

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.23 + phase: 0
(150 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
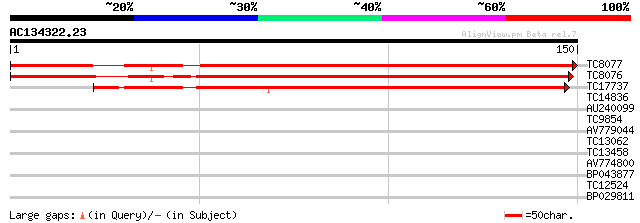
Score E
Sequences producing significant alignments: (bits) Value
TC8077 similar to UP|FER1_PEA (P09911) Ferredoxin I, chloroplast... 211 4e-56
TC8076 similar to UP|FER2_ARATH (O04090) Ferredoxin 2, chloropla... 204 4e-54
TC17737 similar to UP|FER3_MAIZE (P27788) Ferredoxin III, chloro... 154 5e-39
TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precur... 32 0.036
AU240099 27 2.0
TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1 (AT3g1... 25 4.3
AV779044 25 4.3
TC13062 similar to UP|Q889K7 (Q889K7) MaoC-like domain protein, ... 25 5.7
TC13458 weakly similar to UP|Z282_HUMAN (Q9UDV7) Zinc finger pro... 25 5.7
AV774800 24 9.7
BP043877 24 9.7
TC12524 similar to UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kina... 24 9.7
BP029811 24 9.7
>TC8077 similar to UP|FER1_PEA (P09911) Ferredoxin I, chloroplast
precursor, partial (81%)
Length = 703
Score = 211 bits (537), Expect = 4e-56
Identities = 112/154 (72%), Positives = 120/154 (77%), Gaps = 4/154 (2%)
Frame = +1
Query: 1 MATTPALYGTAVSTSFLRRQPMPVSISTTTKAFPSG----FGLKSKTGKRGDLAVAMATY 56
MATTPAL GT V+TSFLRRQP+ KAFP+ FGLKS G R V MA Y
Sbjct: 76 MATTPALSGTMVNTSFLRRQPL--------KAFPNVGQALFGLKSGCGGR----VTMAAY 219
Query: 57 KVKLVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSF 116
KVKL+TPEG EF+CP DVYILDHAEE GID+PYSCRAGSCSSCAGKV G VDQSDGSF
Sbjct: 220 KVKLITPEGPFEFECPDDVYILDHAEEQGIDIPYSCRAGSCSSCAGKVVGGNVDQSDGSF 399
Query: 117 LDDDQIEEGWVLTCVAYPTSDVTIETHKEEELTA 150
LDDDQI+ G+VLTCVAYP SDV IETHKEEELTA
Sbjct: 400 LDDDQIDAGFVLTCVAYPQSDVVIETHKEEELTA 501
>TC8076 similar to UP|FER2_ARATH (O04090) Ferredoxin 2, chloroplast
precursor, partial (78%)
Length = 717
Score = 204 bits (520), Expect = 4e-54
Identities = 109/153 (71%), Positives = 117/153 (76%), Gaps = 4/153 (2%)
Frame = +2
Query: 1 MATTPALYGTAVSTSFLRRQPMPVSISTTTKAFPSG----FGLKSKTGKRGDLAVAMATY 56
MAT PA G VSTSFLRRQP+ AFP+ FG+K G RG V MA Y
Sbjct: 32 MATAPAFSGATVSTSFLRRQPVA--------AFPNVGQVMFGVKG--GSRGG-RVTMAAY 178
Query: 57 KVKLVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSF 116
KVKL+TPEG QEF+CP DVYILD AEEVGID+PYSCRAGSCSSCAGKV GAVDQSDGSF
Sbjct: 179 KVKLITPEGPQEFECPDDVYILDQAEEVGIDIPYSCRAGSCSSCAGKVVEGAVDQSDGSF 358
Query: 117 LDDDQIEEGWVLTCVAYPTSDVTIETHKEEELT 149
LDDDQ+E G+VLTCVAYP SDV IETHKEEELT
Sbjct: 359 LDDDQVEGGFVLTCVAYPKSDVVIETHKEEELT 457
>TC17737 similar to UP|FER3_MAIZE (P27788) Ferredoxin III, chloroplast
precursor (Fd III), partial (72%)
Length = 736
Score = 154 bits (390), Expect = 5e-39
Identities = 76/127 (59%), Positives = 96/127 (74%), Gaps = 1/127 (0%)
Frame = +2
Query: 23 PVSISTTTKAFPSGFGLKSKTGKRGDLAVAMATYKVKLVTPEGTQ-EFDCPSDVYILDHA 81
P+S + K+ FGLKS T R AMA YK+KLV P+G + EF+ DVYILD A
Sbjct: 149 PISFGSV-KSVSRSFGLKSSTPSR---VTAMAAYKIKLVGPDGKENEFEAEDDVYILDAA 316
Query: 82 EEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVTIE 141
E G++LPYSCRAG+CS+CAGK+ G+VDQSDGSFLDD+Q+++G+VLTCV+YPT+D IE
Sbjct: 317 ENAGVELPYSCRAGACSTCAGKIVTGSVDQSDGSFLDDNQLKDGFVLTCVSYPTADCVIE 496
Query: 142 THKEEEL 148
THKE EL
Sbjct: 497 THKEGEL 517
>TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precursor ,
partial (92%)
Length = 1416
Score = 32.3 bits (72), Expect = 0.036
Identities = 18/42 (42%), Positives = 24/42 (56%)
Frame = +2
Query: 87 DLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWVL 128
D PY+ R G+C K+AA + S GS LD+DQI V+
Sbjct: 716 DYPYTGRDGTCKFDKSKIAASVANFSVGS-LDEDQIAANLVM 838
>AU240099
Length = 300
Score = 26.6 bits (57), Expect = 2.0
Identities = 14/44 (31%), Positives = 17/44 (37%)
Frame = +2
Query: 71 CPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDG 114
C VY++D GI PY CS C G + DG
Sbjct: 149 CDKTVYVVDLLTLEGI--PYHKSCFKCSHCKGNLTMSTYSSMDG 274
>TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1
(AT3g17100/K14A17_22_), partial (43%)
Length = 1108
Score = 25.4 bits (54), Expect = 4.3
Identities = 15/47 (31%), Positives = 21/47 (43%), Gaps = 2/47 (4%)
Frame = +2
Query: 95 GSC--SSCAGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVT 139
G C SSC+G AV+ D D DQI+E P ++ +
Sbjct: 551 GCC**SSCSGGEGEDAVEHGDSHQPD*DQIQEAAAAAAEEAPAAEAS 691
>AV779044
Length = 533
Score = 25.4 bits (54), Expect = 4.3
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = -1
Query: 64 EGTQEFDCPSDVYILDHAEEVGI 86
E TQEF SDV+ LD + VG+
Sbjct: 191 ENTQEFFGSSDVFFLDVSSFVGV 123
>TC13062 similar to UP|Q889K7 (Q889K7) MaoC-like domain protein, partial
(5%)
Length = 390
Score = 25.0 bits (53), Expect = 5.7
Identities = 8/18 (44%), Positives = 13/18 (71%)
Frame = -1
Query: 120 DQIEEGWVLTCVAYPTSD 137
D I GW+L+C+ YP ++
Sbjct: 114 DVII*GWILSCIQYPQNE 61
>TC13458 weakly similar to UP|Z282_HUMAN (Q9UDV7) Zinc finger protein 282
(HTLV-I U5RE binding protein 1) (HUB-1), partial (3%)
Length = 810
Score = 25.0 bits (53), Expect = 5.7
Identities = 13/31 (41%), Positives = 15/31 (47%)
Frame = +2
Query: 7 LYGTAVSTSFLRRQPMPVSISTTTKAFPSGF 37
LY TAV + L PMP T K P G+
Sbjct: 179 LYITAVESRKLDETPMPAVNGTDEKCAPCGY 271
>AV774800
Length = 452
Score = 24.3 bits (51), Expect = 9.7
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = -3
Query: 71 CPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDD 120
C + V +L + + LPY R+ S A VAA +D S S D+D
Sbjct: 240 CDNGVVVLTNNAK----LPYQIRSASFRISASIVAAVNLDDSGTSESDED 103
>BP043877
Length = 526
Score = 24.3 bits (51), Expect = 9.7
Identities = 12/27 (44%), Positives = 14/27 (51%)
Frame = -1
Query: 11 AVSTSFLRRQPMPVSISTTTKAFPSGF 37
AVS F+ P STTT+ F GF
Sbjct: 250 AVSNPFMSLHPAQPVPSTTTRGFSCGF 170
>TC12524 similar to UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kinase 1,
partial (27%)
Length = 834
Score = 24.3 bits (51), Expect = 9.7
Identities = 11/21 (52%), Positives = 11/21 (52%)
Frame = -2
Query: 83 EVGIDLPYSCRAGSCSSCAGK 103
E G P CR GSC S GK
Sbjct: 68 E*GYRSPEGCRLGSCCSSRGK 6
>BP029811
Length = 481
Score = 24.3 bits (51), Expect = 9.7
Identities = 13/47 (27%), Positives = 24/47 (50%)
Frame = -3
Query: 102 GKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVTIETHKEEEL 148
GK+ +D ++G+F+DD ++ G V P S ++ T + L
Sbjct: 353 GKLLVTDLDSTNGTFIDDKRLRPG-----VISPVSSGSLVTFGDTHL 228
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,510,052
Number of Sequences: 28460
Number of extensions: 32729
Number of successful extensions: 122
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 119
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 119
length of query: 150
length of database: 4,897,600
effective HSP length: 82
effective length of query: 68
effective length of database: 2,563,880
effective search space: 174343840
effective search space used: 174343840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 51 (24.3 bits)
Medicago: description of AC134322.23


