

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123572.11 + phase: 0
(475 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
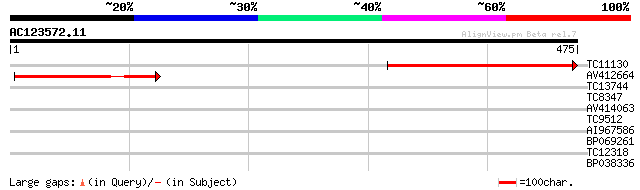
TC11130 weakly similar to UP|Q9LU81 (Q9LU81) Genomic DNA, chromo... 268 1e-72
AV412664 150 3e-37
TC13744 weakly similar to UP|Q39126 (Q39126) HYP1 protein (T4P13... 35 0.035
TC8347 UP|ATPH_LOTJA (P12231) ATP synthase C chain (Lipid-bindi... 32 0.23
AV414063 30 0.66
TC9512 weakly similar to UP|Q9MAB9 (Q9MAB9) T4P13.27 protein, pa... 27 5.6
AI967586 27 7.3
BP069261 27 7.3
TC12318 27 9.5
BP038336 27 9.5
>TC11130 weakly similar to UP|Q9LU81 (Q9LU81) Genomic DNA, chromosome 3, P1
clone: MPE11, partial (44%)
Length = 590
Score = 268 bits (685), Expect = 1e-72
Identities = 130/159 (81%), Positives = 141/159 (87%)
Frame = +2
Query: 317 RQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKYMVAGAAMEINIS 376
RQSFM FADSCLAGESV FFDEVHELSKI E DCVRRIYMARHIIEKY+V GA ME+NIS
Sbjct: 2 RQSFMAFADSCLAGESVFFFDEVHELSKIPEDDCVRRIYMARHIIEKYIVPGAVMEVNIS 181
Query: 377 HRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAKDYWSSMFFLKFQEECDMRCNGYEL 436
HRS+QEI+STS+LAR DLF NALNEI+ LMKTNL DYWSSMFFLKFQEE +MR N YEL
Sbjct: 182 HRSRQEIMSTSNLARPDLFRNALNEIMQLMKTNLVSDYWSSMFFLKFQEESNMRLNDYEL 361
Query: 437 EQMTGWNYSPRLSSVHGTDDPFHHDHLLKNSECGNDTDS 475
EQMT N+SPRLSSVHG+DDPFH +HLLK+S CGNDTDS
Sbjct: 362 EQMTVCNFSPRLSSVHGSDDPFHQEHLLKSSGCGNDTDS 478
>AV412664
Length = 394
Score = 150 bits (380), Expect = 3e-37
Identities = 77/122 (63%), Positives = 93/122 (76%)
Frame = +2
Query: 5 KCAVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVASFNLL 64
+CAV+GGCPTDYVAV +SI+SFI+LL+W+IFPF++H VPRTKGSGFWIPVIQVVASF LL
Sbjct: 59 RCAVEGGCPTDYVAVALSIVSFIVLLLWAIFPFLIHNVPRTKGSGFWIPVIQVVASFILL 238
Query: 65 LSIMTISSNLKRAIGGSHVTCGQVSFHNHTLCLFPVWGEGPLGFGLLLSSRITQAFQLYF 124
LS++ +S K ++C ++ VWGEG LGFGLLLS RITQA QLYF
Sbjct: 239 LSLVLSNSFFKLEKRHWWLSC----------YIWAVWGEGLLGFGLLLSCRITQAAQLYF 388
Query: 125 IF 126
IF
Sbjct: 389 IF 394
>TC13744 weakly similar to UP|Q39126 (Q39126) HYP1 protein (T4P13.21
protein), partial (26%)
Length = 744
Score = 34.7 bits (78), Expect = 0.035
Identities = 25/92 (27%), Positives = 44/92 (47%)
Frame = -3
Query: 182 VATLVGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTAYILNEIHDNISWLQVVSRF 241
VAT + + AA++ + EL WRG+ + + + W+ SW+ V
Sbjct: 487 VATSINLQAAIYLVMN*TSELCK*WRGVTIRPLGILDWIQ-----------SWIIVCISH 341
Query: 242 LLLVLASILVLAFFSISSSQPLLSQISLRRRE 273
L++ L I+++ S+ +Q LS+I L RE
Sbjct: 340 LIIKLRHIMIIWVSSVLLNQ-TLSRIVLEYRE 248
>TC8347 UP|ATPH_LOTJA (P12231) ATP synthase C chain (Lipid-binding
protein) (Subunit III) , complete
Length = 1267
Score = 32.0 bits (71), Expect = 0.23
Identities = 17/51 (33%), Positives = 28/51 (54%)
Frame = +1
Query: 106 LGFGLLLSSRITQAFQLYFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHI 156
L LL+ I F L+++ R L I+ FL +PL+LL W +G ++ +
Sbjct: 409 LSHNLLVDYIINHFFSLFYLENTRNLS*IQ*FLPLPLLLLGWPLGLLLLDL 561
>AV414063
Length = 430
Score = 30.4 bits (67), Expect = 0.66
Identities = 23/68 (33%), Positives = 30/68 (43%), Gaps = 8/68 (11%)
Frame = +3
Query: 46 KGSGFWIPVIQVVAS--FNLLLSIMTISSNLKRAIGGSHV------TCGQVSFHNHTLCL 97
K GFW + ++AS F L L++ S K + G SH+ T VS N C
Sbjct: 201 KCHGFWHNAVLIIASFLFVLYLALQAKQSFRKLSHGRSHIIISYYATLWLVSILNLAWCS 380
Query: 98 FPVWGEGP 105
F VW P
Sbjct: 381 FQVWECSP 404
>TC9512 weakly similar to UP|Q9MAB9 (Q9MAB9) T4P13.27 protein, partial
(26%)
Length = 666
Score = 27.3 bits (59), Expect = 5.6
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 7/51 (13%)
Frame = -2
Query: 2 ANFKCAVKGGCPTDYVAVTVSILSF-------ILLLIWSIFPFIVHKVPRT 45
A F +V GGC T + VTVS S +LL+ IFP +++ + T
Sbjct: 176 AFFSFSVVGGCTTTCIVVTVSGASHNGLWHSPTVLLLSKIFPALINFISFT 24
>AI967586
Length = 405
Score = 26.9 bits (58), Expect = 7.3
Identities = 16/40 (40%), Positives = 23/40 (57%), Gaps = 5/40 (12%)
Frame = -2
Query: 54 VIQVVASF-----NLLLSIMTISSNLKRAIGGSHVTCGQV 88
+I++V SF +L L I+ S AIGG+ V CGQ+
Sbjct: 395 IIEIVGSFMMHVGSLQLRILQCPSIPIHAIGGN*VVCGQI 276
>BP069261
Length = 384
Score = 26.9 bits (58), Expect = 7.3
Identities = 24/97 (24%), Positives = 38/97 (38%)
Frame = -3
Query: 314 KKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKYMVAGAAMEI 373
KK F A S S FF ++S++S H R+ + H +E A
Sbjct: 373 KKSNSQFSMLASSLHFPCSQPFFPLFPKVSRLSYHFRCHRLRLVFHCVEPQNALSAHPFT 194
Query: 374 NISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNL 410
I+H + + L+ DLF EI++ K +
Sbjct: 193 PITHEKR*S*SAYCSLSFVDLFE*NRIEIIYY*KVEI 83
>TC12318
Length = 576
Score = 26.6 bits (57), Expect = 9.5
Identities = 16/50 (32%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Frame = +3
Query: 126 FVKRRLPL-IRSFLLIPLILLPWIIGAAVIHIKKPLSNRCHMSVQWTIPV 174
F +R++ L +RSFLL+ + +LP I A + N H ++ TI +
Sbjct: 66 FKERKMALFVRSFLLLCMFMLPSICNGAFHKTHVKVINSLHGNLDLTIHI 215
>BP038336
Length = 335
Score = 26.6 bits (57), Expect = 9.5
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = +2
Query: 23 ILSFILLLIWSIFPFIVHKVPRTKG 47
I++FILLLIW F + +P T G
Sbjct: 212 IVNFILLLIWLGFKYFCDVLPWTNG 286
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.140 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,183,434
Number of Sequences: 28460
Number of extensions: 165075
Number of successful extensions: 1113
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1107
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1112
length of query: 475
length of database: 4,897,600
effective HSP length: 94
effective length of query: 381
effective length of database: 2,222,360
effective search space: 846719160
effective search space used: 846719160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC123572.11


