

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
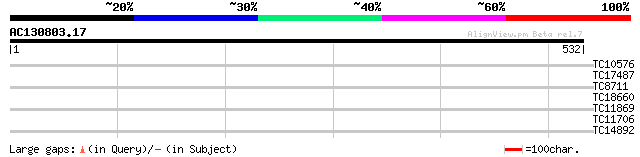
Query= AC130803.17 + phase: 0
(532 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10576 similar to UP|Q93ZQ3 (Q93ZQ3) AT3g63200/F16M2_50, partia... 29 1.7
TC17487 UP|CAE45597 (CAE45597) SAR DNA-binding protein-like prot... 29 2.2
TC8711 weakly similar to UP|VCLC_PEA (P13918) Vicilin precursor,... 28 2.8
TC18660 27 6.3
TC11869 27 8.2
TC11706 homologue to UP|Q8GSM9 (Q8GSM9) Squalene monooxygenase 2... 27 8.2
TC14892 similar to UP|Q9Z0W6 (Q9Z0W6) Pax transcription activati... 27 8.2
>TC10576 similar to UP|Q93ZQ3 (Q93ZQ3) AT3g63200/F16M2_50, partial (39%)
Length = 751
Score = 29.3 bits (64), Expect = 1.7
Identities = 23/89 (25%), Positives = 35/89 (38%), Gaps = 11/89 (12%)
Frame = -2
Query: 348 SKPINLATS---------PEFFFYTNAPTGQRKAFMDAWSKVRRKSVKHLGVRSGVAHEA 398
S P++ +TS P + T + Q+KAF + WS V + GV H
Sbjct: 456 SNPLSFSTSSSVVASVPKPNAWILT*SVRFQQKAFPNIWSTVSETPSSTMPTTDGVEHVR 277
Query: 399 YTQWVIDRAEEIGM--PYPAMRYVSSSTP 425
+ + + P+P R SSTP
Sbjct: 276 WFSFSFSFSHAFAFDDPFPRERTRRSSTP 190
>TC17487 UP|CAE45597 (CAE45597) SAR DNA-binding protein-like protein, complete
Length = 1794
Score = 28.9 bits (63), Expect = 2.2
Identities = 40/138 (28%), Positives = 57/138 (40%), Gaps = 6/138 (4%)
Frame = +1
Query: 379 KVRRKSVKHLGVRSGVAH-----EAYTQWVIDRAEEIG-MPYPAMRYVSSSTPSIPLPLL 432
++R K LG +G A EAY + DR + G + PA Y S+ I
Sbjct: 1312 RLRNLEGKELGRFAGSAKGKPKIEAYDK---DRKKGAGGLITPAKTYNPSADAVISQKSD 1482
Query: 433 PATQDMYQEHLAMESHEKQVWKARYNQAENLIMTLDGRDEQKTHENLMLKKELAKARKEL 492
A + E A + EK+ K + + E + L DE E ++KKE K RKE
Sbjct: 1483 SAMDEDTHEPSADKKKEKKEKKKKKEKNEEKDVPLLA-DEDGDEEKEVVKKEKKKKRKES 1659
Query: 493 EEKDELLLRDSKRAKGRR 510
E EL D K ++
Sbjct: 1660 TENVELQNGDDAGEKKKK 1713
>TC8711 weakly similar to UP|VCLC_PEA (P13918) Vicilin precursor, partial
(74%)
Length = 1591
Score = 28.5 bits (62), Expect = 2.8
Identities = 12/26 (46%), Positives = 16/26 (61%), Gaps = 1/26 (3%)
Frame = +3
Query: 252 RTLKG-RGYILCCISLLYRWFISHLP 276
R LK RG++ CC + WF+S LP
Sbjct: 42 RVLKSKRGFLCCCCWEFFSWFLSPLP 119
>TC18660
Length = 893
Score = 27.3 bits (59), Expect = 6.3
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = +2
Query: 408 EEIGMPYPAMRYVSSSTPSIPLP 430
E MP P+M ++S +TP + LP
Sbjct: 800 EAFDMPTPSMEWISGTTPHLKLP 868
>TC11869
Length = 753
Score = 26.9 bits (58), Expect = 8.2
Identities = 14/43 (32%), Positives = 20/43 (45%)
Frame = +1
Query: 209 LLIYGLILFPNLDNFVDMNAIEVFHSKNPVPTLLADTYHAIHD 251
LL+ G + FV ++V H +PTLLA A+ D
Sbjct: 7 LLLPGQSVLVETSGFVIQQTLQVAHMVTVIPTLLASRVAAVSD 135
>TC11706 homologue to UP|Q8GSM9 (Q8GSM9) Squalene monooxygenase 2 , partial
(15%)
Length = 472
Score = 26.9 bits (58), Expect = 8.2
Identities = 15/51 (29%), Positives = 21/51 (40%)
Frame = +1
Query: 220 LDNFVDMNAIEVFHSKNPVPTLLADTYHAIHDRTLKGRGYILCCISLLYRW 270
L FV + +F P L YH IH L+ +L S+L+ W
Sbjct: 172 LKGFVRCFFLPLFQLITEPPLLQNSEYHRIHFEMLRCSQDVLALFSILWNW 324
>TC14892 similar to UP|Q9Z0W6 (Q9Z0W6) Pax transcription activation domain
interacting protein PTIP, partial (3%)
Length = 1502
Score = 26.9 bits (58), Expect = 8.2
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -1
Query: 467 LDGRDEQKTHENLMLKKELAKARKELEEK 495
L +DEQ TH+NL K ++ +E+ K
Sbjct: 1193 LSPKDEQTTHQNLNFSKHVSHLAREITTK 1107
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,380,575
Number of Sequences: 28460
Number of extensions: 114430
Number of successful extensions: 598
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 597
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 597
length of query: 532
length of database: 4,897,600
effective HSP length: 95
effective length of query: 437
effective length of database: 2,193,900
effective search space: 958734300
effective search space used: 958734300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC130803.17


