

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.7 + phase: 0
(123 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
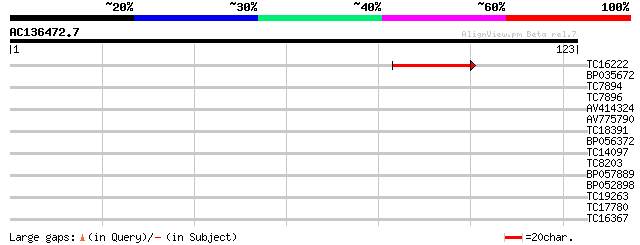
Sequences producing significant alignments: (bits) Value
TC16222 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase (Dihyd... 42 3e-05
BP035672 28 0.34
TC7894 weakly similar to UP|Q93YY0 (Q93YY0) 68 kDa protein HP68,... 28 0.34
TC7896 homologue to UP|Q8GTE3 (Q8GTE3) Ribosomal protein S3a, pa... 28 0.34
AV414324 26 1.7
AV775790 25 3.8
TC18391 similar to UP|Q9SSW9 (Q9SSW9) Small GTP-binding protein ... 25 3.8
BP056372 25 3.8
TC14097 homologue to UP|GCSP_PEA (P26969) Glycine dehydrogenase ... 25 5.0
TC8203 similar to UP|Q8L9C3 (Q8L9C3) Copia-like retroelement pol... 24 6.5
BP057889 24 6.5
BP052898 24 6.5
TC19263 similar to UP|Q8L8A0 (Q8L8A0) Type IIB calcium ATPase MC... 24 6.5
TC17780 similar to GB|AAL11601.1|15983466|AF424607 At1g33990/F12... 24 8.5
TC16367 similar to UP|Q8W3L8 (Q8W3L8) Xyloglucan endo-transglyco... 24 8.5
>TC16222 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase
(Dihydropyrimidine amidohydrolase) , partial (27%)
Length = 570
Score = 42.0 bits (97), Expect = 3e-05
Identities = 18/18 (100%), Positives = 18/18 (100%)
Frame = +2
Query: 84 QKAFGIDDFRKIPNGVNG 101
QKAFGIDDFRKIPNGVNG
Sbjct: 32 QKAFGIDDFRKIPNGVNG 85
>BP035672
Length = 485
Score = 28.5 bits (62), Expect = 0.34
Identities = 16/30 (53%), Positives = 17/30 (56%)
Frame = +2
Query: 4 DAMEEVAKARKSGQRVIGEPVLSGLALDDS 33
DA+ KSG RVI P LSGL LD S
Sbjct: 77 DALSVTRFDTKSGMRVITSPNLSGLFLDGS 166
>TC7894 weakly similar to UP|Q93YY0 (Q93YY0) 68 kDa protein HP68, partial
(9%)
Length = 1599
Score = 28.5 bits (62), Expect = 0.34
Identities = 16/30 (53%), Positives = 17/30 (56%)
Frame = -1
Query: 4 DAMEEVAKARKSGQRVIGEPVLSGLALDDS 33
DA+ KSG RVI P LSGL LD S
Sbjct: 1134 DALSVTRFDTKSGMRVITSPNLSGLFLDGS 1045
>TC7896 homologue to UP|Q8GTE3 (Q8GTE3) Ribosomal protein S3a, partial
(25%)
Length = 592
Score = 28.5 bits (62), Expect = 0.34
Identities = 16/30 (53%), Positives = 17/30 (56%)
Frame = -1
Query: 4 DAMEEVAKARKSGQRVIGEPVLSGLALDDS 33
DA+ KSG RVI P LSGL LD S
Sbjct: 412 DALSVTRFDTKSGMRVITSPNLSGLFLDGS 323
>AV414324
Length = 380
Score = 26.2 bits (56), Expect = 1.7
Identities = 8/21 (38%), Positives = 12/21 (57%)
Frame = -1
Query: 34 WLWHPDFDTAAKYVMSPPIRK 54
W+W DF + Y+ P+RK
Sbjct: 176 WVWIQDFGQQSSYIRRKPVRK 114
>AV775790
Length = 364
Score = 25.0 bits (53), Expect = 3.8
Identities = 9/12 (75%), Positives = 11/12 (91%)
Frame = +2
Query: 97 NGVNGDHSNLRY 108
NG+NG HSNLR+
Sbjct: 224 NGINGYHSNLRH 259
>TC18391 similar to UP|Q9SSW9 (Q9SSW9) Small GTP-binding protein OsRac2,
partial (27%)
Length = 507
Score = 25.0 bits (53), Expect = 3.8
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +3
Query: 40 FDTAAKYVMSPPIRKQ 55
FDTA K V+ PP RK+
Sbjct: 138 FDTAIKVVLQPPRRKE 185
>BP056372
Length = 632
Score = 25.0 bits (53), Expect = 3.8
Identities = 16/31 (51%), Positives = 18/31 (57%), Gaps = 1/31 (3%)
Frame = -3
Query: 89 IDDFRKIPNGVNGDHSNLRY-LRPKYILDDF 118
ID FR + G GDH+ LRP Y LDDF
Sbjct: 630 IDLFRALRGGKAGDHNITNVALRPLY-LDDF 541
>TC14097 homologue to UP|GCSP_PEA (P26969) Glycine dehydrogenase
[decarboxylating], mitochondrial precursor (Glycine
decarboxylase) (Glycine cleavage system P-protein) ,
partial (97%)
Length = 3660
Score = 24.6 bits (52), Expect = 5.0
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = +2
Query: 20 IGEPVLSGLALDDSWLWHPDFDT 42
+ P+ SGLA + +L HP F+T
Sbjct: 1736 VQSPIPSGLARESPFLTHPIFNT 1804
>TC8203 similar to UP|Q8L9C3 (Q8L9C3) Copia-like retroelement pol
polyprotein, partial (96%)
Length = 1198
Score = 24.3 bits (51), Expect = 6.5
Identities = 14/57 (24%), Positives = 26/57 (45%), Gaps = 3/57 (5%)
Frame = +2
Query: 57 HDKALQ---AALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLR 110
HD+ + A +T + + T + QKA IDD K + +N N++ ++
Sbjct: 473 HDQMIMLEGAKATTETVDALRTGASAMKAMQKATNIDDVDKTMDEINEQTENMKQIQ 643
>BP057889
Length = 473
Score = 24.3 bits (51), Expect = 6.5
Identities = 10/25 (40%), Positives = 12/25 (48%)
Frame = -2
Query: 26 SGLALDDSWLWHPDFDTAAKYVMSP 50
+G D SW+W P A YV P
Sbjct: 466 AGFLWDLSWIWGPLCQPAVAYVRPP 392
>BP052898
Length = 480
Score = 24.3 bits (51), Expect = 6.5
Identities = 8/20 (40%), Positives = 13/20 (65%)
Frame = +2
Query: 95 IPNGVNGDHSNLRYLRPKYI 114
+PN N H +LRY++ Y+
Sbjct: 170 LPNLYNAQHVDLRYIQKNYV 229
>TC19263 similar to UP|Q8L8A0 (Q8L8A0) Type IIB calcium ATPase MCA5, partial
(17%)
Length = 536
Score = 24.3 bits (51), Expect = 6.5
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = +3
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKI 95
LST V + + +D + N Q +GI+ F +I
Sbjct: 387 LSTSVTEGISSDADILNRRQLIYGINKFTEI 479
>TC17780 similar to GB|AAL11601.1|15983466|AF424607 At1g33990/F12G12_220
{Arabidopsis thaliana;}, partial (24%)
Length = 611
Score = 23.9 bits (50), Expect = 8.5
Identities = 18/70 (25%), Positives = 34/70 (47%), Gaps = 2/70 (2%)
Frame = +1
Query: 48 MSPPIRKQGHDKALQAALSTGVLQLVGTDHCVFNSTQKAFG--IDDFRKIPNGVNGDHSN 105
+SP ++ +K ++ GV ++ G+DHC F S ++ + + +IP +NG H
Sbjct: 112 LSPDVQ----EKLVRENPPEGVYKIKGSDHCPFFSKPQSLHKILVEIAQIP-*LNGHHDL 276
Query: 106 LRYLRPKYIL 115
P+ L
Sbjct: 277 FLSFHPQLTL 306
>TC16367 similar to UP|Q8W3L8 (Q8W3L8) Xyloglucan endo-transglycosylase,
partial (54%)
Length = 1093
Score = 23.9 bits (50), Expect = 8.5
Identities = 7/17 (41%), Positives = 9/17 (52%)
Frame = +3
Query: 34 WLWHPDFDTAAKYVMSP 50
WLW P+ T + M P
Sbjct: 399 WLWKPELPTVFSHAMGP 449
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,952,589
Number of Sequences: 28460
Number of extensions: 22139
Number of successful extensions: 97
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 97
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 97
length of query: 123
length of database: 4,897,600
effective HSP length: 79
effective length of query: 44
effective length of database: 2,649,260
effective search space: 116567440
effective search space used: 116567440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 49 (23.5 bits)
Medicago: description of AC136472.7


