

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135467.11 - phase: 0
(807 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
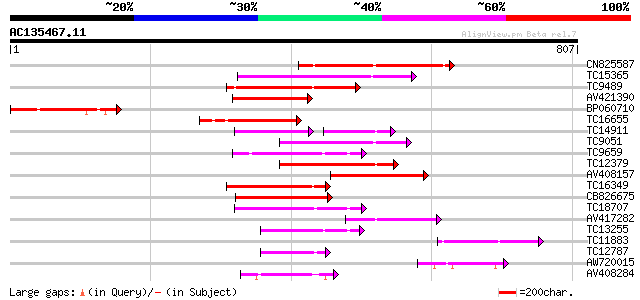
Score E
Sequences producing significant alignments: (bits) Value
CN825587 226 8e-60
TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {... 184 4e-47
TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase... 159 2e-39
AV421390 151 3e-37
BP060710 150 8e-37
TC16655 homologue to UP|O99018 (O99018) Chloroplast protease pre... 145 2e-35
TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), pa... 93 1e-32
TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 ... 136 1e-32
TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATP... 130 8e-31
TC12379 homologue to UP|O04327 (O04327) Cell division protein Ft... 129 1e-30
AV408157 122 2e-28
TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 118 4e-27
CB826675 117 6e-27
TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J... 114 6e-26
AV417282 105 3e-23
TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%) 99 2e-21
TC11883 similar to UP|YME1_YEAST (P32795) YME1 protein (TAT-bin... 85 4e-17
TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (T... 79 4e-15
AW720015 77 1e-14
AV408284 77 1e-14
>CN825587
Length = 663
Score = 226 bits (577), Expect = 8e-60
Identities = 113/222 (50%), Positives = 156/222 (69%)
Frame = +2
Query: 411 ARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTN 470
A+ AP IVFIDEIDA+GR+RG G G NDERE T+NQLL EMDGF +GV+VLA TN
Sbjct: 2 AKSKAPCIVFIDEIDAVGRQRG-AGLGGGNDEREQTINQLLTEMDGFSGNSGVIVLAATN 178
Query: 471 RADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGF 530
R D+LD+ALLRPGRFDR +++D PD+ GR +I Q++ + L + + ++A TPGF
Sbjct: 179 RPDVLDSALLRPGRFDRQVTVDRPDVAGRVKILQVHSRGKALAKDVDF--DKIARRTPGF 352
Query: 531 AGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEA 590
GAD+ N+ NEAA++AAR D +++ D A++RII G EKKN V+S+ +++ VAYHEA
Sbjct: 353 TGADLQNLMNEAAILAARRDLKEISKDEISDALERIIAGPEKKNAVVSEEKKKLVAYHEA 532
Query: 591 GHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSENLLRT 632
GHA+ G + +P+ K++I+PRG A G + PSE L +
Sbjct: 533 GHALVGALMPEYDPVAKISIIPRGQAG-GLTFFAPSEERLES 655
>TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;}, partial (87%)
Length = 1385
Score = 184 bits (468), Expect = 4e-47
Identities = 101/256 (39%), Positives = 152/256 (58%), Gaps = 1/256 (0%)
Frame = +3
Query: 325 VAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVP 383
+ G ++ +EI E + +K+P+ +E LG PKG LL GPPGTGKTLLA+A A +
Sbjct: 333 IGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLARAVAHHTDCT 512
Query: 384 FLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDER 443
F+ +SGS+ ++ ++G G VR LF AR+ APSI+F+DEID+IG R G + E
Sbjct: 513 FIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARMESGSGXGDSEV 692
Query: 444 ESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIF 503
+ T+ +LL ++DGF + + VL TNR DILD ALLRPGR DR I P+ + R I
Sbjct: 693 QRTMLELLNQLDGFEASNKIKVLMATNRIDILDQALLRPGRIDRKIEFPNPNEESRQDIL 872
Query: 504 QIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAI 563
+I+ +++ L +++A G +GA++ VC EA + A R VT + FE A+
Sbjct: 873 KIHSRRMNLMR--GIDLKKIAEKMNGASGAELKAVCTEAGMFALRERRVHVTQEDFEMAV 1046
Query: 564 DRIIGGLEKKNRVISK 579
+++ +KN + K
Sbjct: 1047AKVMKKETEKNMSLRK 1094
>TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase subunit
RPT2a, partial (69%)
Length = 916
Score = 159 bits (401), Expect = 2e-39
Identities = 90/192 (46%), Positives = 118/192 (60%), Gaps = 1/192 (0%)
Frame = +1
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
V KV+K + D+ G + QEI E V L +P+ YE++G K PKG +L G PGT
Sbjct: 346 VMKVEKAPLES--YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGT 519
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTLLAKA A + FL + GS+ ++ ++G GP VR LF+ A +PSIVFIDEIDA+
Sbjct: 520 GKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAV 699
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
G KR SG E + T+ +LL ++DGF + V V+ TNR + LD ALLRPGR DR
Sbjct: 700 GTKR-YDAHSGGEREIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDR 876
Query: 488 TISIDVPDIKGR 499
I +PD K R
Sbjct: 877 KIEFPLPDTKTR 912
>AV421390
Length = 409
Score = 151 bits (382), Expect = 3e-37
Identities = 74/115 (64%), Positives = 89/115 (77%)
Frame = +2
Query: 317 KNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKAT 376
KN FKDV GC++AKQE+ E V +L+NP K+ LG K+PKG LL G PGTGKTLLAKA
Sbjct: 53 KNVKTFKDVKGCDDAKQELEEVVEYLRNPAKFTRLGGKLPKGILLTGAPGTGKTLLAKAI 232
Query: 377 AGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKR 431
AGE+GVPF +GS+F EMFVGVG RVR+LFQ A++ AP I+FIDEIDA+G R
Sbjct: 233 AGEAGVPFFYRAGSEFEEMFVGVGARRVRSLFQAAKKKAPCIIFIDEIDAVGSTR 397
>BP060710
Length = 534
Score = 150 bits (379), Expect = 8e-37
Identities = 98/167 (58%), Positives = 113/167 (66%), Gaps = 9/167 (5%)
Frame = -1
Query: 1 MIFSRIGRSLSRSSRVKNLLHGETRLGTLYGVSRTNVFVDDVEKGLGFVRGYVSSA-IAR 59
MIFSRIGRSLSRSSR +NL G+ RLGTL SRTNV E GLGF RGY SS AR
Sbjct: 477 MIFSRIGRSLSRSSRARNLFCGDGRLGTLGVSSRTNVAA---EGGLGFFRGYASSVGAAR 307
Query: 60 NNG-FGSNLYDFKSIAAN-RMLHRMFSSESPKKKNYEKFYPKEKKEVPKG---EEKKSES 114
++G SNL FKS+AAN R+ HR+FSSE+PKK NYE FYPKEKK+ PKG +KK ES
Sbjct: 306 SSGSLLSNLAAFKSVAANPRLHHRLFSSEAPKK-NYENFYPKEKKDAPKGNGNNDKKYES 130
Query: 115 KDESKSNTEDGGSFHEAFIK---QFQNYLTPLLVVGLFLSSLSLGPR 158
KDES +N+++ E F K PLLV+GLF SS S GPR
Sbjct: 129 KDESNTNSDE----QEIFKKLS*NISKSTHPLLVMGLFFSSFSFGPR 1
>TC16655 homologue to UP|O99018 (O99018) Chloroplast protease precursor,
partial (39%)
Length = 934
Score = 145 bits (366), Expect = 2e-35
Identities = 80/146 (54%), Positives = 102/146 (69%)
Frame = +3
Query: 270 LLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCE 329
++L+G L+ + R+ GG G G G G F KA K V F DVAG +
Sbjct: 516 IILIGGLFLLTRR-PGGMG---GPGGPGFPLTFGQSKA---KFQMEPNTGVTFDDVAGVD 674
Query: 330 EAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISG 389
EAKQ+ ME V FLK P+++ +GA+IPKG LL+GPPGTGKTLLAKA AGE+GVPF SISG
Sbjct: 675 EAKQDFMEVVEFLKKPERFTAVGARIPKGVLLIGPPGTGKTLLAKAIAGEAGVPFFSISG 854
Query: 390 SDFMEMFVGVGPSRVRNLFQEARQCA 415
S+F+EMFVGVG SRVR+LF++A++ A
Sbjct: 855 SEFVEMFVGVGASRVRDLFKKAKENA 932
>TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), partial (32%)
Length = 1418
Score = 93.2 bits (230), Expect(2) = 1e-32
Identities = 46/113 (40%), Positives = 67/113 (58%)
Frame = +2
Query: 320 VYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGE 379
V ++DV G E+ K+ I++ V K G + G LL GPPGTGKTLLAKA A E
Sbjct: 215 VKWEDVGGLEDVKKSILDTVQLPLLHKDLFASGLRKRSGVLLYGPPGTGKTLLAKAVATE 394
Query: 380 SGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRG 432
+ FLS+ G + + M++G VR++FQ+AR P ++F DE+D++ G
Sbjct: 395 CSLNFLSVKGPELINMYIGESEKNVRDIFQKARSARPCVIFFDELDSLAPASG 553
Score = 64.3 bits (155), Expect(2) = 1e-32
Identities = 37/104 (35%), Positives = 60/104 (57%), Gaps = 2/104 (1%)
Frame = +3
Query: 447 LNQLLVEMDGFGTTA-GVVVLAGTNRADILDNALLRPGRFDRTISIDV-PDIKGRDQIFQ 504
L+Q+L E+DG ++ + ++ +NR D++D ALLRPGRFD+ + + V D R+++ +
Sbjct: 597 LSQMLAEIDGLSDSSQDLFIIGASNRPDLIDPALLRPGRFDKLLYVGVNSDASYRERVLK 776
Query: 505 IYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAAR 548
+K KL + S YS P F GAD +C +A AA+
Sbjct: 777 ALTRKFKLHEDVSLYS-IAKKCPPNFTGADTYALCADAWFHAAK 905
>TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (46%)
Length = 867
Score = 136 bits (342), Expect = 1e-32
Identities = 74/188 (39%), Positives = 109/188 (57%)
Frame = +2
Query: 385 LSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERE 444
+ + GS+ ++ +VG G VR LFQ AR IVF DE+DAIG R G G N E +
Sbjct: 2 IRVIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDAIGGARFDDGVGGDN-EVQ 178
Query: 445 STLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQ 504
T+ +++ ++DGF + VL TNR D LD ALLRPGR DR + +PD++ R QIF+
Sbjct: 179 RTMLEIVNQLDGFDARGNIKVLMATNRPDTLDPALLRPGRLDRKVEFGLPDLESRTQIFK 358
Query: 505 IYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAID 564
I+ + + + + + + L+ L P GADI +VC EA + A R VT F A++
Sbjct: 359 IHTRTMNCERDIRF--ELLSRLCPNSTGADIRSVCTEAGMYAIRARRKTVTEKDFLDAVN 532
Query: 565 RIIGGLEK 572
++I G +K
Sbjct: 533 KVIKGYQK 556
>TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATPase,
partial (74%)
Length = 1060
Score = 130 bits (327), Expect = 8e-31
Identities = 83/195 (42%), Positives = 116/195 (58%), Gaps = 4/195 (2%)
Frame = +1
Query: 317 KNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEEL--GAKIP-KGALLVGPPGTGKTLLA 373
K + + DVAG E AKQ + E V P K+ + G + P + LL GPPGTGK+ LA
Sbjct: 445 KPNIKWNDVAGLESAKQSLQEAVIL---PVKFPQFFTGKRRPWRAFLLYGPPGTGKSYLA 615
Query: 374 KATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGR 433
KA A E+ F S+S SD ++G V NLFQ AR+ APSI+F+DEID++ +RG
Sbjct: 616 KAVATEADSTFFSVSSSDLGSKWMGESEKLVSNLFQMARESAPSIIFVDEIDSLCGQRGE 795
Query: 434 GGFSGSNDERESTLNQLLVEMDGFGTT-AGVVVLAGTNRADILDNALLRPGRFDRTISID 492
G S ++ ++ +LLV+M G G V+VLA TN LD A+ R RFD+ I I
Sbjct: 796 GNESEASRRIKT---ELLVQMQGVGNNDQKVLVLAATNTPYALDQAIRR--RFDKRIYIP 960
Query: 493 VPDIKGRDQIFQIYL 507
+PD+K R +F+++L
Sbjct: 961 LPDVKARQHMFKVHL 1005
>TC12379 homologue to UP|O04327 (O04327) Cell division protein FtsH isolog,
partial (15%)
Length = 505
Score = 129 bits (325), Expect = 1e-30
Identities = 74/169 (43%), Positives = 110/169 (64%)
Frame = +1
Query: 385 LSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERE 444
LS+S S + + + + S R L+QEA++ APS+VFIDE+DA+GR+RG SG ER+
Sbjct: 1 LSLSLSLSLSLSLSLSLSLSRALYQEAKENAPSVVFIDELDAVGRERGLIKGSGGQ-ERD 177
Query: 445 STLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQ 504
+TLNQLLV +DGF V+ +A TNR DILD AL+RPGRFDR I I P + GR +I +
Sbjct: 178 ATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGRIEILK 357
Query: 505 IYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQ 553
++ +K + + Y + +A++T G GA++AN+ AA+ R ++
Sbjct: 358 VHARKKPIAGDVDYMA--VASMTDGMVGAELANIVEVAAINMMRDTRTE 498
>AV408157
Length = 423
Score = 122 bits (306), Expect = 2e-28
Identities = 62/140 (44%), Positives = 88/140 (62%)
Frame = +3
Query: 457 FGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEP 516
F + + V+VL TNRAD+LD AL RPGRFDR + ++ PD GR+ I ++++ + +L
Sbjct: 3 FDSNSAVIVLGATNRADVLDPALRRPGRFDRVVMVEAPDRIGREAILKVHVSRKELPLAK 182
Query: 517 SYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRV 576
+A +T GF GAD+AN+ NEAAL+A R ++ V F A++R I G+EKK
Sbjct: 183 DVDLDDIARMTTGFTGADLANLVNEAALLAGRKNKVVVERIDFIQAVERSIAGIEKKTAK 362
Query: 577 ISKRERRTVAYHEAGHAVAG 596
+ E+ VA HEAGHAV G
Sbjct: 363 LQGSEKAVVARHEAGHAVVG 422
>TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (65%)
Length = 901
Score = 118 bits (295), Expect = 4e-27
Identities = 61/149 (40%), Positives = 98/149 (64%), Gaps = 1/149 (0%)
Frame = +3
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
++ + ++ K V + D+ GC+ KQEI E V L + + Y+++G P+G LL GPPGT
Sbjct: 456 ISLLSQSEKPDVTYNDIGGCDIQKQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGT 635
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKT+LAKA A + F+ + GS+F++ ++G GP VR++F+ A++ AP+I+FIDE+DAI
Sbjct: 636 GKTMLAKAVANHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLAKENAPAIIFIDEVDAI 815
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDG 456
R +G++ E + L +LL +MDG
Sbjct: 816 ATAR-FDAQTGADREVQRILMELLNQMDG 899
>CB826675
Length = 551
Score = 117 bits (294), Expect = 6e-27
Identities = 60/139 (43%), Positives = 93/139 (66%), Gaps = 1/139 (0%)
Frame = +1
Query: 322 FKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGES 380
+ DV G E+ QE++E + + + +++++LG + PKG LL GPPGTGKTL+A+A A ++
Sbjct: 133 YNDVGGLEKQIQELVEAIVLPMTHKERFQKLGVRPPKGVLLYGPPGTGKTLMARACAAQT 312
Query: 381 GVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSN 440
FL ++G ++MF+G G VR+ FQ A++ +P I+FIDEIDAIG KR SG
Sbjct: 313 NATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRFDSEVSGDR 492
Query: 441 DERESTLNQLLVEMDGFGT 459
E + T+ +LL ++DGF +
Sbjct: 493 -EVQRTMLELLNQLDGFSS 546
>TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J8.110 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 771
Score = 114 bits (285), Expect = 6e-26
Identities = 69/191 (36%), Positives = 106/191 (55%), Gaps = 3/191 (1%)
Frame = +3
Query: 320 VYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAG 378
V F D+ +E K+ + E V L+ P ++ K +G LL GPPGTGKT+LA A A
Sbjct: 192 VTFGDIGAMDEIKESLQELVMLPLRRPDLFKGGLLKPCRGILLFGPPGTGKTMLAXAIAN 371
Query: 379 ESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSG 438
E+G F+++S S + G VR LF A + AP+I+F+DE+D++ +R R G
Sbjct: 372 EAGASFINVSMSTITSKWFGEDEKNVRALFSLAAKVAPTIIFVDEVDSMLGQRTR---VG 542
Query: 439 SNDERESTLNQLLVEMDGFGTTAG--VVVLAGTNRADILDNALLRPGRFDRTISIDVPDI 496
++ N+ + DG T G ++VLA TNR LD A++R RF+R I + +P +
Sbjct: 543 EHEAMRKIKNEFMTHWDGLLTAPGEQILVLAATNRPFDLDEAIIR--RFERRIMVGLPSV 716
Query: 497 KGRDQIFQIYL 507
+ R+ I + L
Sbjct: 717 ENREMILKTLL 749
>AV417282
Length = 407
Score = 105 bits (262), Expect = 3e-23
Identities = 58/136 (42%), Positives = 82/136 (59%)
Frame = +3
Query: 479 LLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANV 538
L+RPGRFDR + + PD++GR QI + ++ K+ + +A TPGF+GAD+AN+
Sbjct: 3 LVRPGRFDRHVVVPNPDVEGRRQILESHMSKVLKADDVDLMI--IARGTPGFSGADLANM 176
Query: 539 CNEAALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWF 598
N AAL AA V+M E A D+I+ G E+K+ VIS R+ A+HE GHA+
Sbjct: 177 VNIAALKAAMDGAKAVSMHDLEFAKDKIMMGSERKSAVISDESRKMTAFHEGGHALVAIH 356
Query: 599 LEHCEPLLKVTIVPRG 614
+ P+ K TIVPRG
Sbjct: 357 TDGALPVHKATIVPRG 404
>TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%)
Length = 532
Score = 99.4 bits (246), Expect = 2e-21
Identities = 65/149 (43%), Positives = 86/149 (57%), Gaps = 1/149 (0%)
Frame = +1
Query: 357 KGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAP 416
KG LL GPPGTGKT+LAKA A E F +IS S + + G V+ LFQ AR AP
Sbjct: 37 KGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAP 216
Query: 417 SIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVV-VLAGTNRADIL 475
S +F+DEIDAI +RG R T +LL++MDG T +V VLA TN L
Sbjct: 217 STIFLDEIDAIISQRGEARSEHEASRRLKT--ELLIQMDGLTRTDELVFVLAATNLPWEL 390
Query: 476 DNALLRPGRFDRTISIDVPDIKGRDQIFQ 504
D A+LR R ++ I + +P+ + R +F+
Sbjct: 391 DAAMLR--RLEKRILVPLPEPEARVAMFE 471
>TC11883 similar to UP|YME1_YEAST (P32795) YME1 protein (TAT-binding
homolog 11) (OSD1 protein) , partial (7%)
Length = 551
Score = 85.1 bits (209), Expect = 4e-17
Identities = 53/154 (34%), Positives = 86/154 (55%), Gaps = 3/154 (1%)
Frame = +2
Query: 609 TIVPRGTAALGFAQYVPSENLLRT-KEQLLDMTCMTLGGRAAEQVLIGA--ISTGAQNDL 665
TIVPRG A LG +P + T ++Q+L + +GGR AE+++ G +++GA +DL
Sbjct: 5 TIVPRGMA-LGMVTQLPEVDQTSTSRKQMLARLDVCMGGRVAEELIFGESEVTSGASSDL 181
Query: 666 EKVTKMTYAQVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTV 725
TK+ A V YG S +VGL + N+D G+ S +T +I++EVR+ + AY
Sbjct: 182 SHATKLARAMVTQYGMSSEVGLATHNYNDD--GRSMSSETRLLIEKEVRNLLERAYTNAK 355
Query: 726 QLIEEHKEKLAQIAELLLEKEVLHQEDLVRILGE 759
++ H ++L +A LLE E L + +L +
Sbjct: 356 TILTTHDKELQALANALLEHETLTGSQINALLAK 457
>TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (TAT-binding
homolog 4), partial (20%)
Length = 389
Score = 78.6 bits (192), Expect = 4e-15
Identities = 40/100 (40%), Positives = 59/100 (59%)
Frame = +3
Query: 357 KGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAP 416
KG LL GPPGTGKT+LAKA A E+G F++IS S + G G V+ +F A + +P
Sbjct: 84 KGILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKISP 263
Query: 417 SIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDG 456
S++F+DE+D + +R G ++ N+ +V DG
Sbjct: 264 SVIFVDEVDXMLXRREN---PGEHEAMRKMXNEFMVHWDG 374
>AW720015
Length = 542
Score = 77.0 bits (188), Expect = 1e-14
Identities = 53/144 (36%), Positives = 78/144 (53%), Gaps = 14/144 (9%)
Frame = +1
Query: 581 ERRTVAYHEAGHAVAGWFLEHC---EPLL-KVTIVPRGTAALGFAQYVPSEN---LLRTK 633
E+ VA HEAGHAV G + + +P + K++I+PR ALGF Y+P N L
Sbjct: 25 EKAVVARHEAGHAVVGTAVANLIAGQPRVEKLSILPRSGGALGFT-YIPPTNEDRYLLFI 201
Query: 634 EQLLDMTCMTLGGRAAEQVLI-GAISTGAQNDLEKVTKMTYAQVAIYGFSEKVGLLSF-- 690
++L LGGRAAE+V+ G +STGA +D+ + T M Y +A YG S+ +G +S
Sbjct: 202 DELRGRLVTLLGGRAAEEVVYSGRVSTGALDDIRRATDMAYKAIAEYGLSQTIGPVSIGT 381
Query: 691 ----PQNEDQFGKPYSGDTGNIID 710
+E P+ D G ++D
Sbjct: 382 LSNGGMDESGGSVPWGRDQGRLVD 453
>AV408284
Length = 425
Score = 76.6 bits (187), Expect = 1e-14
Identities = 57/149 (38%), Positives = 81/149 (54%), Gaps = 10/149 (6%)
Frame = +1
Query: 329 EEAKQEIMEFVHF-LKNPKKYEE---LGAKIPKGALLVGPPGTGKTLLAKATAGESGVPF 384
E K+ + E V LK P + LG + KG LL GPPGTGKT+LAKA A ESG F
Sbjct: 4 ESIKEALFELVILPLKRPDLFSHGKLLGPQ--KGVLLYGPPGTGKTMLAKAIAKESGAVF 177
Query: 385 LSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDA-IGRKRGRGGFSGSNDER 443
+++ S+ M + G V +F A + P+I+FIDE+D+ +G++R +
Sbjct: 178 INVRISNLMSKWFGDAQKLVAAVFSLAHKLQPAIIFIDEVDSFLGQRR--------TTDH 333
Query: 444 ESTLN---QLLVEMDGFGT--TAGVVVLA 467
E+ LN + + DGF T A V+VLA
Sbjct: 334 EALLNMKTEFMALWDGFTTDQNARVMVLA 420
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,014,486
Number of Sequences: 28460
Number of extensions: 130787
Number of successful extensions: 783
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 732
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 752
length of query: 807
length of database: 4,897,600
effective HSP length: 98
effective length of query: 709
effective length of database: 2,108,520
effective search space: 1494940680
effective search space used: 1494940680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135467.11


