

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
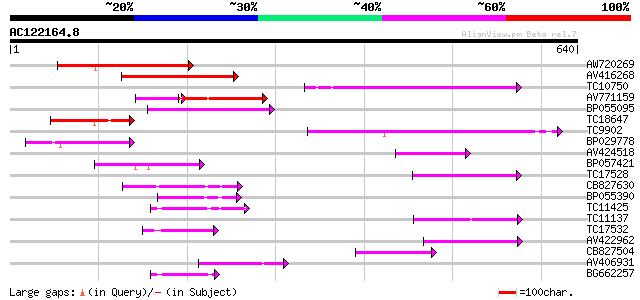
Query= AC122164.8 + phase: 0 /pseudo
(640 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW720269 161 3e-40
AV416268 106 1e-23
TC10750 similar to UP|AAP80385 (AAP80385) ABC transporter, parti... 100 6e-22
AV771159 74 8e-22
BP055095 90 1e-18
TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partia... 72 4e-13
TC9902 similar to UP|BAD07483 (BAD07483) PDR-type ABC transporte... 69 2e-12
BP029778 69 3e-12
AV424518 66 2e-11
BP057421 64 1e-10
TC17528 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Ar... 60 1e-09
CB827630 54 1e-07
BP055390 52 4e-07
TC11425 similar to UP|Q9M1Q9 (Q9M1Q9) P-glycoprotein-like proeti... 50 9e-07
TC11137 similar to GB|CAD59563.1|27368811|OSA535041 PDR-like ABC... 49 2e-06
TC17532 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like prote... 49 3e-06
AV422962 47 7e-06
CB827504 46 2e-05
AV406931 46 2e-05
BG662257 46 2e-05
>AW720269
Length = 524
Score = 161 bits (407), Expect = 3e-40
Identities = 85/158 (53%), Positives = 121/158 (75%), Gaps = 5/158 (3%)
Frame = +3
Query: 55 HVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQK---GSVLLNQKPVDK-SQF 110
++LK+VSF AR EI+A+VGPSG GKS+LL I+AG+ + + ++ +N P+ +Q
Sbjct: 51 NILKSVSFVARSSEIVAVVGPSGTGKSTLLRIIAGRVKDRDFDPKTISINDHPMTSPAQL 230
Query: 111 RKLSGYVTQKDTLFPLLTVEETMMFSAKLRLK-LPQQQQCSRVKSLIKELGLDHVAGTRI 169
RK+ G+V Q+D L PLLTV+ET++FSAK RLK + + RV+SL++ELGL HV+ + +
Sbjct: 231 RKICGFVAQEDNLLPLLTVKETLLFSAKFRLKEMTPNDREMRVESLMQELGLFHVSDSFV 410
Query: 170 GDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGL 207
GD+ RGISGGER+RVSIGV++IH+P +L+LDEPTSGL
Sbjct: 411 GDEENRGISGGERKRVSIGVDMIHNPPILLLDEPTSGL 524
>AV416268
Length = 424
Score = 106 bits (265), Expect = 1e-23
Identities = 58/136 (42%), Positives = 88/136 (64%), Gaps = 4/136 (2%)
Frame = +3
Query: 127 LTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVRGISGGERRR 184
LTV E + +SA L+L + + ++ R IKE+GL TRIG +G+SGG++RR
Sbjct: 12 LTVGEAVYYSAHLQLPDSMSKSEKRERADFTIKEMGLQDAINTRIGGWGSKGVSGGQKRR 191
Query: 185 VSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRG--RTIILSIHQPGFRIV 242
VSI +E++ P++L LDEPTSGLDS ++ +I + + + G RTI+ SIHQP I
Sbjct: 192 VSICIEILTHPRLLFLDEPTSGLDSAASYHVISRISSLNKKDGIQRTIVASIHQPSNEIF 371
Query: 243 KLFNSLLLLANGSVLH 258
+LF+SL LL++G ++
Sbjct: 372 QLFHSLCLLSSGKTVY 419
>TC10750 similar to UP|AAP80385 (AAP80385) ABC transporter, partial (38%)
Length = 807
Score = 100 bits (250), Expect = 6e-22
Identities = 66/247 (26%), Positives = 126/247 (50%), Gaps = 2/247 (0%)
Frame = +1
Query: 333 TLQQLFQQSKVIDEDIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRT 392
T QQ + + +DE I GT ++ A S L ++ LT R N+ R + R
Sbjct: 49 TSQQSYAARQKVDE-ISQFKGTVLEAGGSEA-SFLMQSYTLTKRSFINMSRDFGYYWLRL 222
Query: 393 IQMLISGLVLGSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKET 452
+ ++ + +G+I+ N+ R +F+ F+ SI P F+++ ++ +E
Sbjct: 223 VIYIVVTICIGTIYLNVGTGYNSILARGSCASFVFGFVTFMSIGGFPSFVEDMKVFQRER 402
Query: 453 SCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTA 512
G Y V+++ ++N + PFL+++ L Y++V L+ F+ ++ F+L ++ +
Sbjct: 403 LNGHYGVTAFVVSNTVSATPFLVLITFLSGTICYFMVRLHPGFSHYVFFVLCLYASVTVV 582
Query: 513 NSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPS-YWIF-MHYISLFKYPF 570
S+++ +++VPNF++G + G+ G F L SGYF H+IP W + M YIS +
Sbjct: 583 ESLMMAIASIVPNFLMGIIIGAGIQGIFMLVSGYFRLPHDIPKPVWRYPMSYISFHFWAL 762
Query: 571 EGFLINE 577
+G N+
Sbjct: 763 QGQYQND 783
>AV771159
Length = 486
Score = 73.6 bits (179), Expect(2) = 8e-22
Identities = 39/101 (38%), Positives = 64/101 (62%)
Frame = -3
Query: 191 VIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIHQPGFRIVKLFNSLLL 250
+IH P +L+LDEPTSGLDST+AL+I++ + +A T GRT++ +IHQP R+ +F+ ++L
Sbjct: 337 LIH-PSLLLLDEPTSGLDSTTALRILNTIPRLA-TGGRTVVTTIHQPSSRLYYMFDKVVL 164
Query: 251 LANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSID 291
L+ G +++G A +G + VN + +D D
Sbjct: 163 LSEGCPIYYGPASTALEYFSSVGFSTCVTVNPADLLLDLAD 41
Score = 47.4 bits (111), Expect(2) = 8e-22
Identities = 24/57 (42%), Positives = 34/57 (59%)
Frame = -2
Query: 143 LPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLI 199
L + ++ V+ +I ELGL + IG +RGISG E+RRVSIG E H K ++
Sbjct: 485 LTRDEKVQHVERVITELGLSGCRSSMIGGPLLRGISGAEKRRVSIGQETAHPSKPVV 315
>BP055095
Length = 538
Score = 90.1 bits (222), Expect = 1e-18
Identities = 46/144 (31%), Positives = 79/144 (53%)
Frame = +3
Query: 156 IKELGLDHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQI 215
+K LGLD A IGD+ RGISGG+++RV+ G ++ K L +DE ++GLDS++ QI
Sbjct: 39 LKVLGLDICADVMIGDEMRRGISGGQKKRVTTGEMLVGPAKALFMDEISTGLDSSTTFQI 218
Query: 216 IDMLKVMAETRGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLE 275
++ M T+++S+ QP +LF+ ++LL+ G +++ G + + MG +
Sbjct: 219 CKFMRQMVHIMDVTMVISLLQPAPETFELFDDIILLSEGQIVYQGPRENVLEFFEYMGFK 398
Query: 276 LPLHVNVVEFAIDSIDVIQQQQQW 299
P +F + Q+Q W
Sbjct: 399 CPERKGAADFLQEVTSKKDQEQYW 470
>TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partial (17%)
Length = 430
Score = 71.6 bits (174), Expect = 4e-13
Identities = 41/100 (41%), Positives = 61/100 (61%), Gaps = 5/100 (5%)
Frame = +1
Query: 47 EKGCSGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQ-----KGSVLLN 101
+K VR +LKNVS +A+P +LAI+GPSG+GK++LL +LAG+ G + N
Sbjct: 79 DKSSKSVRFLLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLAASPRLHLSGLLEFN 258
Query: 102 QKPVDKSQFRKLSGYVTQKDTLFPLLTVEETMMFSAKLRL 141
KP K+ ++ YV Q+D F LTV ET+ + +L+L
Sbjct: 259 GKPGSKNAYK--FAYVRQEDLFFSQLTVRETLSLATELQL 372
>TC9902 similar to UP|BAD07483 (BAD07483) PDR-type ABC transporter 1,
partial (19%)
Length = 1207
Score = 69.3 bits (168), Expect = 2e-12
Identities = 60/294 (20%), Positives = 124/294 (41%), Gaps = 6/294 (2%)
Frame = +1
Query: 337 LFQQSKVIDEDIINKTGTGMD--FSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQ 394
LF+++K + +++ D F+ F+ L + + + +R A R
Sbjct: 13 LFRRNKQLIQELGEPAPDSKDLYFATQFSQPFLIQCQACLWKQRWSYWRNPPYTAVRFFF 192
Query: 395 MLISGLVLGSIFCNLKDDLRGTQERVG----LFAFILTFLLSSSIEALPIFLQEREILMK 450
++ G+IF +L + Q+ + +++ +L + +S P+ ER + +
Sbjct: 193 TTFIAVMFGTIFWDLGGKHKRRQDLLNAVGSMYSAVLFLGVQNSSSVQPVVSVERTVFYR 372
Query: 451 ETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLY 510
E + G Y YA A LV LP++ A+ + V +Y ++G + F +L ++ L
Sbjct: 373 EKAAGMYSALPYAFAQILVELPYIFFQAVTYGVIVYAMIGFDWTAEKFFWYLFFMYFTLL 552
Query: 511 TANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPF 570
+ A+ PN V + V + LFSG+ + IP +W + ++ +
Sbjct: 553 YFTFYGMMGVAVTPNHHVASIVAAAFYAIWNLFSGFVVPRPSIPVWWRWYYWACPVAWTI 732
Query: 571 EGFLINEFSNSKKCLEYMFGACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMV 624
G + ++F + ++ G V + E+ YG K+ + VC ++V
Sbjct: 733 YGLIASQFGDITTVMDTEGGKTV----KMFLEDYYG-----IKHSFIGVCAVVV 867
>BP029778
Length = 438
Score = 68.6 bits (166), Expect = 3e-12
Identities = 47/131 (35%), Positives = 75/131 (56%), Gaps = 7/131 (5%)
Frame = -2
Query: 18 HTNKAEH-PFKIFSKSPQLVNTNVQETEEVEKGCSGVRH----VLKNVSFQARPWEILAI 72
H NK PF+ F+ + VN V +E+ K G+ +L NVS P + A+
Sbjct: 401 HKNKGMILPFQQFTVTFHNVNYYVDMPQEIRK--QGIIETKLKLLSNVSGVFAPGVLTAL 228
Query: 73 VGPSGAGKSSLLEILAGKHRPQ--KGSVLLNQKPVDKSQFRKLSGYVTQKDTLFPLLTVE 130
+G SGAGK++L+++LAG+ +G + ++ P + F +++GYV Q D P +TVE
Sbjct: 227 MGSSGAGKTTLMDVLAGRKTGGYIEGDIKISGYPKVQHTFARVAGYVEQNDIHSPQVTVE 48
Query: 131 ETMMFSAKLRL 141
E+++FSA LRL
Sbjct: 47 ESLLFSALLRL 15
>AV424518
Length = 261
Score = 66.2 bits (160), Expect = 2e-11
Identities = 32/85 (37%), Positives = 49/85 (57%)
Frame = +1
Query: 436 EALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNF 495
EA+P+FLQER I M+ET+ +YR SSY +++ ++ LP L+ L+ F +W VGL
Sbjct: 7 EAIPVFLQERYIFMRETAYNAYRRSSYVLSHSVIALPALIFLSFAFAATTFWSVGLAGGT 186
Query: 496 TAFLHFLLLIWLVLYTANSVVVCFS 520
+ F + L I + +S V S
Sbjct: 187 SGFFFYFLTIVASFWAGSSFVTFLS 261
>BP057421
Length = 547
Score = 63.5 bits (153), Expect = 1e-10
Identities = 43/158 (27%), Positives = 76/158 (47%), Gaps = 33/158 (20%)
Frame = +3
Query: 96 GSVLLNQKPVDKSQFRKLSGYVTQKDTLFPLLTVEETMMFSAKLR--------------- 140
G + N +++ RK + Y++Q D +TV+ET+ FSA+ +
Sbjct: 51 GEITYNGHKLNEFVPRKTAAYISQNDVHVGEMTVKETLDFSARCQGVGTRYDXLSELGRR 230
Query: 141 ---------LKLPQQQQCSRVKSL---------IKELGLDHVAGTRIGDDRVRGISGGER 182
+L + + +K +K LGLD T +GD+ RG+SGG++
Sbjct: 231 EKEAGIFPEAELDLFMKATALKGTESSLITDYTLKILGLDICKDTIVGDEMHRGVSGGQK 410
Query: 183 RRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLK 220
+RV+ G ++ K L +DE ++GLDS++ QI+ L+
Sbjct: 411 KRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQ 524
>TC17528 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Arabidopsis
thaliana;}, partial (20%)
Length = 442
Score = 60.1 bits (144), Expect = 1e-09
Identities = 37/125 (29%), Positives = 66/125 (52%), Gaps = 2/125 (1%)
Frame = +1
Query: 455 GSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANS 514
G Y V++Y +AN L PFL+ +A+ + Y +V + F+ F L I+ + S
Sbjct: 16 GYYGVAAYILANFLSSFPFLVAIALTTSTITYNMVKFRPGLSHFVFFTLNIYSSISVIES 195
Query: 515 VVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPS-YWIF-MHYISLFKYPFEG 572
+++ ++LVPNF++G G+IG + SG+F ++P W + + YIS + +G
Sbjct: 196 LMMVVASLVPNFLMGIITGAGIIGIMMMTSGFFRLLSDLPKPVWRYPISYISYGAWAIQG 375
Query: 573 FLINE 577
N+
Sbjct: 376 SYKND 390
>CB827630
Length = 405
Score = 53.5 bits (127), Expect = 1e-07
Identities = 41/135 (30%), Positives = 68/135 (50%)
Frame = +2
Query: 128 TVEETMMFSAKLRLKLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVRGISGGERRRVSI 187
T E ++F +L+ L V+ +K L L H + D + SGG +RR+ +
Sbjct: 2 TGREHLLFYGRLK-NLKGLVLTQAVEESLKSLNLFHGG---VADKQAGKYSGGMKRRLIV 169
Query: 188 GVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIHQPGFRIVKLFNS 247
+ +I DP+V+ +DEP++GLD S + +++K+ R R IIL+ H L +
Sbjct: 170 AISLIGDPRVVYMDEPSTGLDPASRKSLWNVVKL--AKRDRAIILTTHSME-EAEALCDR 340
Query: 248 LLLLANGSVLHHGTA 262
L + NGS+ G A
Sbjct: 341 LGIXVNGSLQCVGNA 385
>BP055390
Length = 488
Score = 51.6 bits (122), Expect = 4e-07
Identities = 32/97 (32%), Positives = 58/97 (58%), Gaps = 2/97 (2%)
Frame = +3
Query: 167 TRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETR 226
T++G V+ +SGG+++R++I ++ +P +L+LDE TS LD+ S + + L+
Sbjct: 204 TQVGQRGVQ-LSGGQKQRIAIARAILKNPPILLLDEATSALDTESEKLVQEALE--TAMH 374
Query: 227 GRTIILSIHQPGFRIVKLFNS--LLLLANGSVLHHGT 261
GRT+IL H R+ + N+ + ++ NG V+ GT
Sbjct: 375 GRTVILIAH----RLSTVVNADVIAVVENGQVVETGT 473
>TC11425 similar to UP|Q9M1Q9 (Q9M1Q9) P-glycoprotein-like proetin, partial
(9%)
Length = 712
Score = 50.4 bits (119), Expect = 9e-07
Identities = 37/112 (33%), Positives = 61/112 (54%), Gaps = 1/112 (0%)
Frame = +2
Query: 160 GLDHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDML 219
G D V G +R +SGG+++RV+I +I P +L+LDE TS LD+ S + D L
Sbjct: 71 GYDTVVG-----ERGTLLSGGQKQRVAIARAIIKTPNILLLDEATSALDAESERVVQDAL 235
Query: 220 -KVMAETRGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLR 270
KVM RT ++ H+ +K + + +L NG ++ G + L +N++
Sbjct: 236 DKVMV---NRTTVIVAHR--LSTIKNADVITVLKNGVIVEKGRHETL-INIK 373
>TC11137 similar to GB|CAD59563.1|27368811|OSA535041 PDR-like ABC
transporter {Oryza sativa (japonica cultivar-group);},
partial (15%)
Length = 786
Score = 49.3 bits (116), Expect = 2e-06
Identities = 29/123 (23%), Positives = 58/123 (46%), Gaps = 1/123 (0%)
Frame = +1
Query: 457 YRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFL-HFLLLIWLVLYTANSV 515
Y YAIA +P++ +++ +Y +V + F +F + + LY
Sbjct: 253 YAPLPYAIAQVFTEIPYVFAQTTFYSLIVYAMVSFDWKVDKFFWYFFVSFFSFLYFTYYG 432
Query: 516 VVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLI 575
++ S + PN V + G F LFSG+FI +IP +W++ ++I + G ++
Sbjct: 433 MMTVS-ITPNHQVASIFAAAFYGLFNLFSGFFIPRPKIPGWWVWYYWICPVAWTVYGLIV 609
Query: 576 NEF 578
+++
Sbjct: 610 SQY 618
>TC17532 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like protein
(At5g02270), partial (48%)
Length = 476
Score = 48.5 bits (114), Expect = 3e-06
Identities = 32/85 (37%), Positives = 45/85 (52%)
Frame = +2
Query: 151 RVKSLIKELGLDHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDST 210
R LIK LG+D R+ +S G+RRRV I + ++ KVL+LDE T LD
Sbjct: 113 RRAELIKVLGIDL-------SWRLHKVSDGQRRRVQICMGLLKPFKVLLLDEITVDLDVL 271
Query: 211 SALQIIDMLKVMAETRGRTIILSIH 235
+ ++ LK + RG TII + H
Sbjct: 272 ARADLLSFLKKECDERGATIIYATH 346
>AV422962
Length = 382
Score = 47.4 bits (111), Expect = 7e-06
Identities = 25/111 (22%), Positives = 53/111 (47%)
Frame = +3
Query: 468 LVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFI 527
L P ++F LY + L+ F F ++ + + A+++ + A+VP
Sbjct: 12 LAEAPIGAAFPLMFGAVLYPMARLHPTLMRFGKFCGIVTMESFAASAMGLTVGAMVPTTE 191
Query: 528 VGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEF 578
+V ++ F +F GY+++ P + ++ +SL ++ F+G +NEF
Sbjct: 192 AAMAVGPSLMTVFIVFGGYYVNRENTPIIFRWIPSVSLIRWAFQGLCVNEF 344
>CB827504
Length = 536
Score = 46.2 bits (108), Expect = 2e-05
Identities = 31/93 (33%), Positives = 49/93 (52%), Gaps = 2/93 (2%)
Frame = +2
Query: 391 RTIQMLISGLVLGSIFCNL-KDDLRGTQERVGLFAFILTFL-LSSSIEALPIFLQEREIL 448
R Q+L + ++LG ++ + +G Q++ GL FI F A+ F QER +L
Sbjct: 248 RITQVLSTAIILGLLWWQSDSSNPKGLQDQAGLLFFIAVFWGFFPVFTAIFTFPQERAML 427
Query: 449 MKETSCGSYRVSSYAIANGLVYLPFLLILAILF 481
KE + YR+S+Y +A LP L+L +LF
Sbjct: 428 AKERASDMYRLSAYFLARTTSDLPLDLVLPVLF 526
>AV406931
Length = 438
Score = 45.8 bits (107), Expect = 2e-05
Identities = 25/101 (24%), Positives = 48/101 (46%)
Frame = +3
Query: 214 QIIDMLKVMAETRGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMG 273
QI+ L+ T ++S+ QP LF+ ++L+++G V++HG + + MG
Sbjct: 3 QIVSSLRQYVHILNGTAVISLLQPAPETYDLFDDIILISDGQVVYHGPREYVLDFFESMG 182
Query: 274 LELPLHVNVVEFAIDSIDVIQQQQQWQVETETPRRLQGTTQ 314
+ P +F + + + Q+Q+ V + P R TQ
Sbjct: 183 FKCPERKGAADF-LQEVTSKKDQEQYWVRRDEPYRFVTVTQ 302
>BG662257
Length = 336
Score = 45.8 bits (107), Expect = 2e-05
Identities = 30/78 (38%), Positives = 45/78 (57%), Gaps = 1/78 (1%)
Frame = +1
Query: 160 GLDHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDML 219
G D V G DR +SGG+++RV+I ++ PK+L+LDE TS LD+ S + D L
Sbjct: 103 GYDTVVG-----DRGIQLSGGQKQRVAIARAIVKSPKILLLDEATSALDAESEKVVQDAL 267
Query: 220 -KVMAETRGRTIILSIHQ 236
+V + RT I+ H+
Sbjct: 268 DRVRVD---RTTIVVAHR 312
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.140 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,022,135
Number of Sequences: 28460
Number of extensions: 159775
Number of successful extensions: 981
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 971
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 974
length of query: 640
length of database: 4,897,600
effective HSP length: 96
effective length of query: 544
effective length of database: 2,165,440
effective search space: 1177999360
effective search space used: 1177999360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC122164.8


