGRG G GG
Sbjct: 404 DTGRYSGGGGGRGRG------GGSIRG-GGRGGRGGGFGG 502
Score = 32.0 bits (71), Expect = 0.24
Identities = 14/23 (60%), Positives = 15/23 (64%)
Frame = +2
Query: 25 FSTSSGFGGGGGDGRGGGRGRGG 47
+ T GGGGG GRGGG RGG
Sbjct: 401 YDTGRYSGGGGGRGRGGGSIRGG 469
Score = 28.9 bits (63), Expect = 2.0
Identities = 13/20 (65%), Positives = 13/20 (65%)
Frame = +2
Query: 30 GFGGGGGDGRGGGRGRGGSG 49
G G GGG RGGGRG G G
Sbjct: 434 GRGRGGGSIRGGGRGGRGGG 493
>TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%)
Length = 562
Score = 39.7 bits (91), Expect = 0.001
Identities = 20/35 (57%), Positives = 20/35 (57%)
Frame = +2
Query: 253 DDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRG 287
D G G GRGR GR GRG G GGRGGRG
Sbjct: 194 DSGRGGGRGRFGSGGRGGDRGGRGGSGGGGRGGRG 298
Score = 36.6 bits (83), Expect = 0.010
Identities = 22/41 (53%), Positives = 22/41 (53%)
Frame = +2
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGG 292
R G SG GRG GR GRG G GGRGG GRGG
Sbjct: 149 RSGGGRSGGGRG-GRFDSGRGGGRGRFGSGGRGGDRGGRGG 268
Score = 35.4 bits (80), Expect = 0.022
Identities = 26/47 (55%), Positives = 26/47 (55%), Gaps = 6/47 (12%)
Frame = +2
Query: 252 RDDGDGSGR---GRGRGRGRDVYARGRGDRG--RGGRGGR-GDGRGG 292
R G GR GRG GRGR G G RG RGGRGG G GRGG
Sbjct: 164 RSGGGRGGRFDSGRGGGRGRF----GSGGRGGDRGGRGGSGGGGRGG 292
Score = 30.8 bits (68), Expect = 0.54
Identities = 22/47 (46%), Positives = 22/47 (46%)
Frame = +2
Query: 254 DGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
D GSG G GR GRG R GRGG G GR G G DR
Sbjct: 128 DNQGSGGRSGGGRS----GGGRGGRFDSGRGG-GRGRFGSGGRGGDR 253
Score = 30.8 bits (68), Expect = 0.54
Identities = 14/23 (60%), Positives = 15/23 (64%)
Frame = +2
Query: 27 TSSGFGGGGGDGRGGGRGRGGSG 49
+ G GG GRGGGRGR GSG
Sbjct: 167 SGGGRGGRFDSGRGGGRGRFGSG 235
Score = 27.7 bits (60), Expect = 4.6
Identities = 17/26 (65%), Positives = 17/26 (65%), Gaps = 5/26 (19%)
Frame = +2
Query: 29 SGFGGGGGD----GRGGGR-GRGGSG 49
SG GGG G GRGG R GRGGSG
Sbjct: 197 SGRGGGRGRFGSGGRGGDRGGRGGSG 274
>TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein S2,
partial (88%)
Length = 1134
Score = 39.7 bits (91), Expect = 0.001
Identities = 34/112 (30%), Positives = 47/112 (41%)
Frame = +3
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDRTSHQDIARSNA 311
R GD G GRG G GRGDRGRG R GRGG +R G R +
Sbjct: 63 RGGGDRGGFGRGFG--------GRGDRGRGDR-----GRGGRRRTGGRRDEEEKWVPVTK 203
Query: 312 DGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDI 363
G V D + L + + +II+ ++ L+DE ++ M +
Sbjct: 204 LGRLVRDGK-----IRSLEQIYLHSLPIKEHQIIDMLVGPTLKDEVMKIMPV 344
>TC12354 similar to UP|O23702 (O23702) Dehydrogenase, partial (25%)
Length = 488
Score = 39.7 bits (91), Expect = 0.001
Identities = 20/41 (48%), Positives = 23/41 (55%)
Frame = -1
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGG 292
R G G RG+ R RGR RGRG +G G G R +GR G
Sbjct: 281 RGGGRGQQRGKSRRRGRRSRGRGRGGKGGDGAGRRVEGRRG 159
Score = 30.8 bits (68), Expect = 0.54
Identities = 18/35 (51%), Positives = 19/35 (53%), Gaps = 3/35 (8%)
Frame = -1
Query: 260 RGRGRGRGRD---VYARGRGDRGRGGRGGRGDGRG 291
R RG GRG+ RGR RGRG G GDG G
Sbjct: 287 RERGGGRGQQRGKSRRRGRRSRGRGRGGKGGDGAG 183
>TC16032 similar to BAC87886 (BAC87886) PGL-3, partial (4%)
Length = 631
Score = 39.3 bits (90), Expect = 0.002
Identities = 32/86 (37%), Positives = 40/86 (46%), Gaps = 21/86 (24%)
Frame = +3
Query: 229 VRQTPADAESDNVPRRQPMNRFVRDDGDG-----SGRGRGRGRG------RDVYARGRGD 277
+ Q DA+ + PRR N+ G G +GRGRG RG R+ YA G+
Sbjct: 147 LNQPDVDAKDQHYPRRGYQNQRGGRGGGGRRGYSNGRGRGGSRGGYYPNGRNQYADQPGN 326
Query: 278 ---------RGRGGRGGRG-DGRGGF 293
RGRGGRG G +G GGF
Sbjct: 327 YYPRNNYNNRGRGGRGSYGNNGSGGF 404
Score = 32.7 bits (73), Expect = 0.14
Identities = 20/45 (44%), Positives = 21/45 (46%), Gaps = 3/45 (6%)
Frame = +3
Query: 30 GFGGGGGDGRGGGRGRGGSGTVTFNFGEKA---APGNPNPTPNVN 71
G GGGG G GRGRGGS + G PGN P N N
Sbjct: 216 GRGGGGRRGYSNGRGRGGSRGGYYPNGRNQYADQPGNYYPRNNYN 350
Score = 28.1 bits (61), Expect = 3.5
Identities = 17/41 (41%), Positives = 20/41 (48%), Gaps = 6/41 (14%)
Frame = +3
Query: 271 YARGRGDRGRGGR------GGRGDGRGGFKRYGDDRTSHQD 305
Y RG RG GGR GRG RGG+ Y + R + D
Sbjct: 198 YQNQRGGRGGGGRRGYSNGRGRGGSRGGY--YPNGRNQYAD 314
>TC16065 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich
glycoprotein DZ-HRGP precursor, partial (20%)
Length = 608
Score = 38.5 bits (88), Expect = 0.003
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = -1
Query: 32 GGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKP 75
GGGGG G GGGRG GG G+ P + P P ++++ P
Sbjct: 155 GGGGGYGGGGGRGGGGMGSGRGVSPSHGGPPSRPPAPELHQATP 24
Score = 33.1 bits (74), Expect = 0.11
Identities = 35/118 (29%), Positives = 43/118 (35%), Gaps = 4/118 (3%)
Frame = -1
Query: 15 SNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGG--SGTVTFNFGEKAAPGNPNPTPNVNE 72
S++T ++ P ++ GGGG G GGRG GG G +G PG
Sbjct: 425 SHSTERSAPPPQQAAAAPGGGGPGPQGGRGYGGPPQGGRGGGYGGGGGPGYGGGGRG-GH 249
Query: 73 SKPDATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPG--TGRGRGRGRGFDP 128
P P GR GRGG +S G G G G GRG P
Sbjct: 248 GVPQQQYGGPPEYQGR--GRGGPSQQGGRGGYSGGGGGYGGGGGRGGGGMGSGRGVSP 81
Score = 30.8 bits (68), Expect = 0.54
Identities = 17/32 (53%), Positives = 18/32 (56%), Gaps = 1/32 (3%)
Frame = -1
Query: 262 RGRGRGRDVYARGRGDRGRGGRG-GRGDGRGG 292
+GRGRG GRG GG G G G GRGG
Sbjct: 209 QGRGRGGPSQQGGRGGYSGGGGGYGGGGGRGG 114
Score = 28.1 bits (61), Expect = 3.5
Identities = 19/46 (41%), Positives = 22/46 (47%), Gaps = 5/46 (10%)
Frame = -1
Query: 257 GSGRGRGRGRGRDVYA-----RGRGDRGRGGRGGRGDGRGGFKRYG 297
G GRG G G + Y +GRG G +GGRG GG YG
Sbjct: 269 GGGRG-GHGVPQQQYGGPPEYQGRGRGGPSQQGGRGGYSGGGGGYG 135
>TC18628 weakly similar to PIR|T09527|T09527 glycine-rich protein 1 -
chickpea {Cicer arietinum;} , partial (32%)
Length = 424
Score = 37.7 bits (86), Expect = 0.004
Identities = 16/24 (66%), Positives = 18/24 (74%)
Frame = +3
Query: 274 GRGDRGRGGRGGRGDGRGGFKRYG 297
GRG GRGG GGRG G GG+ R+G
Sbjct: 15 GRGHGGRGGGGGRGGGGGGYCRHG 86
Score = 32.3 bits (72), Expect = 0.18
Identities = 14/20 (70%), Positives = 14/20 (70%)
Frame = +3
Query: 30 GFGGGGGDGRGGGRGRGGSG 49
G GG G GRGGG GRGG G
Sbjct: 6 GGGGRGHGGRGGGGGRGGGG 65
>TC18335 similar to UP|Q7XJ13 (Q7XJ13) Chloroplast nucleoid DNA-binding
protein-like protein, partial (19%)
Length = 588
Score = 36.6 bits (83), Expect = 0.010
Identities = 22/57 (38%), Positives = 28/57 (48%), Gaps = 6/57 (10%)
Frame = +3
Query: 245 QPMNRFVRDDGDGS------GRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKR 295
+P+N + +DG+ S GRGRGRGRG Y G + G G G GG R
Sbjct: 405 KPLNEY-EEDGEXSPXMPXXGRGRGRGRGXXFYNNGGMEYGGGDGWDXGXXYGGXSR 572
Score = 29.3 bits (64), Expect = 1.6
Identities = 18/52 (34%), Positives = 24/52 (45%), Gaps = 9/52 (17%)
Frame = +3
Query: 51 VTFNFGEK----AAPGNPNPTPN-----VNESKPDATDSPIPPGAGRGHGRG 93
+T N +K ++ G P P +NE + D SP P GRG GRG
Sbjct: 327 ITINLSKKELDTSSTGYQPPLPADQVKPLNEYEEDGEXSPXMPXXGRGRGRG 482
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.137 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,354,595
Number of Sequences: 28460
Number of extensions: 107293
Number of successful extensions: 2837
Number of sequences better than 10.0: 266
Number of HSP's better than 10.0 without gapping: 1485
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2333
length of query: 502
length of database: 4,897,600
effective HSP length: 94
effective length of query: 408
effective length of database: 2,222,360
effective search space: 906722880
effective search space used: 906722880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC137839.10


