

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
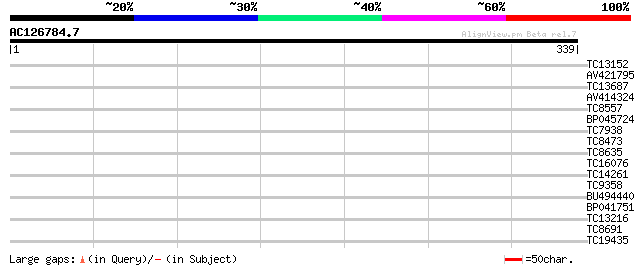
Query= AC126784.7 - phase: 0
(339 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial... 31 0.34
AV421795 30 0.58
TC13687 similar to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determin... 29 0.99
AV414324 29 0.99
TC8557 similar to UP|Q9LK52 (Q9LK52) Dbj|BAA90629.1, complete 28 2.2
BP045724 27 4.9
TC7938 homologue to UP|Q8LJW0 (Q8LJW0) 40S ribosomal S4 protein,... 27 4.9
TC8473 similar to UP|BAD11591 (BAD11591) 27k vesicle-associated ... 27 4.9
TC8635 similar to UP|Q8VYT9 (Q8VYT9) ER1-like receptor kinase (F... 27 6.4
TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 27 6.4
TC14261 weakly similar to UP|Q05929 (Q05929) EDGP precursor (Fra... 27 6.4
TC9358 weakly similar to GB|AAH10781.1|14789753|BC010781 RIO kin... 26 8.4
BU494440 26 8.4
BP041751 26 8.4
TC13216 UP|Q8LJW0 (Q8LJW0) 40S ribosomal S4 protein, partial (51%) 26 8.4
TC8691 similar to UP|Q940T5 (Q940T5) AT5g59550/f2o15_210, partia... 26 8.4
TC19435 similar to UP|Q39036 (Q39036) NADPH-ferrihemoprotein red... 26 8.4
>TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial (60%)
Length = 453
Score = 30.8 bits (68), Expect = 0.34
Identities = 20/76 (26%), Positives = 34/76 (44%), Gaps = 1/76 (1%)
Frame = +2
Query: 169 TSVKWA-SPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKH 227
T V+++ S AT + + WD PG + F G+ S V S+D HP+++
Sbjct: 98 TDVRFSPSMPRLATSSFDKTVKVWDVDNPGYSLRTFTGH------SASVMSLDFHPNKED 259
Query: 228 TCLAAGSLGTVLAWDL 243
+ S G + W +
Sbjct: 260 LICSCDSDGEIRYWSI 307
>AV421795
Length = 267
Score = 30.0 bits (66), Expect = 0.58
Identities = 21/64 (32%), Positives = 30/64 (46%), Gaps = 2/64 (3%)
Frame = +3
Query: 50 IHSFKPNPLSFEFQSSWTSPSP--ISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARL 107
+H +P P S+EF T PSP +SSI + SS +L F F+ S L
Sbjct: 45 LHLLQPIPSSYEFPDPATVPSP*LLSSING--------FKRANSSTTLSFNFSSSNSEPL 200
Query: 108 VSEV 111
V+ +
Sbjct: 201 VASI 212
>TC13687 similar to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determining MinD)
(At5g24020), partial (45%)
Length = 470
Score = 29.3 bits (64), Expect = 0.99
Identities = 14/28 (50%), Positives = 18/28 (64%)
Frame = +3
Query: 65 SWTSPSPISSIKSSQFLQKSIIASSTSS 92
+W SPSP S+ +SS S A+STSS
Sbjct: 129 TWASPSPASASRSSPSTPMSACATSTSS 212
>AV414324
Length = 380
Score = 29.3 bits (64), Expect = 0.99
Identities = 14/28 (50%), Positives = 20/28 (71%)
Frame = +3
Query: 301 DGILGVIEQGEEPIELLAEPCAINSFDI 328
DG+ G E G P+EL ++PC +N+FDI
Sbjct: 288 DGLFGACENGG-PVEL-SKPCNLNAFDI 365
>TC8557 similar to UP|Q9LK52 (Q9LK52) Dbj|BAA90629.1, complete
Length = 1064
Score = 28.1 bits (61), Expect = 2.2
Identities = 24/72 (33%), Positives = 34/72 (46%), Gaps = 2/72 (2%)
Frame = -2
Query: 31 NRFAVLATSDFDSNLSSIEIHSFKP--NPLSFEFQSSWTSPSPISSIKSSQFLQKSIIAS 88
N F +++ F + SS SF+ + LSF SS S + +SS SS S + +
Sbjct: 679 NGFPRRSSASFTLSSSSCSSRSFRSFSSALSFFICSSSASKASVSSSSSSSSSLSSSLLT 500
Query: 89 STSSGSLHFLFA 100
STS S L A
Sbjct: 499 STSESSASTLRA 464
>BP045724
Length = 490
Score = 26.9 bits (58), Expect = 4.9
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = +2
Query: 88 SSTSSGSLHFLFADSTDAR 106
+S+SS SLHFLF D D +
Sbjct: 326 ASSSSSSLHFLFTDEGDGK 382
>TC7938 homologue to UP|Q8LJW0 (Q8LJW0) 40S ribosomal S4 protein, complete
Length = 1142
Score = 26.9 bits (58), Expect = 4.9
Identities = 14/45 (31%), Positives = 25/45 (55%)
Frame = +2
Query: 128 LMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVK 172
L+DG V G +++V++ +N N+R L+D+ G SV+
Sbjct: 260 LVDGKVRTDKTYPSGFMDVVSIPKTNENFRLLYDTKGRFRLHSVR 394
>TC8473 similar to UP|BAD11591 (BAD11591) 27k vesicle-associated membrane
protein-associated protein-like, partial (81%)
Length = 1232
Score = 26.9 bits (58), Expect = 4.9
Identities = 17/38 (44%), Positives = 22/38 (57%), Gaps = 2/38 (5%)
Frame = -1
Query: 62 FQSSWTSPSPI--SSIKSSQFLQKSIIASSTSSGSLHF 97
F SS TSPSPI S+ FL+ S AS+++S F
Sbjct: 890 FHSSMTSPSPIILGPASSAGFLRASRAASASASCRFSF 777
>TC8635 similar to UP|Q8VYT9 (Q8VYT9) ER1-like receptor kinase (Fragment),
partial (87%)
Length = 752
Score = 26.6 bits (57), Expect = 6.4
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = -2
Query: 298 CSEDGILGVIEQGEEPIELLAEPCAI 323
C +L + EQ E+P LL +PC +
Sbjct: 277 CQLADVLRIREQAEKPFSLLPDPCML 200
>TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (24%)
Length = 1048
Score = 26.6 bits (57), Expect = 6.4
Identities = 14/66 (21%), Positives = 25/66 (37%)
Frame = +3
Query: 178 EFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGT 237
+ AT + WD P V ++ G+ S + S+D HP + S
Sbjct: 162 QLATASIDKSVRLWDAANPTYCVQEYNGH------SSAIMSLDFHPKKTDIFCFCDSANE 323
Query: 238 VLAWDL 243
+ W++
Sbjct: 324 IRYWNI 341
>TC14261 weakly similar to UP|Q05929 (Q05929) EDGP precursor (Fragment),
partial (84%)
Length = 1604
Score = 26.6 bits (57), Expect = 6.4
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +3
Query: 190 WWDQRKPGGPVSQFKG 205
WWDQ K GGP+ +G
Sbjct: 894 WWDQGKHGGPLHGSRG 941
>TC9358 weakly similar to GB|AAH10781.1|14789753|BC010781 RIO kinase 2 {Mus
musculus;} , partial (10%)
Length = 1117
Score = 26.2 bits (56), Expect = 8.4
Identities = 25/72 (34%), Positives = 35/72 (47%), Gaps = 4/72 (5%)
Frame = -3
Query: 24 LPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPN----PLSFEFQSSWTSPSPISSIKSSQ 79
+P+LS + + SD SS+ I SF+ + P EF+S PS I S+
Sbjct: 632 IPLLSCSS-----SCSDKYEKSSSLMIASFRSSTESTPSLSEFESLSGVPSMNL*ISSAS 468
Query: 80 FLQKSIIASSTS 91
FL K + ASS S
Sbjct: 467 FLVKPLAASSLS 432
>BU494440
Length = 508
Score = 26.2 bits (56), Expect = 8.4
Identities = 17/43 (39%), Positives = 24/43 (55%), Gaps = 3/43 (6%)
Frame = -2
Query: 39 SDFDS---NLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSS 78
SDF S N S+ + S +P P F + +SPSP SS+ +S
Sbjct: 249 SDFASTCFNTSNRRLSSKQPRPGP*IFPQTSSSPSPSSSVSAS 121
>BP041751
Length = 471
Score = 26.2 bits (56), Expect = 8.4
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +3
Query: 41 FDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPI 72
+ +L+S++I S KP PLSF S + S I
Sbjct: 60 YPHSLTSLKIKSPKPKPLSFSLMESSIAASRI 155
>TC13216 UP|Q8LJW0 (Q8LJW0) 40S ribosomal S4 protein, partial (51%)
Length = 450
Score = 26.2 bits (56), Expect = 8.4
Identities = 14/45 (31%), Positives = 25/45 (55%)
Frame = +1
Query: 128 LMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVK 172
L+DG V G +++V++ +N N+R L+D+ G SV+
Sbjct: 256 LVDGKVRTDKTYPAGFMDVVSIPKTNENFRLLYDTKGRFRLHSVR 390
>TC8691 similar to UP|Q940T5 (Q940T5) AT5g59550/f2o15_210, partial (16%)
Length = 942
Score = 26.2 bits (56), Expect = 8.4
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = -1
Query: 147 VTVGDSNLNYRRLFDSGGLVSYTSVKWASP 176
+TVG++ L R F + YTS KW SP
Sbjct: 270 LTVGETLLGARFEFTPPSISVYTSGKWLSP 181
>TC19435 similar to UP|Q39036 (Q39036) NADPH-ferrihemoprotein reductase ,
partial (12%)
Length = 459
Score = 26.2 bits (56), Expect = 8.4
Identities = 17/40 (42%), Positives = 19/40 (47%)
Frame = +3
Query: 55 PNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGS 94
P P S SSW+SP P S S ASS SSG+
Sbjct: 174 PAPSSSRMSSSWSSPPPSPS---------SSAASSFSSGA 266
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,268,732
Number of Sequences: 28460
Number of extensions: 88796
Number of successful extensions: 571
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 569
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 571
length of query: 339
length of database: 4,897,600
effective HSP length: 91
effective length of query: 248
effective length of database: 2,307,740
effective search space: 572319520
effective search space used: 572319520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC126784.7


