

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125473.10 + phase: 0
(257 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
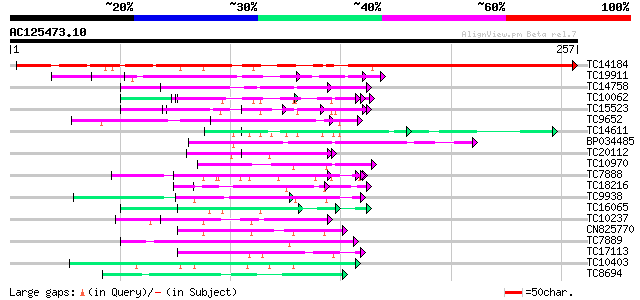
Sequences producing significant alignments: (bits) Value
TC14184 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-l... 279 3e-76
TC19911 weakly similar to PIR|A44805|A44805 eggshell protein pre... 69 6e-13
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 60 3e-10
TC10062 similar to BAD08700 (BAD08700) Cold shock domain protein... 56 7e-09
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 55 1e-08
TC9652 weakly similar to UP|AAP46157 (AAP46157) Latex abundant p... 49 7e-07
TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding p... 47 2e-06
BP034485 46 7e-06
TC20112 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich... 45 9e-06
TC10970 similar to GB|AAA32722.1|166570|ATH18RAB glycine rich pr... 45 9e-06
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 45 1e-05
TC18216 similar to GB|CAA28991.1|34041|HSKER65A keratin type II ... 45 1e-05
TC9938 weakly similar to GB|AAM97131.1|22531255|AY136466 ribonuc... 45 2e-05
TC16065 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich... 44 2e-05
TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precu... 43 6e-05
CN825770 42 1e-04
TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 42 1e-04
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 40 3e-04
TC10403 similar to UP|O80815 (O80815) T8F5.22 protein, partial (... 40 4e-04
TC8694 similar to UP|IF38_MEDTR (Q9XHM1) Eukaryotic translation ... 40 4e-04
>TC14184 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-like
protein, partial (57%)
Length = 1141
Score = 279 bits (714), Expect = 3e-76
Identities = 164/271 (60%), Positives = 188/271 (68%), Gaps = 17/271 (6%)
Frame = +3
Query: 4 EENRSRGLFHHKKHDDDDNRPIDTDSGYNKTSSYSTGDD---DYN---KKTSYGNDNSGG 57
EENR RGLFHH K++D RP DT++GYNKTS YST + DY+ KK+SY + SGG
Sbjct: 87 EENRDRGLFHHHKNED---RPTDTETGYNKTS-YSTDEPSGGDYDSGYKKSSY--ETSGG 248
Query: 58 DYETGYNKNTDNYSNNETS---GGYGSGGAKT--TTGGGYGGGYGDSDTKTTTGGGY--G 110
YETGYNK T +YS +E + GGY SG KT +T GGGYG GGGY G
Sbjct: 249 GYETGYNK-TSSYSTDEPNSGGGGYDSGYNKTAYSTDEPSGGGYGTGGA----GGGYSTG 413
Query: 111 GGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDD----RRE 166
GGYG TGGG GG Y +TGGG GGGY +T GGGYG+D RR+
Sbjct: 414 GGYG-------TGGGGGGEY-------STGGG-GGGY------STGGGGYGNDDSGNRRD 530
Query: 167 DVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGS 226
+VDY++EEK H K EHLGELG AAG +AL+EKHKSEKDPENAHRHK+EEEVAA AAVG+
Sbjct: 531 EVDYKEEEKHHKKLEHLGELGTTAAGAYALHEKHKSEKDPENAHRHKLEEEVAAAAAVGA 710
Query: 227 GGFAFHEHHEKKETKEEEEEAHGKKKHHFFG 257
GGFAFHEHHEKKE KEEEEE+HGKK HH FG
Sbjct: 711 GGFAFHEHHEKKEAKEEEEESHGKKHHHLFG 803
>TC19911 weakly similar to PIR|A44805|A44805 eggshell protein precursor -
fluke (Schistosoma haematobium) {Schistosoma
haematobium;} , partial (24%)
Length = 350
Score = 69.3 bits (168), Expect = 6e-13
Identities = 46/118 (38%), Positives = 60/118 (49%)
Frame = +1
Query: 53 DNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGG 112
+NS G+ TG++ + +++ S G TTG G GG D D ++ GG GGG
Sbjct: 13 ENSAGN--TGHSGGGSHGNDHRHSDNEHGGRRSETTGHGLSGGGYDDDNRS--GGRTGGG 180
Query: 113 YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDY 170
YGD GG GGYGD ++ GG GGGYGD ++ GGYGD R V Y
Sbjct: 181 YGD-------GGHSAGGYGDHGGRS---GGGGGGYGDGGL-SSGSGGYGDGGRSGVGY 321
Score = 65.1 bits (157), Expect = 1e-11
Identities = 45/125 (36%), Positives = 61/125 (48%)
Frame = +1
Query: 38 STGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYG 97
S G+ ++ S+GND+ D E G + ++ + + GGY +GG GGGYG
Sbjct: 19 SAGNTGHSGGGSHGNDHRHSDNEHG-GRRSETTGHGLSGGGYDDDNR---SGGRTGGGYG 186
Query: 98 DSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTG 157
D GG GGYGD ++ GG GGGYGD + + GGYGD +G
Sbjct: 187 D-------GGHSAGGYGDHGGRS---GGGGGGYGDGGLSSGS-----GGYGDGG---RSG 312
Query: 158 GGYGD 162
GYGD
Sbjct: 313 VGYGD 327
Score = 47.4 bits (111), Expect = 2e-06
Identities = 38/114 (33%), Positives = 51/114 (44%), Gaps = 1/114 (0%)
Frame = +1
Query: 20 DDNRPIDTDSGYNKTSSYSTGDDDYNKKTSYGNDN-SGGDYETGYNKNTDNYSNNETSGG 78
+D+R D + G ++ + G Y +DN SGG GY ++GG
Sbjct: 61 NDHRHSDNEHGGRRSETTGHGLSG----GGYDDDNRSGGRTGGGYG------DGGHSAGG 210
Query: 79 YGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGD 132
YG G ++ GG GGGYGD + + GGYGD GG G GYGD
Sbjct: 211 YGDHGGRS---GGGGGGYGDGGLSSGS-----GGYGD-------GGRSGVGYGD 327
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 60.5 bits (145), Expect = 3e-10
Identities = 42/96 (43%), Positives = 46/96 (47%)
Frame = +1
Query: 69 NYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGG 128
N + NE G GG GGGYGGG G + GGYGGG G S GGGYGG
Sbjct: 292 NITVNEAQARGGGGGGGGRGGGGYGGGGGRRE------GGYGGG-GGSRGYGGGGGGYGG 450
Query: 129 GYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDR 164
G G GGYGG G S + GGG G+ R
Sbjct: 451 G-GGGGYGGRREGGYGGDGGGSRYGSGGGGGGGNWR 555
Score = 52.4 bits (124), Expect = 8e-08
Identities = 37/96 (38%), Positives = 40/96 (41%)
Frame = +1
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG 110
G GG Y G + Y S GYG G GGGYGGG GGGYG
Sbjct: 337 GGGRGGGGYGGGGGRREGGYGGGGGSRGYGGG------GGGYGGG---------GGGGYG 471
Query: 111 GGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 146
G GGYGG G S + +GGG GGG
Sbjct: 472 G---------RREGGYGGDGGGS--RYGSGGGGGGG 546
Score = 28.1 bits (61), Expect = 1.6
Identities = 18/49 (36%), Positives = 21/49 (42%), Gaps = 2/49 (4%)
Frame = +1
Query: 49 SYGNDNSGGDYETGYNKNTDNYSNNETSGGYG--SGGAKTTTGGGYGGG 95
S G GG GY GGYG GG++ +GGG GGG
Sbjct: 412 SRGYGGGGG----GYGGGGGGGYGGRREGGYGGDGGGSRYGSGGGGGGG 546
>TC10062 similar to BAD08700 (BAD08700) Cold shock domain protein 2, partial
(66%)
Length = 481
Score = 55.8 bits (133), Expect = 7e-09
Identities = 42/88 (47%), Positives = 42/88 (47%), Gaps = 3/88 (3%)
Frame = +3
Query: 77 GGYGSGGAKTTTGGGYGGG-YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGG--GYGDS 133
GG G GG GGGYGGG YG GGGYGGG GGGYGG GYG S
Sbjct: 282 GGAGGGGRG---GGGYGGGGYG--------GGGYGGG-------GRGGGGYGGGGGYGGS 407
Query: 134 DTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
G G GGGYG GGG G
Sbjct: 408 G-----GYGGGGGYGGGGRGGGRGGGGG 476
Score = 53.5 bits (127), Expect = 3e-08
Identities = 41/87 (47%), Positives = 41/87 (47%), Gaps = 2/87 (2%)
Frame = +3
Query: 77 GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGY--GGGYGDSDTKTTTGGGYGGGYGDSD 134
GGYG GG GGGYGGG GGGY GGGYG S GGYGGG G
Sbjct: 312 GGYGGGGYG---GGGYGGG-------GRGGGGYGGGGGYGGS-------GGYGGGGGYG- 437
Query: 135 TKTTTGGGYGGGYGDSDTKTTTGGGYG 161
GGG GGG G GGG G
Sbjct: 438 -----GGGRGGGRG--------GGGGG 479
Score = 53.5 bits (127), Expect = 3e-08
Identities = 42/95 (44%), Positives = 43/95 (45%), Gaps = 3/95 (3%)
Frame = +3
Query: 74 ETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGG-YGDSDTKTTTGGGYGGGYGD 132
E +G G T GGG GGG GGGYGGG YG GGGYGGG G
Sbjct: 234 EVTGPNGQAVQGTRRGGGAGGG-------GRGGGGYGGGGYG--------GGGYGGG-GR 365
Query: 133 SDTKTTTGGGYG--GGYGDSDTKTTTGGGYGDDRR 165
GGGYG GGYG GGGYG R
Sbjct: 366 GGGGYGGGGGYGGSGGYGG-------GGGYGGGGR 449
Score = 53.1 bits (126), Expect = 5e-08
Identities = 39/86 (45%), Positives = 41/86 (47%), Gaps = 2/86 (2%)
Frame = +3
Query: 76 SGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG--GGYGDS 133
+GG G GG GG GGGYG GG GGGYG GGGYG GGYG
Sbjct: 288 AGGGGRGGGGYGGGGYGGGGYGG-------GGRGGGGYGG-------GGGYGGSGGYGG- 422
Query: 134 DTKTTTGGGYGGGYGDSDTKTTTGGG 159
GGGYGGG G + GGG
Sbjct: 423 ------GGGYGGG-GRGGGRGGGGGG 479
Score = 41.2 bits (95), Expect = 2e-04
Identities = 30/81 (37%), Positives = 31/81 (38%)
Frame = +3
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG 110
G GG Y G GGYG GG +GG YGG GGGYG
Sbjct: 294 GGGRGGGGYGGGGYGGGGYGGGGRGGGGYGGGGGYGGSGG-YGG-----------GGGYG 437
Query: 111 GGYGDSDTKTTTGGGYGGGYG 131
GG GGG GGG G
Sbjct: 438 GG--------GRGGGRGGGGG 476
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 55.1 bits (131), Expect = 1e-08
Identities = 39/97 (40%), Positives = 47/97 (48%), Gaps = 6/97 (6%)
Frame = +2
Query: 72 NNETSGGYGSGGAKTTTGGGYGG-----GYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGY 126
N + G G GG GGGYGG GYG +GGG GGGYG S GGGY
Sbjct: 578 NEAQARGRGGGGG----GGGYGGGRREGGYGGGGGSRYSGGG-GGGYGGSG-----GGGY 727
Query: 127 GGGYGDSDTKTTTGGGYGGG-YGDSDTKTTTGGGYGD 162
GG + GGGYGG G ++ ++GG + D
Sbjct: 728 GG------RREGGGGGYGGSREGGGGSRYSSGGSWRD 820
Score = 52.4 bits (124), Expect = 8e-08
Identities = 31/75 (41%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Frame = +2
Query: 70 YSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 129
Y GGYG GG +GGG GGGYG S GGGYGG + GGGYGG
Sbjct: 626 YGGGRREGGYGGGGGSRYSGGG-GGGYGGSG-----GGGYGG------RREGGGGGYGGS 769
Query: 130 -YGDSDTKTTTGGGY 143
G ++ ++GG +
Sbjct: 770 REGGGGSRYSSGGSW 814
Score = 47.8 bits (112), Expect = 2e-06
Identities = 30/61 (49%), Positives = 32/61 (52%), Gaps = 2/61 (3%)
Frame = +2
Query: 106 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG--GGYGDSDTKTTTGGGYGDD 163
GGG GGGYG + GGYGGG G S GGGYG GG G + GGGYG
Sbjct: 605 GGGGGGGYGGGRRE----GGYGGG-GGSRYSGGGGGGYGGSGGGGYGGRREGGGGGYGGS 769
Query: 164 R 164
R
Sbjct: 770 R 772
Score = 39.7 bits (91), Expect = 5e-04
Identities = 26/77 (33%), Positives = 34/77 (43%), Gaps = 1/77 (1%)
Frame = +2
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG 110
G GG E GY + + GGYG G GGGYGG + GGGYG
Sbjct: 617 GGGYGGGRREGGYGGGGGSRYSGGGGGGYGGSG-----GGGYGG------RREGGGGGYG 763
Query: 111 GG-YGDSDTKTTTGGGY 126
G G ++ ++GG +
Sbjct: 764 GSREGGGGSRYSSGGSW 814
>TC9652 weakly similar to UP|AAP46157 (AAP46157) Latex abundant protein 1,
partial (62%)
Length = 576
Score = 49.3 bits (116), Expect = 7e-07
Identities = 34/121 (28%), Positives = 52/121 (42%), Gaps = 2/121 (1%)
Frame = +3
Query: 29 SGYNKTSSYSTG--DDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKT 86
S + + S+Y+ D +Y+ + + + GY++ + Y + GYG
Sbjct: 228 SSHAEPSAYANEALDVEYSSYSRPKPRPAPASFNPGYDRPSSEYES-----GYGRKPESE 392
Query: 87 TTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 146
+G YG GYG TG YG GYG +++ YG GYG K + YG G
Sbjct: 393 YSGLEYGSGYGRKQEHQVTGSEYGSGYGGRKQESSE---YGSGYG---RKQESESEYGSG 554
Query: 147 Y 147
Y
Sbjct: 555 Y 557
Score = 47.0 bits (110), Expect = 3e-06
Identities = 25/70 (35%), Positives = 33/70 (46%), Gaps = 1/70 (1%)
Frame = +3
Query: 92 YGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG-DS 150
Y GYG +G YG GYG TG YG GYG +++ YG GYG
Sbjct: 357 YESGYGRKPESEYSGLEYGSGYGRKQEHQVTGSEYGSGYGGRKQESSE---YGSGYGRKQ 527
Query: 151 DTKTTTGGGY 160
++++ G GY
Sbjct: 528 ESESEYGSGY 557
Score = 32.7 bits (73), Expect = 0.063
Identities = 19/60 (31%), Positives = 24/60 (39%), Gaps = 4/60 (6%)
Frame = +3
Query: 126 YGGGYGDSDTKTTTGGGYGGGYGDSDTKTTT----GGGYGDDRREDVDYEKEEKRHSKHE 181
Y GYG +G YG GYG T G GYG ++E +Y R + E
Sbjct: 357 YESGYGRKPESEYSGLEYGSGYGRKQEHQVTGSEYGSGYGGRKQESSEYGSGYGRKQESE 536
>TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding protein,
partial (20%)
Length = 760
Score = 47.4 bits (111), Expect = 2e-06
Identities = 44/128 (34%), Positives = 50/128 (38%), Gaps = 34/128 (26%)
Frame = +3
Query: 89 GGGYGGGYGDSD----------TKTTTGG-----GYGGG----YGDSDTKTTTGGGYGGG 129
G GYGGGYG T+ TGG GYGGG YG+ G GYGG
Sbjct: 3 GSGYGGGYGGRSEEENPGYGRKTEYETGGSEYESGYGGGRKPSYGEEQ-----GSGYGGR 167
Query: 130 YGDSDTKTTTG-GGYGG----GYG----------DSDTKTTTGGGYGDDRREDVDYEKEE 174
+ K + G Y G YG D D + YGDD +D D K
Sbjct: 168 SEYEEKKPSYGRSNYEGQEEVEYGRERPPRLDDDDDDNEGYARKKYGDDGSDDDDERK-- 341
Query: 175 KRHSKHEH 182
K H KH H
Sbjct: 342 KHHHKHHH 365
Score = 42.0 bits (97), Expect = 1e-04
Identities = 43/162 (26%), Positives = 59/162 (35%), Gaps = 19/162 (11%)
Frame = +3
Query: 106 GGGYGGGYGDSD----------TKTTTGG-----GYGGG----YGDSDTKTTTGGGYGGG 146
G GYGGGYG T+ TGG GYGGG YG+ G GY GG
Sbjct: 3 GSGYGGGYGGRSEEENPGYGRKTEYETGGSEYESGYGGGRKPSYGEEQ-----GSGY-GG 164
Query: 147 YGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDP 206
+ + K + G + +E+V+Y +E + G A Y S+ D
Sbjct: 165 RSEYEEKKPSYGRSNYEGQEEVEYGRERPPRLDDDDDDNEGYAR----KKYGDDGSDDDD 332
Query: 207 ENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAH 248
E H H+HH +K +E + H
Sbjct: 333 ERKKHH-------------------HKHHHRKNYDDE*TKCH 401
>BP034485
Length = 485
Score = 45.8 bits (107), Expect = 7e-06
Identities = 40/137 (29%), Positives = 57/137 (41%), Gaps = 6/137 (4%)
Frame = +3
Query: 82 GGAKTTTGGGYGGGYGDSD------TKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDT 135
GG K+ G GGG GDS TK GGG+ + K+ GG GGG G++
Sbjct: 6 GGNKSKEGKQKGGGGGDSSSDKKKKTKDKNGGGFLVKFLGLGKKSKKGG--GGGGGETAN 179
Query: 136 KTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFA 195
K GG G++ K GG G + + +D++ ++ H G+ G GG
Sbjct: 180 KNKNKNKNNGG-GNNKGKEGKKGGGGGGKLDKIDFDFQDFDIPSH---GKSGGKGGGG-- 341
Query: 196 LYEKHKSEKDPENAHRH 212
K +N H H
Sbjct: 342 ------GGKGNKNGHDH 374
Score = 37.7 bits (86), Expect = 0.002
Identities = 36/132 (27%), Positives = 46/132 (34%), Gaps = 13/132 (9%)
Frame = +3
Query: 43 DYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGY-----GSGGAKTTTGGGYGGGYG 97
D K+ G GG ++ +K N GG+ G G GGG GG
Sbjct: 3 DGGNKSKEGKQKGGGGGDSSSDKKKKTKDKN--GGGFLVKFLGLGKKSKKGGGGGGGETA 176
Query: 98 DSDTKTTTGGGYGGGYGDSDTKTTTGGGYGG-------GYGDSDTKT-TTGGGYGGGYGD 149
+ + G G G K GGG GG + D D + GG GGG G
Sbjct: 177 NKNKNKNKNNGGGNNKGKEGKK---GGGGGGKLDKIDFDFQDFDIPSHGKSGGKGGGGGG 347
Query: 150 SDTKTTTGGGYG 161
K G+G
Sbjct: 348 KGNKNGHDHGHG 383
Score = 35.8 bits (81), Expect = 0.007
Identities = 35/120 (29%), Positives = 47/120 (39%), Gaps = 8/120 (6%)
Frame = +3
Query: 46 KKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGG-------GYGD 98
KK+ G GG+ NKN + N+ +GG + G + GGG GG + D
Sbjct: 132 KKSKKGGGGGGGETA---NKNKNK---NKNNGGGNNKGKEGKKGGGGGGKLDKIDFDFQD 293
Query: 99 SDTKT-TTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTG 157
D + GG GGG G K G+ G+G GG G G D + G
Sbjct: 294 FDIPSHGKSGGKGGGGGGKGNK----NGHDHGHG-------IGGNMNGNGGHMDPMSQLG 440
Score = 26.9 bits (58), Expect = 3.5
Identities = 22/95 (23%), Positives = 34/95 (35%), Gaps = 3/95 (3%)
Frame = +3
Query: 32 NKTSSYSTGDDDYNKKTSYGNDNSGGDYETG---YNKNTDNYSNNETSGGYGSGGAKTTT 88
NK + + + N K G GG + ++ + ++ SGG G GG
Sbjct: 177 NKNKNKNKNNGGGNNKGKEGKKGGGGGGKLDKIDFDFQDFDIPSHGKSGGKGGGGGGKGN 356
Query: 89 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTG 123
G+ G+G GG G G D + G
Sbjct: 357 KNGHDHGHG-------IGGNMNGNGGHMDPMSQLG 440
>TC20112 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich
glycoprotein DZ-HRGP precursor, partial (11%)
Length = 523
Score = 45.4 bits (106), Expect = 9e-06
Identities = 28/68 (41%), Positives = 33/68 (48%), Gaps = 2/68 (2%)
Frame = +2
Query: 81 SGGAKTTTGGGYGGGYGDS--DTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTT 138
SGG GGG+GG GD DT +GGG G G G + GG GGG+
Sbjct: 341 SGGFSVGRGGGFGGRGGDRKYDTGRYSGGGGGRGRGGGSIRGGGRGGRGGGF-------- 496
Query: 139 TGGGYGGG 146
GG +GGG
Sbjct: 497 -GGRHGGG 517
Score = 39.7 bits (91), Expect = 5e-04
Identities = 21/45 (46%), Positives = 25/45 (54%), Gaps = 2/45 (4%)
Frame = +2
Query: 106 GGGYGGGYGDS--DTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 148
GGG+GG GD DT +GGG G G G + GG GGG+G
Sbjct: 365 GGGFGGRGGDRKYDTGRYSGGGGGRGRGGGSIRGGGRGGRGGGFG 499
Score = 31.6 bits (70), Expect = 0.14
Identities = 19/41 (46%), Positives = 22/41 (53%), Gaps = 2/41 (4%)
Frame = +2
Query: 123 GGGYGGGYGDS--DTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
GGG+GG GD DT +GGG G G G + GGG G
Sbjct: 365 GGGFGGRGGDRKYDTGRYSGGGGGRGRGGGSIR---GGGRG 478
Score = 30.4 bits (67), Expect = 0.31
Identities = 17/48 (35%), Positives = 22/48 (45%), Gaps = 4/48 (8%)
Frame = -2
Query: 67 TDNYSNNETSGGYG----SGGAKTTTGGGYGGGYGDSDTKTTTGGGYG 110
T+N T+GG+G +G TG YG G T+ TGG G
Sbjct: 357 TENPPEGGTNGGFGLACAAGPKPLGTGAAYGAP*GTGPTRLATGGRIG 214
Score = 29.3 bits (64), Expect = 0.70
Identities = 27/85 (31%), Positives = 30/85 (34%), Gaps = 2/85 (2%)
Frame = -2
Query: 122 TGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG--DDRREDVDYEKEEKRHSK 179
T GG+G TG YG G T+ TGG G R E+ E R SK
Sbjct: 333 TNGGFGLACAAGPKPLGTGAAYGAP*GTGPTRLATGGRIGPITSLRRSRYEEESEYRLSK 154
Query: 180 HEHLGELGAAAAGGFALYEKHKSEK 204
GF L E SEK
Sbjct: 153 -------------GFML*EARVSEK 118
Score = 28.9 bits (63), Expect = 0.91
Identities = 17/41 (41%), Positives = 20/41 (48%), Gaps = 2/41 (4%)
Frame = +2
Query: 76 SGGYGSGGAKTTTGGG--YGGGYGDSDTKTTTGGGYGGGYG 114
+G Y GG GGG GGG G GGG+GG +G
Sbjct: 407 TGRYSGGGGGRGRGGGSIRGGGRGG------RGGGFGGRHG 511
Score = 27.7 bits (60), Expect = 2.0
Identities = 14/40 (35%), Positives = 16/40 (40%)
Frame = -2
Query: 105 TGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 144
T GG+G TG YG G T+ TGG G
Sbjct: 333 TNGGFGLACAAGPKPLGTGAAYGAP*GTGPTRLATGGRIG 214
Score = 27.7 bits (60), Expect = 2.0
Identities = 14/40 (35%), Positives = 16/40 (40%)
Frame = -2
Query: 88 TGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 127
T GG+G TG YG G T+ TGG G
Sbjct: 333 TNGGFGLACAAGPKPLGTGAAYGAP*GTGPTRLATGGRIG 214
Score = 27.7 bits (60), Expect = 2.0
Identities = 14/47 (29%), Positives = 20/47 (41%)
Frame = +2
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYG 97
G GGD + + + GG GG + GGG+GG +G
Sbjct: 371 GFGGRGGDRKYDTGRYSGGGGGRGRGGGSIRGGGRGGRGGGFGGRHG 511
Score = 26.2 bits (56), Expect = 5.9
Identities = 14/27 (51%), Positives = 16/27 (58%), Gaps = 2/27 (7%)
Frame = +2
Query: 140 GGGYGGGYGDS--DTKTTTGGGYGDDR 164
GGG+GG GD DT +GGG G R
Sbjct: 365 GGGFGGRGGDRKYDTGRYSGGGGGRGR 445
>TC10970 similar to GB|AAA32722.1|166570|ATH18RAB glycine rich protein
{Arabidopsis thaliana;} , partial (25%)
Length = 697
Score = 45.4 bits (106), Expect = 9e-06
Identities = 37/101 (36%), Positives = 43/101 (41%), Gaps = 20/101 (19%)
Frame = +2
Query: 86 TTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG-GGYGDSDTKTTTGGG-- 142
T TG GYG G +D TT G+G T TG GYG GG+G T TGGG
Sbjct: 2 TGTGTGYGTGGHGTDYGTTGTHGHG--------TTGTGTGYGTGGHGTEYGTTGTGGGVM 157
Query: 143 -----------------YGGGYGDSDTKTTTGGGYGDDRRE 166
YG G G + T TTT G +G+ E
Sbjct: 158 DKIKEKLPCAGSTATHEYGSG-GTTGTTTTTTGHHGEQHHE 277
Score = 36.2 bits (82), Expect = 0.006
Identities = 33/99 (33%), Positives = 40/99 (40%), Gaps = 2/99 (2%)
Frame = +2
Query: 48 TSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGG 107
T YG G DY T +G +G G T TG G GG+G T TGG
Sbjct: 14 TGYGTGGHGTDYGT--------------TGTHGHGTTGTGTGYG-TGGHGTEYGTTGTGG 148
Query: 108 GYGGGYGDS--DTKTTTGGGYGGGYGDSDTKTTTGGGYG 144
G + +T YG G G + T TTT G +G
Sbjct: 149 GVMDKIKEKLPCAGSTATHEYGSG-GTTGTTTTTTGHHG 262
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 45.1 bits (105), Expect = 1e-05
Identities = 39/98 (39%), Positives = 48/98 (48%), Gaps = 11/98 (11%)
Frame = -1
Query: 75 TSGGYGSGGAKTTTGGGY-GGGYGDSDTKT----TTGG-GYGGGYGDSDTKT-----TTG 123
T+GG+G+ G +TT G G GGG+ T T GG G GG+G T T G
Sbjct: 589 TTGGFGT*GGETTGGLGT*GGGFTTGGLGT*GGFTIGGLGTWGGFGT*GGFTIGGFGTWG 410
Query: 124 GGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
G GG+G TT G G GG+GD GGG+G
Sbjct: 409 GFTTGGFGT*GGLTTGGLG**GGFGDG-----LGGGHG 311
Score = 43.9 bits (102), Expect = 3e-05
Identities = 38/99 (38%), Positives = 44/99 (44%), Gaps = 11/99 (11%)
Frame = -1
Query: 75 TSGGYGSGGAKTTTGGGYG--GGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGD 132
T+GG G+ G T T GG+G GG T GG GG G T G G GG+G
Sbjct: 628 TTGGLGTYGGVTGTTGGFGT*GGETTGGLGT*GGGFTTGGLGT*GGFTIGGLGTWGGFGT 449
Query: 133 SDTKT-----TTGGGYGGGYGDSDTKTTTG----GGYGD 162
T T GG GG+G TT G GG+GD
Sbjct: 448 *GGFTIGGFGTWGGFTTGGFGT*GGLTTGGLG**GGFGD 332
Score = 42.4 bits (98), Expect = 8e-05
Identities = 36/100 (36%), Positives = 45/100 (45%)
Frame = -1
Query: 47 KTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTG 106
+T+ G GG + TG + T GG G+ G T GG GG+G T G
Sbjct: 559 ETTGGLGT*GGGFTTGGLGT*GGF----TIGGLGTWGGFGT*GGFTIGGFG------TWG 410
Query: 107 GGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 146
G GG+G TT G G GG+GD GGG+G G
Sbjct: 409 GFTTGGFGT*GGLTTGGLG**GGFGDG-----LGGGHGSG 305
Score = 41.2 bits (95), Expect = 2e-04
Identities = 35/94 (37%), Positives = 41/94 (43%), Gaps = 9/94 (9%)
Frame = -1
Query: 75 TSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSD 134
T+GG G+ G T GG GG G T GG GG+G T GG GG+G
Sbjct: 820 TTGGSGT*GGG*TIGGFTIGGVGT*GGG*TIGGFTIGGFGT*GGG*TMGGFTMGGFGT*G 641
Query: 135 TKTTTGGGYG---------GGYGDSDTKTTTGGG 159
T T GG G GG+G +TT G G
Sbjct: 640 GVTGTTGGLGTYGGVTGTTGGFGT*GGETTGGLG 539
Score = 41.2 bits (95), Expect = 2e-04
Identities = 35/87 (40%), Positives = 37/87 (42%)
Frame = -1
Query: 75 TSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSD 134
T GG G GG T G G GG G T GG GG G T GG GG+G
Sbjct: 868 TIGGVGVGGV-TIGGDGTTGGSGT*GGG*TIGGFTIGGVGT*GGG*TIGGFTIGGFGT*G 692
Query: 135 TKTTTGGGYGGGYGDSDTKTTTGGGYG 161
T GG GG+G T T GG G
Sbjct: 691 GG*TMGGFTMGGFGT*GGVTGTTGGLG 611
Score = 38.9 bits (89), Expect = 9e-04
Identities = 31/77 (40%), Positives = 36/77 (46%), Gaps = 5/77 (6%)
Frame = -1
Query: 75 TSGGYGSGGAKT-----TTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 129
T GG+G+ G T T GG GG+G TT G G GG+GD GGG+G G
Sbjct: 469 TWGGFGT*GGFTIGGFGTWGGFTTGGFGT*GGLTTGGLG**GGFGDG-----LGGGHGSG 305
Query: 130 YGDSDTKTTTGGGYGGG 146
D GGY GG
Sbjct: 304 P*D--------GGYFGG 278
Score = 38.1 bits (87), Expect = 0.002
Identities = 29/71 (40%), Positives = 34/71 (47%)
Frame = -1
Query: 75 TSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSD 134
T+GG+G+ G TT G G GG+GD GGG+G G D GGY GG
Sbjct: 403 TTGGFGT*GGLTTGGLG**GGFGDG-----LGGGHGSGP*D--------GGYFGGL*TGG 263
Query: 135 TKTTTGGGYGG 145
T GG GG
Sbjct: 262 --FTIGGCCGG 236
>TC18216 similar to GB|CAA28991.1|34041|HSKER65A keratin type II (AA1-215)
{Homo sapiens;} , partial (16%)
Length = 501
Score = 45.1 bits (105), Expect = 1e-05
Identities = 28/72 (38%), Positives = 37/72 (50%), Gaps = 1/72 (1%)
Frame = +2
Query: 75 TSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGG-GYGGGYGDS 133
TSGG+G + TGGG+G T GG+GGG+G + GG G GGG+G
Sbjct: 14 TSGGFGGAAS---TGGGFGAA------AAPTSGGFGGGFGAVPSAGGFGGAGSGGGFGAF 166
Query: 134 DTKTTTGGGYGG 145
+ GG+GG
Sbjct: 167 SNPGS--GGFGG 196
Score = 43.1 bits (100), Expect = 5e-05
Identities = 28/82 (34%), Positives = 38/82 (46%), Gaps = 1/82 (1%)
Frame = +2
Query: 84 AKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGG-G 142
A T GG+GG +TGGG+G T GG+GGG+G + GG G
Sbjct: 2 AAAPTSGGFGGA-------ASTGGGFGA------AAAPTSGGFGGGFGAVPSAGGFGGAG 142
Query: 143 YGGGYGDSDTKTTTGGGYGDDR 164
GGG+G + GG+G +
Sbjct: 143 SGGGFGAFSNPGS--GGFGGSK 202
>TC9938 weakly similar to GB|AAM97131.1|22531255|AY136466
ribonucleoprotein-like {Arabidopsis thaliana;}, partial
(9%)
Length = 633
Score = 44.7 bits (104), Expect = 2e-05
Identities = 34/95 (35%), Positives = 40/95 (41%), Gaps = 9/95 (9%)
Frame = +1
Query: 76 SGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGG---GYGD 132
SG YG G++ GGY G G + G+ G YG S ++ GGYGG YG
Sbjct: 25 SGAYGGYGSEFGGYGGYAGAMG--PYRGDPSLGFAGRYGGSFSRGYDLGGYGGASESYGA 198
Query: 133 SDTKTTTGGG------YGGGYGDSDTKTTTGGGYG 161
GGG Y GGY S GGYG
Sbjct: 199 YGAGAAAGGGSSAAGAYQGGYDAS-----LAGGYG 288
Score = 41.6 bits (96), Expect = 1e-04
Identities = 34/103 (33%), Positives = 41/103 (39%), Gaps = 3/103 (2%)
Frame = +1
Query: 30 GYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSG---GAKT 86
GY GD YG S G GY +++Y G YG+G G +
Sbjct: 70 GYAGAMGPYRGDPSLGFAGRYGGSFSRGYDLGGYGGASESY------GAYGAGAAAGGGS 231
Query: 87 TTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 129
+ G Y GGY S GGYGG G S + GGYG G
Sbjct: 232 SAAGAYQGGYDAS-----LAGGYGGASGGSFYGSR--GGYGAG 339
Score = 37.7 bits (86), Expect = 0.002
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 1/91 (1%)
Frame = +1
Query: 57 GDYETGY-NKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGD 115
GD G+ + ++S GGYG + + G YG G + ++ G Y GGY
Sbjct: 100 GDPSLGFAGRYGGSFSRGYDLGGYGGA---SESYGAYGAG-AAAGGGSSAAGAYQGGYDA 267
Query: 116 SDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 146
S GGYGG G S + GGYG G
Sbjct: 268 S-----LAGGYGGASGGSFYGSR--GGYGAG 339
>TC16065 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich
glycoprotein DZ-HRGP precursor, partial (20%)
Length = 608
Score = 44.3 bits (103), Expect = 2e-05
Identities = 35/94 (37%), Positives = 36/94 (38%), Gaps = 6/94 (6%)
Frame = -1
Query: 77 GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGG------YGDSDTKTTTGGGYGGGY 130
GG G GG GG GGGYG GGG GG YG G G
Sbjct: 347 GGRGYGGPPQ---GGRGGGYGGGGGPGYGGGGRGGHGVPQQQYGGPPEYQGRGRGGPSQQ 177
Query: 131 GDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDR 164
G + GGGYGGG G GGG G R
Sbjct: 176 GGRGGYSGGGGGYGGGGGRG------GGGMGSGR 93
Score = 43.5 bits (101), Expect = 4e-05
Identities = 31/80 (38%), Positives = 32/80 (39%), Gaps = 6/80 (7%)
Frame = -1
Query: 77 GGYGSGGAKTTTGGGYGGG------YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGY 130
GGYG GG GGG GG YG G G G + GGGYGGG
Sbjct: 305 GGYGGGGGPGYGGGGRGGHGVPQQQYGGPPEYQGRGRGGPSQQGGRGGYSGGGGGYGGGG 126
Query: 131 GDSDTKTTTGGGYGGGYGDS 150
G GGG G G G S
Sbjct: 125 GRG------GGGMGSGRGVS 84
Score = 40.0 bits (92), Expect = 4e-04
Identities = 30/84 (35%), Positives = 31/84 (36%), Gaps = 1/84 (1%)
Frame = -1
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTG-GGYGGGYGDSDTKTTTGGGY 109
G GG G+ Y G G GG G GGY GG GGGY
Sbjct: 287 GGPGYGGGGRGGHGVPQQQYGGPPEYQGRGRGGPSQQGGRGGYSGG----------GGGY 138
Query: 110 GGGYGDSDTKTTTGGGYGGGYGDS 133
GGG G GGG G G G S
Sbjct: 137 GGGGGRG------GGGMGSGRGVS 84
>TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75)
(NGM-75), partial (49%)
Length = 803
Score = 42.7 bits (99), Expect = 6e-05
Identities = 30/80 (37%), Positives = 36/80 (44%), Gaps = 2/80 (2%)
Frame = -1
Query: 69 NYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGG--YGGGYGDSDTKTTTGGGY 126
+Y T GG+ GG+ + G YGGGY GGG YGG Y + + GG Y
Sbjct: 293 SYGGEYTGGGFS*GGSYSGGGFSYGGGY--------AGGGFQYGGRY--TGGGFS*GGSY 144
Query: 127 GGGYGDSDTKTTTGGGYGGG 146
GG D GG Y GG
Sbjct: 143 SGGGWYCDGGFL*GGWYSGG 84
Score = 40.0 bits (92), Expect = 4e-04
Identities = 35/104 (33%), Positives = 46/104 (43%), Gaps = 6/104 (5%)
Frame = -1
Query: 49 SYGNDNSGGDYETG--YNKNTDNYSNNETSGGYGSGGAKT----TTGGGYGGGYGDSDTK 102
SYG + +GG + G Y+ +Y GG+ GG T + GG Y GG D
Sbjct: 293 SYGGEYTGGGFS*GGSYSGGGFSYGGGYAGGGFQYGGRYTGGGFS*GGSYSGGGWYCDGG 114
Query: 103 TTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 146
GG Y GG G S ++GG GG+ + GG Y GG
Sbjct: 113 FL*GGWYSGG-GFSWGGFSSGGFSKGGF-------S*GGWYLGG 6
Score = 36.2 bits (82), Expect = 0.006
Identities = 34/88 (38%), Positives = 39/88 (43%), Gaps = 4/88 (4%)
Frame = -1
Query: 77 GGYGSGGAKTTTGGGYGGGY---GDSDTKTTTGG-GYGGGYGDSDTKTTTGGGYGGGYGD 132
GG+ SGG + GG Y GG G GG YGG Y TGGG+ G
Sbjct: 395 GGWYSGGG-FS*GG*YTGGR**NGG*GVGCPVGGLSYGGEY--------TGGGFS*GGSY 243
Query: 133 SDTKTTTGGGYGGGYGDSDTKTTTGGGY 160
S + GGGY GG G TGGG+
Sbjct: 242 SGGGFSYGGGYAGG-GFQYGGRYTGGGF 162
>CN825770
Length = 586
Score = 42.0 bits (97), Expect = 1e-04
Identities = 31/94 (32%), Positives = 44/94 (45%), Gaps = 17/94 (18%)
Frame = +3
Query: 77 GGYGSGGAKT-TTGGGYGGGYGDSDTKTTTGGG--YGGGYGDSDTKTTTGGGYG------ 127
G YG+ G +T +T G G S+ + + G G G GYGD++ + GYG
Sbjct: 12 GVYGAVGGRTGSTPNSNASGPGGSELQGSGGSGSYMGSGYGDANGNS----GYGNAAWRS 179
Query: 128 ------GGYGDSDTKTTTGG--GYGGGYGDSDTK 153
G YG + GG GYGGGYG + ++
Sbjct: 180 EQAQATGNYGTPQGNGSHGGQVGYGGGYGGAQSR 281
Score = 31.6 bits (70), Expect = 0.14
Identities = 23/66 (34%), Positives = 30/66 (44%), Gaps = 1/66 (1%)
Frame = +3
Query: 30 GYNKTSSY-STGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTT 88
G + SY +G D N + YGN + + + T NY T G GS G +
Sbjct: 93 GSGGSGSYMGSGYGDANGNSGYGN----AAWRSEQAQATGNYG---TPQGNGSHGGQVGY 251
Query: 89 GGGYGG 94
GGGYGG
Sbjct: 252 GGGYGG 269
Score = 30.0 bits (66), Expect = 0.41
Identities = 27/88 (30%), Positives = 36/88 (40%), Gaps = 17/88 (19%)
Frame = +3
Query: 91 GYGGGYGDSDTKT-TTGGGYGGGYGDSDTKTTTGGG--YGGGYGDSDTKTTTG------- 140
G G YG +T +T G G S+ + + G G G GYGD++ + G
Sbjct: 3 GNPGVYGAVGGRTGSTPNSNASGPGGSELQGSGGSGSYMGSGYGDANGNSGYGNAAWRSE 182
Query: 141 -----GGYG--GGYGDSDTKTTTGGGYG 161
G YG G G + GGGYG
Sbjct: 183 QAQATGNYGTPQGNGSHGGQVGYGGGYG 266
Score = 26.6 bits (57), Expect = 4.5
Identities = 20/66 (30%), Positives = 25/66 (37%), Gaps = 5/66 (7%)
Frame = +3
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGG-----AKTTTGGGYGGGYGDSDTKTTT 105
G+ SG +GY N GYG+ A+ T G G G +
Sbjct: 93 GSGGSGSYMGSGYGDANGN-------SGYGNAAWRSEQAQATGNYGTPQGNGSHGGQVGY 251
Query: 106 GGGYGG 111
GGGYGG
Sbjct: 252 GGGYGG 269
>TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (66%)
Length = 1158
Score = 42.0 bits (97), Expect = 1e-04
Identities = 40/111 (36%), Positives = 45/111 (40%), Gaps = 3/111 (2%)
Frame = -2
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG 110
G GGD TG + T GG+ GG T GG GG+ T GG
Sbjct: 401 GGVTIGGDGTTG---GSGT*GGG*TIGGFTIGGVGT*GGG*TIGGFTIGGFGT*GGG*TM 231
Query: 111 GGYGDSDTKTTTGGG---YGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGG 158
GG+G T T GG YGG G + T GG GG G TTGG
Sbjct: 230 GGFGT*GGVTGTTGGLGTYGGVTGTTGGFGT*GGETTGGLGT*GGGFTTGG 78
Score = 38.1 bits (87), Expect = 0.002
Identities = 35/87 (40%), Positives = 37/87 (42%)
Frame = -2
Query: 75 TSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSD 134
T GG G GG T G G GG G T GG GG G T GG GG+G
Sbjct: 422 TIGGVGVGGV-TIGGDGTTGGSGT*GGG*TIGGFTIGGVGT*GGG*TIGGFTIGGFG--- 255
Query: 135 TKTTTGGGYGGGYGDSDTKTTTGGGYG 161
T GG GG+G T T GG G
Sbjct: 254 --T*GGG*TMGGFGT*GGVTGTTGGLG 180
Score = 33.9 bits (76), Expect = 0.028
Identities = 31/84 (36%), Positives = 32/84 (37%), Gaps = 6/84 (7%)
Frame = -2
Query: 81 SGGAKTTTGGGYGGGYGDSDTKTTTGGGYG------GGYGDSDTKTTTGGGYGGGYGDSD 134
+GG TTGGG G G T GG G GG G T G G GG G
Sbjct: 524 AGGGDGTTGGGGGHGVSVGGLGGVTTGGVGVGGVTIGGVGVGGV-TIGGDGTTGGSGT*G 348
Query: 135 TKTTTGGGYGGGYGDSDTKTTTGG 158
T GG GG G T GG
Sbjct: 347 GG*TIGGFTIGGVGT*GGG*TIGG 276
Score = 29.3 bits (64), Expect = 0.70
Identities = 20/46 (43%), Positives = 26/46 (56%), Gaps = 6/46 (13%)
Frame = -2
Query: 75 TSGGYGSGGAKTTTGGGY-GGGYGDSDTKT----TTGG-GYGGGYG 114
T+GG+G+ G +TT G G GGG+ T T GG G GG+G
Sbjct: 158 TTGGFGT*GGETTGGLGT*GGGFTTGGLGT*GGFTIGGLGTWGGFG 21
Score = 28.5 bits (62), Expect = 1.2
Identities = 18/41 (43%), Positives = 22/41 (52%), Gaps = 1/41 (2%)
Frame = -2
Query: 74 ETSGGYGSGGAKTTTGG-GYGGGYGDSDTKTTTGGGYGGGY 113
ET+GG G+ G TTGG G GG+ T G G GG+
Sbjct: 128 ETTGGLGT*GGGFTTGGLGT*GGFTIGGLGTWGGFGT*GGF 6
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 40.4 bits (93), Expect = 3e-04
Identities = 36/93 (38%), Positives = 40/93 (42%), Gaps = 8/93 (8%)
Frame = -2
Query: 77 GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGG----GYGGGY--GDSDTKTTTGGGYGGGY 130
GG +GG GGG GGG G T GG G GG + G + GGG GGG
Sbjct: 809 GGGNNGGKGGKNGGGGGGGGGGVKPGITIGGKGNKGEGGCFDGGGWNGLGINGGGLGGG* 630
Query: 131 GDSDTKTTTGGGYGG--GYGDSDTKTTTGGGYG 161
+GGG GG G G S GGG G
Sbjct: 629 NGLG---ISGGGLGG*NGLGKS------GGGLG 558
Score = 35.8 bits (81), Expect = 0.007
Identities = 36/103 (34%), Positives = 40/103 (37%), Gaps = 13/103 (12%)
Frame = -2
Query: 72 NNETSGGYGSGGAKTTTGGGYGGG------YGDSDTKTTTGGGYGGGY-GDSDTKTTTGG 124
NN GG GG GGG GGG G K G GGG+ G GG
Sbjct: 800 NNGGKGGKNGGG-----GGGGGGGVKPGITIGGKGNKGEGGCFDGGGWNGLGINGGGLGG 636
Query: 125 G------YGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
G GGG G + +GGG GG G +GGG G
Sbjct: 635 G*NGLGISGGGLGG*NGLGKSGGGLGGWNGSG----KSGGGLG 519
>TC10403 similar to UP|O80815 (O80815) T8F5.22 protein, partial (15%)
Length = 1392
Score = 40.0 bits (92), Expect = 4e-04
Identities = 41/153 (26%), Positives = 57/153 (36%), Gaps = 21/153 (13%)
Frame = +3
Query: 28 DSGYNKTSSYSTGDDDYNKKTSYGNDNSG--------GDYETGYNKNTDNYSNNETS-GG 78
D + T TG DY + SG G Y+ + + SNNE G
Sbjct: 723 DRDRSSTPGSRTGRADYRNNGNRDEHPSGVPRPYGGRGRGRGSYSSSRGHNSNNERQDSG 902
Query: 79 YGSG--GAKTTTGGGYGGGYGDSDTKTTTG-----GGYGGGYGDS----DTKTTTGGGYG 127
YG+G G + G + + + + G GG+GG G S T GG+G
Sbjct: 903 YGAGRWGPASKDGNDGLSNFPGAKVQNSPGREAFPGGWGGSDGGSGWGGGASTGDNGGWG 1082
Query: 128 -GGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGG 159
G GD + G+G G + TGGG
Sbjct: 1083QGSGGDQNDAQEGSSGWGTGTKKAGEDGWTGGG 1181
Score = 33.9 bits (76), Expect = 0.028
Identities = 31/121 (25%), Positives = 47/121 (38%), Gaps = 2/121 (1%)
Frame = +3
Query: 7 RSRGLFHHKKHDDDDNRPIDTDSGYNKTSSYST-GDDDYNKKTSYGNDNSGGDYETGYNK 65
R RG + + + +N D+ G + S G+D + NS G
Sbjct: 837 RGRGSYSSSRGHNSNNERQDSGYGAGRWGPASKDGNDGLSNFPGAKVQNS-----PGREA 1001
Query: 66 NTDNYSNNETSGGYGSGGAKTTTGGGYG-GGYGDSDTKTTTGGGYGGGYGDSDTKTTTGG 124
+ ++ G+G GGA T GG+G G GD + G+G G + TGG
Sbjct: 1002FPGGWGGSDGGSGWG-GGASTGDNGGWGQGSGGDQNDAQEGSSGWGTGTKKAGEDGWTGG 1178
Query: 125 G 125
G
Sbjct: 1179G 1181
Score = 28.1 bits (61), Expect = 1.6
Identities = 22/95 (23%), Positives = 33/95 (34%), Gaps = 1/95 (1%)
Frame = +3
Query: 71 SNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG-GGYGDSDTKTTTGGGYGGG 129
+ + GSG + + GG+ G D D +T G G Y ++ + G
Sbjct: 642 AGGSSGASVGSGWGGSNSEGGWKGYSYDRDRSSTPGSRTGRADYRNNGNRDEHPSGVPRP 821
Query: 130 YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDR 164
YG + G Y G + GYG R
Sbjct: 822 YGG---RGRGRGSYSSSRGHNSNNERQDSGYGAGR 917
>TC8694 similar to UP|IF38_MEDTR (Q9XHM1) Eukaryotic translation initiation
factor 3 subunit 8 (eIF3 p110) (eIF3c), partial (23%)
Length = 986
Score = 40.0 bits (92), Expect = 4e-04
Identities = 30/111 (27%), Positives = 42/111 (37%)
Frame = +2
Query: 43 DYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTK 102
DY + G +GG ++ + + GYGSGG + GGY ++
Sbjct: 467 DYAAAAAGGGGGAGGRWQ---DLSLSQPRQGSGRAGYGSGGGRPFAFNQAAGGY----SR 625
Query: 103 TTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTK 153
TG G GGG GGY YG S G G +GD ++
Sbjct: 626 DRTGRGGGGG-----------GGYQSQYGGSGRAHQGGSALRGHHGDGSSR 745
Score = 37.4 bits (85), Expect = 0.003
Identities = 24/70 (34%), Positives = 27/70 (38%)
Frame = +2
Query: 78 GYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKT 137
G A GGG GG + D G GYG + GGY S +T
Sbjct: 461 GQDYAAAAAGGGGGAGGRWQDLSLSQPRQGSGRAGYGSGGGRPFAFNQAAGGY--SRDRT 634
Query: 138 TTGGGYGGGY 147
GGG GGGY
Sbjct: 635 GRGGGGGGGY 664
Score = 30.4 bits (67), Expect = 0.31
Identities = 18/56 (32%), Positives = 20/56 (35%)
Frame = +2
Query: 106 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
GGG GG + D G GYG + GGY T GGG G
Sbjct: 494 GGGAGGRWQDLSLSQPRQGSGRAGYGSGGGRPFAFNQAAGGYSRDRTGRGGGGGGG 661
Score = 28.9 bits (63), Expect = 0.91
Identities = 23/76 (30%), Positives = 25/76 (32%), Gaps = 4/76 (5%)
Frame = +2
Query: 93 GGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGG----GYGGGYG 148
GGG D + G Y GGG GG + D G GYG G G
Sbjct: 422 GGGGLDLPPRRRDGQDYAAAAAGG------GGGAGGRWQDLSLSQPRQGSGRAGYGSGGG 583
Query: 149 DSDTKTTTGGGYGDDR 164
GGY DR
Sbjct: 584 RPFAFNQAAGGYSRDR 631
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.303 0.131 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,267,059
Number of Sequences: 28460
Number of extensions: 69961
Number of successful extensions: 1963
Number of sequences better than 10.0: 198
Number of HSP's better than 10.0 without gapping: 593
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1285
length of query: 257
length of database: 4,897,600
effective HSP length: 88
effective length of query: 169
effective length of database: 2,393,120
effective search space: 404437280
effective search space used: 404437280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC125473.10


