

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
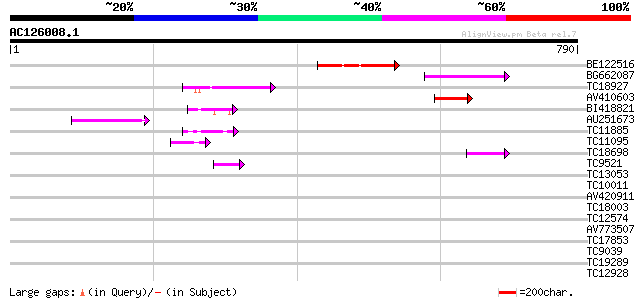
Query= AC126008.1 - phase: 0 /pseudo
(790 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE122516 96 2e-20
BG662087 91 5e-19
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 75 4e-14
AV410603 70 1e-12
BI418821 56 2e-08
AU251673 55 4e-08
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 54 7e-08
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 43 2e-04
TC18698 43 2e-04
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 42 4e-04
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 40 0.001
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 40 0.001
AV420911 39 0.002
TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 39 0.004
TC12574 36 0.027
AV773507 32 0.39
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 32 0.39
TC9039 32 0.51
TC19289 similar to GB|AAK15470.1|13194578|AF308596 endoribonucle... 30 1.1
TC12928 30 1.5
>BE122516
Length = 364
Score = 96.3 bits (238), Expect = 2e-20
Identities = 55/114 (48%), Positives = 74/114 (64%)
Frame = +2
Query: 430 GMDWLLAFGVSINCLTKSVTFSKPVEELGREFLTAEQVKKSLDGEACVFMMFASLKVSGE 489
GM+WL A ++NC K+VTF + R T ++V K+ + E+ V + S +
Sbjct: 17 GMNWLTANDATLNCRKKTVTFGTSEGDAKRVKRT-DKVGKASECESDVLLGALETDKS-D 190
Query: 490 KGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASEL 543
G +PVV+EF +VFPE+++ELP EREVEF+ID VPGT PISIAPYRMS EL
Sbjct: 191 TGVEGIPVVREFSDVFPEEVSELPPEREVEFSID*VPGTGPISIAPYRMSLVEL 352
>BG662087
Length = 373
Score = 91.3 bits (225), Expect = 5e-19
Identities = 47/119 (39%), Positives = 67/119 (55%)
Frame = +1
Query: 578 GSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISK 637
G R+ VDY LNK K+ YPLP ID +D E+ S +D SGYHQI++ D K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 638 TAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEH 696
TAF T +Y Y +PFG+ NA + M+R+F + R + V++D+++V S H
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 75.1 bits (183), Expect = 4e-14
Identities = 46/144 (31%), Positives = 68/144 (46%), Gaps = 14/144 (9%)
Frame = -2
Query: 241 KPYSKDKGKKRESGGGS-------RPSLAD---VKCYRCGTLGHYANECMNDLNCHKCGK 290
KP+ + + + SG RP+ +D + C+RC GH+AN C DL C C K
Sbjct: 434 KPFQRPQNRGTSSGYSHSFGNFVPRPTQSDTSEIVCHRCSKKGHFANRCP-DLVCWNCQK 258
Query: 291 AGHKAVDCR--VVAREITCYNCGEKGHISTKPKK--AAGKVFALNAKEVEQPDNLIRGMC 346
GH DC V + K K+ A+ +V+ ++ E + D LIR +
Sbjct: 257 TGHSGKDCTNPKVEAATNAIAARRPAPAANKGKRPVASARVYTVSGAESHRADGLIRSVG 78
Query: 347 FINSTPLIAIIDTGATHSFISASC 370
+N PL + D+GATHSFI +C
Sbjct: 77 SVNCKPLTILFDSGATHSFIDLAC 6
>AV410603
Length = 162
Score = 70.1 bits (170), Expect = 1e-12
Identities = 30/53 (56%), Positives = 42/53 (78%)
Frame = +1
Query: 593 TIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYG 645
T+K+ +P+P +D+ +D+L G++ FSK+DLRSGYHQI VK ED KT FRT +G
Sbjct: 4 TVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>BI418821
Length = 614
Score = 56.2 bits (134), Expect = 2e-08
Identities = 30/84 (35%), Positives = 37/84 (43%), Gaps = 14/84 (16%)
Frame = +2
Query: 248 GKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLN---------CHKCGKAGHKAVDC 298
G++R GGG CY CG GH A +C N C+ CG AGH A DC
Sbjct: 347 GERRNGGGGGGGGGG---CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDC 517
Query: 299 RVVARE-----ITCYNCGEKGHIS 317
CYNCG+ GH++
Sbjct: 518 NRSNNNSGGGGAGCYNCGDTGHLA 589
Score = 28.5 bits (62), Expect = 4.3
Identities = 19/79 (24%), Positives = 27/79 (34%), Gaps = 6/79 (7%)
Frame = +2
Query: 212 CRMFDDAGKAKSNYYKVQNEKKGKGH------GVGKPYSKDKGKKRESGGGSRPSLADVK 265
C D G + ++ N G G G ++D + + GG
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDCNRSNNNSGGGGAG----- 556
Query: 266 CYRCGTLGHYANECMNDLN 284
CY CG GH A +C N
Sbjct: 557 CYNCGDTGHLARDCNRSNN 613
>AU251673
Length = 413
Score = 55.1 bits (131), Expect = 4e-08
Identities = 34/111 (30%), Positives = 56/111 (49%), Gaps = 1/111 (0%)
Frame = +2
Query: 86 DDQKVRLATHMFAEEAEYWWINAKGRLELGGDVVTWAKFKVEFLRKYFPEDLRTKKEVEF 145
D + V LA+ A W+ +G TWA F EF+ ++ P+ +R +F
Sbjct: 11 DTRAVELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRDF 190
Query: 146 LNLKQGI-MSVAEYAAKFEELSRFCPYINAEDAMVSKCVKFESGLRPVIYQ 195
L+Q M+V+EY+A F LSR+ PY E+ V + V+ GL+ +++
Sbjct: 191 ERLEQAEGMTVSEYSAHFTHLSRYVPYPLLEEERVKRFVR---GLKEYLFK 334
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 54.3 bits (129), Expect = 7e-08
Identities = 29/77 (37%), Positives = 39/77 (49%)
Frame = +2
Query: 242 PYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVV 301
PY +D + G SR +L C C GH+A EC N CH CG GH A +C
Sbjct: 299 PYRRDSRR-----GFSRDNL----CKNCKRPGHFARECPNVAICHNCGLPGHIASEC--- 442
Query: 302 AREITCYNCGEKGHIST 318
+ C+NC E GH+++
Sbjct: 443 TTKSLCWNCKEPGHMAS 493
Score = 37.4 bits (85), Expect = 0.009
Identities = 14/33 (42%), Positives = 17/33 (51%)
Frame = +2
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDC 298
C+ CG GH A+EC C C + GH A C
Sbjct: 401 CHNCGLPGHIASECTTKSLCWNCKEPGHMASSC 499
Score = 34.3 bits (77), Expect = 0.078
Identities = 19/50 (38%), Positives = 25/50 (50%)
Frame = +2
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKAAGKVFALNAKE 334
C C + GH A +C VA C+NCG GHI++ + K N KE
Sbjct: 344 CKNCKRPGHFARECPNVA---ICHNCGLPGHIAS---ECTTKSLCWNCKE 475
Score = 25.8 bits (55), Expect(2) = 0.24
Identities = 8/13 (61%), Positives = 10/13 (76%)
Frame = +1
Query: 286 HKCGKAGHKAVDC 298
H CGK GH+A +C
Sbjct: 517 HTCGKVGHRAREC 555
Score = 25.4 bits (54), Expect(2) = 0.24
Identities = 10/25 (40%), Positives = 12/25 (48%)
Frame = +2
Query: 266 CYRCGTLGHYANECMNDLNCHKCGK 290
C+ C GH A+ C ND C K
Sbjct: 458 CWNCKEPGHMASSCPNDGICTPVAK 532
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 43.1 bits (100), Expect = 2e-04
Identities = 21/56 (37%), Positives = 29/56 (51%)
Frame = +1
Query: 224 NYYKVQNEKKGKGHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANEC 279
N ++V+ +G G G + G+ GGGS D+KCY CG GH+A EC
Sbjct: 268 NGWRVELSHNSRGGGGGGGGGRGGGRSGGGGGGS-----DMKCYECGEPGHFAREC 420
>TC18698
Length = 808
Score = 42.7 bits (99), Expect = 2e-04
Identities = 20/60 (33%), Positives = 33/60 (54%)
Frame = -2
Query: 637 KTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEH 696
KT + +Y Y VMP G+ N + M++IFH + + V V+++D++V S E H
Sbjct: 801 KTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*FH 622
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 42.0 bits (97), Expect = 4e-04
Identities = 18/43 (41%), Positives = 23/43 (52%)
Frame = +1
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKAAGKV 327
C CG GH A DC+ + CY CGE+GHI K + K+
Sbjct: 391 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKL 519
Score = 29.3 bits (64), Expect = 2.5
Identities = 10/17 (58%), Positives = 10/17 (58%)
Frame = +1
Query: 265 KCYRCGTLGHYANECMN 281
KCYRCG GH C N
Sbjct: 454 KCYRCGERGHIEKNCKN 504
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 40.4 bits (93), Expect = 0.001
Identities = 17/36 (47%), Positives = 20/36 (55%), Gaps = 1/36 (2%)
Frame = +3
Query: 264 VKCYRCGTLGHYANECMNDLN-CHKCGKAGHKAVDC 298
V C C LGH + +CM L CH CG GH A +C
Sbjct: 3 VVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 110
Score = 33.1 bits (74), Expect = 0.17
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = +3
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGHIS 317
C C + GH + DC + + C+NCG +GH++
Sbjct: 9 CRNCQQLGHMSRDC--MGPLMICHNCGGRGHLA 101
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 40.4 bits (93), Expect = 0.001
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = +2
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKAAGK 326
C CG GH A DC+ + CY CG++GH+ K + K
Sbjct: 371 CFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCKNSPKK 496
Score = 29.3 bits (64), Expect = 2.5
Identities = 10/17 (58%), Positives = 10/17 (58%)
Frame = +2
Query: 265 KCYRCGTLGHYANECMN 281
KCYRCG GH C N
Sbjct: 434 KCYRCGDRGHVERNCKN 484
>AV420911
Length = 418
Score = 39.3 bits (90), Expect = 0.002
Identities = 28/90 (31%), Positives = 38/90 (42%)
Frame = +2
Query: 190 RPVIYQYMCVQEIRDFDTLVHKCRMFDDAGKAKSNYYKVQNEKKGKGHGVGKPYSKDKGK 249
RP Y ++ + RD +H GK N ++V+ KG G G+
Sbjct: 206 RPPGYAFLEFDDKRDALDAIHALD-----GK---NGWRVELSHNSKGGGGGR-------- 337
Query: 250 KRESGGGSRPSLADVKCYRCGTLGHYANEC 279
GGG D+KCY CG GH+A EC
Sbjct: 338 ----GGGRGRGGEDLKCYECGEPGHFAREC 415
Score = 29.6 bits (65), Expect = 1.9
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = +2
Query: 282 DLNCHKCGKAGHKAVDCR 299
DL C++CG+ GH A +CR
Sbjct: 365 DLKCYECGEPGHFARECR 418
>TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (42%)
Length = 587
Score = 38.5 bits (88), Expect = 0.004
Identities = 20/56 (35%), Positives = 26/56 (45%)
Frame = +1
Query: 224 NYYKVQNEKKGKGHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANEC 279
N ++V+ KG G G+ GGG D+KCY CG GH+A EC
Sbjct: 76 NGWRVELSHNSKGGGGGR------------GGGRGRGGEDLKCYECGEPGHFAREC 207
Score = 29.6 bits (65), Expect = 1.9
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = +1
Query: 282 DLNCHKCGKAGHKAVDCR 299
DL C++CG+ GH A +CR
Sbjct: 157 DLKCYECGEPGHFARECR 210
>TC12574
Length = 325
Score = 35.8 bits (81), Expect = 0.027
Identities = 14/33 (42%), Positives = 23/33 (69%)
Frame = +2
Query: 663 FMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEE 695
F +N IF + + F++VFI+DIL Y++ +EE
Sbjct: 2 FKNSVNHIFESFFEHFMIVFINDILSYTEDKEE 100
>AV773507
Length = 496
Score = 32.0 bits (71), Expect = 0.39
Identities = 23/105 (21%), Positives = 45/105 (41%), Gaps = 9/105 (8%)
Frame = +2
Query: 203 RDFDTLVHKCRMFDDAGKAKSNYYKVQNEKKGKGHGVGKPYSKDKGKKRESGGGSRPSLA 262
RD L + M D G+ + + +++G + + + ++ G + S ++P +
Sbjct: 77 RDGKILECQIMMERDTGRPRGFGFITFADRRGMEDAIKEMHGREIGDRIISVNKAQPKMG 256
Query: 263 DVKC--------YRCGTLGHY-ANECMNDLNCHKCGKAGHKAVDC 298
Y G G Y A + + +C +CG+ GH+A DC
Sbjct: 257 GDDADQGYRGGGYSSGGRGSYGAGDRVGQDDCFECGRPGHRARDC 391
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 32.0 bits (71), Expect = 0.39
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 2/39 (5%)
Frame = +2
Query: 246 DKGKKRESGG--GSRPSLADVKCYRCGTLGHYANECMND 282
D+G + S G GSR S +C++CG GH+A EC ++
Sbjct: 356 DRGDRDRSRGYGGSRGSNGG-ECFKCGKPGHFARECPSE 469
Score = 30.8 bits (68), Expect = 0.87
Identities = 19/64 (29%), Positives = 25/64 (38%)
Frame = +2
Query: 235 KGHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLNCHKCGKAGHK 294
K G +D G + G R D R + G+ + N C KCGK GH
Sbjct: 281 KAQPQGSSRDRDDGDRYRERGRDRDDRGD----RDRSRGYGGSRGSNGGECFKCGKPGHF 448
Query: 295 AVDC 298
A +C
Sbjct: 449 AREC 460
>TC9039
Length = 1218
Score = 31.6 bits (70), Expect = 0.51
Identities = 30/144 (20%), Positives = 56/144 (38%), Gaps = 11/144 (7%)
Frame = +2
Query: 147 NLKQGIMSVAEYAAKFEELSRFCPYINAEDAMVSKCV--KFESGLRPVIYQYMCVQEIRD 204
N+++ IM ++ A+K + L + D ++ V + Y C +E
Sbjct: 503 NIREYIMGMSNIASKLKALK-----LELSDDLLIHLVLLSLPAQFSQFKISYNCPKEKWS 667
Query: 205 FDTLVHKCRMFDDAGKAKSNYYKVQNEKKGKGHGVGKPYSKDKGKKRES---------GG 255
+ L+ C ++ +++ E+K H V SKDKGK++++
Sbjct: 668 LNELISFCVQEEE---------RLKQERKESAHFVST--SKDKGKRKKTVEPKNEAADAP 814
Query: 256 GSRPSLADVKCYRCGTLGHYANEC 279
+ D CY C GH +C
Sbjct: 815 APKKQKEDDTCYFCNVSGHMKKKC 886
>TC19289 similar to GB|AAK15470.1|13194578|AF308596 endoribonuclease/protein
kinase IRE1-like protein {Arabidopsis thaliana;},
partial (18%)
Length = 470
Score = 30.4 bits (67), Expect = 1.1
Identities = 19/57 (33%), Positives = 30/57 (52%), Gaps = 3/57 (5%)
Frame = +1
Query: 706 VLKENQLCAKLSK---CEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFE 759
++KE LCAKLS + LK++S LGH + GG S W++P+ + +
Sbjct: 7 IIKERSLCAKLSDMGISKRLLKDMSSLGHNATGGGSS----------GWQAPEQLVQ 147
>TC12928
Length = 755
Score = 30.0 bits (66), Expect = 1.5
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = +3
Query: 38 RNEGEAEERRLDRFLRNNPPTFKGRFDPEGAQTWVQGM 75
R+EGE R+D F+ + P RFD A TW GM
Sbjct: 6 RHEGETSFHRMDMFINFSDPFIAKRFD-VNACTWAFGM 116
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,864,030
Number of Sequences: 28460
Number of extensions: 173295
Number of successful extensions: 827
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 754
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 804
length of query: 790
length of database: 4,897,600
effective HSP length: 98
effective length of query: 692
effective length of database: 2,108,520
effective search space: 1459095840
effective search space used: 1459095840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126008.1


