

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135565.4 - phase: 0
(137 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
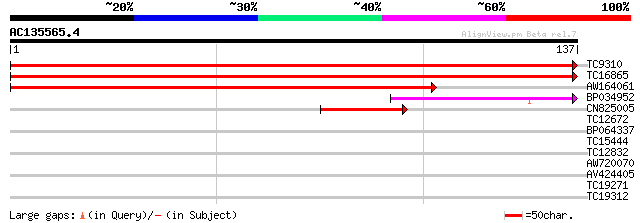
Score E
Sequences producing significant alignments: (bits) Value
TC9310 similar to UP|RS24_ARATH (Q9SS17) 40S ribosomal protein S... 265 2e-72
TC16865 similar to UP|RS24_ARATH (Q9SS17) 40S ribosomal protein ... 264 5e-72
AW164061 199 2e-52
BP034952 66 2e-12
CN825005 41 6e-05
TC12672 weakly similar to UP|Q9LV59 (Q9LV59) Genomic DNA, chromo... 27 1.6
BP064337 27 1.6
TC15444 26 2.1
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 25 3.6
AW720070 25 3.6
AV424405 25 6.2
TC19271 similar to UP|Q8S531 (Q8S531) Cytosolic aldehyde dehydro... 25 6.2
TC19312 similar to UP|Q9XIS4 (Q9XIS4) Branching enzyme 3 , part... 24 8.1
>TC9310 similar to UP|RS24_ARATH (Q9SS17) 40S ribosomal protein S24,
complete
Length = 706
Score = 265 bits (678), Expect = 2e-72
Identities = 130/137 (94%), Positives = 136/137 (98%)
Frame = +1
Query: 1 MADKAVTIRTRKFMTNRLLSRKQFVIDVLHPGRANVSKAELKEKLARIYDVKDPNTVFVF 60
MADKAVTIRTRKFMTNRLLSRKQF++DVLHPGRANVSKAELKEKLAR+YDVKDPNTVFVF
Sbjct: 85 MADKAVTIRTRKFMTNRLLSRKQFIVDVLHPGRANVSKAELKEKLARLYDVKDPNTVFVF 264
Query: 61 KFRTHFGGGKSTGFGLIYDSVENAKKYEPKYRLVRNGLDTKVEKSRKQMKERKNRAKKIR 120
KFRTHFGGGKSTGFGLIYDS+ENAKKYEPKYRL+RNGLDTKVEKSRKQ+KERKNRAKKIR
Sbjct: 265 KFRTHFGGGKSTGFGLIYDSLENAKKYEPKYRLIRNGLDTKVEKSRKQLKERKNRAKKIR 444
Query: 121 GVKKTKAGDAAKAGKKK 137
GVKKTKA DAAKAGKKK
Sbjct: 445 GVKKTKAADAAKAGKKK 495
>TC16865 similar to UP|RS24_ARATH (Q9SS17) 40S ribosomal protein S24,
complete
Length = 634
Score = 264 bits (674), Expect = 5e-72
Identities = 129/137 (94%), Positives = 135/137 (98%)
Frame = +2
Query: 1 MADKAVTIRTRKFMTNRLLSRKQFVIDVLHPGRANVSKAELKEKLARIYDVKDPNTVFVF 60
MADKAVTIRTRKFMTNRLLSRKQF++DVLHPGRANVSKAELKEKLAR+YDVKDPNTVFVF
Sbjct: 68 MADKAVTIRTRKFMTNRLLSRKQFIVDVLHPGRANVSKAELKEKLARLYDVKDPNTVFVF 247
Query: 61 KFRTHFGGGKSTGFGLIYDSVENAKKYEPKYRLVRNGLDTKVEKSRKQMKERKNRAKKIR 120
KFRTHFGGGKSTGFGLIYDS+ENAKKYEPKYRL+RNGLDTKVEKSRKQ+KERKNRAKKIR
Sbjct: 248 KFRTHFGGGKSTGFGLIYDSLENAKKYEPKYRLIRNGLDTKVEKSRKQLKERKNRAKKIR 427
Query: 121 GVKKTKAGDAAKAGKKK 137
GVKKTKA DAAK GKKK
Sbjct: 428 GVKKTKASDAAKGGKKK 478
>AW164061
Length = 340
Score = 199 bits (505), Expect = 2e-52
Identities = 95/103 (92%), Positives = 101/103 (97%)
Frame = +1
Query: 1 MADKAVTIRTRKFMTNRLLSRKQFVIDVLHPGRANVSKAELKEKLARIYDVKDPNTVFVF 60
MADKAVTIRTRKFMTNRLLSRKQF++DVLHPGRANVSK ELKEKLAR+YDVKDPNTVFVF
Sbjct: 31 MADKAVTIRTRKFMTNRLLSRKQFIVDVLHPGRANVSKTELKEKLARLYDVKDPNTVFVF 210
Query: 61 KFRTHFGGGKSTGFGLIYDSVENAKKYEPKYRLVRNGLDTKVE 103
KFRTHFGGGKSTGFGLIY S+ENA+KYEPKYRL+RNGLDTKVE
Sbjct: 211 KFRTHFGGGKSTGFGLIYHSLENAQKYEPKYRLIRNGLDTKVE 339
>BP034952
Length = 450
Score = 66.2 bits (160), Expect = 2e-12
Identities = 40/73 (54%), Positives = 42/73 (56%), Gaps = 28/73 (38%)
Frame = -3
Query: 93 LVRNGLDTKVEKSRKQMKERKNRAKKIRGVKK---------------------------- 124
L +NGLDTKVEKSRKQ+KERKNRAKKIRGVKK
Sbjct: 334 LFQNGLDTKVEKSRKQLKERKNRAKKIRGVKKVIKAYNS*WFLTLYLGFVELNWDVLFLQ 155
Query: 125 TKAGDAAKAGKKK 137
TKA DAAK GKKK
Sbjct: 154 TKASDAAKGGKKK 116
>CN825005
Length = 665
Score = 41.2 bits (95), Expect = 6e-05
Identities = 17/21 (80%), Positives = 19/21 (89%)
Frame = +2
Query: 76 LIYDSVENAKKYEPKYRLVRN 96
LIYDS+ENAKKYEPKYR + N
Sbjct: 599 LIYDSLENAKKYEPKYRFIMN 661
>TC12672 weakly similar to UP|Q9LV59 (Q9LV59) Genomic DNA, chromosome 3, P1
clone: MOB24, partial (14%)
Length = 548
Score = 26.6 bits (57), Expect = 1.6
Identities = 13/40 (32%), Positives = 24/40 (59%), Gaps = 1/40 (2%)
Frame = +3
Query: 74 FGLIYDSVENAKKYE-PKYRLVRNGLDTKVEKSRKQMKER 112
FG IY+ +EN K+ + + +R + ++E +KQ+ ER
Sbjct: 3 FGKIYEKIENTKRQQMMELEKMRLDFNRELELQKKQILER 122
>BP064337
Length = 485
Score = 26.6 bits (57), Expect = 1.6
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = -2
Query: 46 ARIYDVKDPNTVFVFKFRTHFG 67
AR+Y V TVF FK + +FG
Sbjct: 160 ARMYQVNGDETVFFFKAKDYFG 95
>TC15444
Length = 641
Score = 26.2 bits (56), Expect = 2.1
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = +2
Query: 20 SRKQFVIDVLHPGRANVSK 38
+R V+DVLH GRAN SK
Sbjct: 86 NRVLHVVDVLHRGRANKSK 142
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 25.4 bits (54), Expect = 3.6
Identities = 15/43 (34%), Positives = 23/43 (52%)
Frame = +2
Query: 11 RKFMTNRLLSRKQFVIDVLHPGRANVSKAELKEKLARIYDVKD 53
++F N + +V D+L G ELK +LAR +D+KD
Sbjct: 65 KRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREFDMKD 193
>AW720070
Length = 484
Score = 25.4 bits (54), Expect = 3.6
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = +2
Query: 105 SRKQMKERKNRAKKIRGVKKTKAGDAAKAGKKK 137
+ K+ K++K + KK + VKK + K KKK
Sbjct: 104 AEKKRKKKKKKRKKKKQVKKESSQQQEKKKKKK 202
>AV424405
Length = 335
Score = 24.6 bits (52), Expect = 6.2
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = -1
Query: 103 EKSRKQMKERKNRAKKIRGVKKTKAGDAAKAGKK 136
+KS K M + +NR+K+ R +K K+ + GKK
Sbjct: 110 QKSLKTMCDTRNRSKQ*RLEEKHKSWKEQEEGKK 9
>TC19271 similar to UP|Q8S531 (Q8S531) Cytosolic aldehyde dehydrogenase
RF2C, partial (25%)
Length = 485
Score = 24.6 bits (52), Expect = 6.2
Identities = 14/33 (42%), Positives = 16/33 (48%)
Frame = +1
Query: 55 NTVFVFKFRTHFGGGKSTGFGLIYDSVENAKKY 87
N F F FGG K +GFG Y +E KY
Sbjct: 229 NCFFAFDIDCPFGGYKMSGFGRDY-GLEALHKY 324
>TC19312 similar to UP|Q9XIS4 (Q9XIS4) Branching enzyme 3 , partial (4%)
Length = 523
Score = 24.3 bits (51), Expect = 8.1
Identities = 10/25 (40%), Positives = 17/25 (68%)
Frame = -2
Query: 34 ANVSKAELKEKLARIYDVKDPNTVF 58
ANVS +LK +R ++ +DP+ +F
Sbjct: 213 ANVSMVQLKLLTSRSFEAEDPSGMF 139
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,460,954
Number of Sequences: 28460
Number of extensions: 13983
Number of successful extensions: 92
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 92
length of query: 137
length of database: 4,897,600
effective HSP length: 81
effective length of query: 56
effective length of database: 2,592,340
effective search space: 145171040
effective search space used: 145171040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 50 (23.9 bits)
Medicago: description of AC135565.4


