

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121244.7 - phase: 0
(418 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
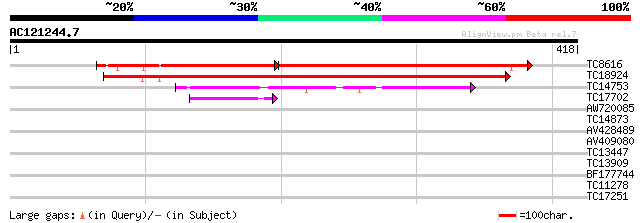
Score E
Sequences producing significant alignments: (bits) Value
TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like pro... 228 e-110
TC18924 homologue to UP|Q94FN3 (Q94FN3) Phosphatidylinositol tra... 358 e-100
TC14753 similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial... 62 1e-10
TC17702 weakly similar to GB|AAN73306.1|25141223|BT002309 At3g51... 48 3e-06
AW720085 39 0.002
TC14873 weakly similar to GB|BAB08982.1|9758453|AB008271 seleniu... 37 0.006
AV428489 37 0.008
AV409080 37 0.008
TC13447 33 0.067
TC13909 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, ... 32 0.19
BF177744 30 0.57
TC11278 similar to UP|NFS1_ARATH (O49543) Cysteine desulfurase, ... 29 1.3
TC17251 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP... 28 2.8
>TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like protein III,
complete
Length = 2302
Score = 228 bits (581), Expect(2) = e-110
Identities = 120/220 (54%), Positives = 144/220 (64%), Gaps = 33/220 (15%)
Frame = +3
Query: 199 IERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGI 258
+ER K+D NKLMQVTT+DRYV+YH Q E+ AIKFPACT+A+KRHIDSS TILD+QG+
Sbjct: 603 LERLGKVDPNKLMQVTTMDRYVRYHVQEFEKSFAIKFPACTMAAKRHIDSSTTILDVQGV 782
Query: 259 GFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVH 318
G N ++ RE++ R K+ DNYP+T Q FIIN G R L + + F+DPK SK+H
Sbjct: 783 GLKNFTKSARELITRLQKVDGDNYPETLCQMFIINAGPGFRLLWNTVKSFLDPKTTSKIH 962
Query: 319 VIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPWNNPEILK---------- 368
V+G++Y K L+VIDASELP LGG CTC +QGGCLRSDKGPW NPEILK
Sbjct: 963 VLGNKYHSKWLEVIDASELPECLGGACTCEDQGGCLRSDKGPWKNPEILKMVLNGEPRRA 1142
Query: 369 -----------------------VKGSDTLTAESSSEAED 385
VKGSDT TAES SEAED
Sbjct: 1143RQVVKVLNSEGKVIAYAKPRYPTVKGSDTSTAESGSEAED 1262
Score = 186 bits (471), Expect(2) = e-110
Identities = 99/162 (61%), Positives = 118/162 (72%), Gaps = 27/162 (16%)
Frame = +2
Query: 65 TGSDENRKERRSDF--------------KKKLLNAFSKFKHSFKKKR------------- 97
+GSDE R ERRSDF KKK L+A +KFKHS +KK
Sbjct: 125 SGSDEKR-ERRSDFENSEDERRTRIGSLKKKALSASTKFKHSLRKKSSRRKSDGRVSSVS 301
Query: 98 VDELQAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLE 157
+++++ V + AV DAFR+ LIMDELLPQ DDYHMMLRFLKARKFDI KAKHMWA+ML+
Sbjct: 302 IEDVRDVEELQAV-DAFRQSLIMDELLPQAFDDYHMMLRFLKARKFDIEKAKHMWAEMLQ 478
Query: 158 WRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFI 199
WRKEFGADTIM+DFEF EL+EV++Y PHG+HGVDKEGRPV+I
Sbjct: 479 WRKEFGADTIMQDFEFQELDEVVRYYPHGHHGVDKEGRPVYI 604
>TC18924 homologue to UP|Q94FN3 (Q94FN3) Phosphatidylinositol transfer-like
protein II, complete
Length = 1976
Score = 358 bits (920), Expect = e-100
Identities = 169/306 (55%), Positives = 227/306 (73%), Gaps = 6/306 (1%)
Frame = +3
Query: 70 NRKERRSDFKKKLLNAFSKFKHSFKKK--RVDELQAVNVVPA----VVDAFRKLLIMDEL 123
++ ++ KKK ++ S ++ +V ++ +V A VD FR+ LI+DEL
Sbjct: 75 DKSDKAGSLKKKAMSLRSSLSRKSRRSSSKVMSVEIEDVRDAEDLKAVDEFRQALILDEL 254
Query: 124 LPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYN 183
LP+KHDDYH +LRFLKARKFD K+K MW+DML+WRKEFGADTI+EDF+F E++EV+KY
Sbjct: 255 LPEKHDDYHQLLRFLKARKFDTEKSKQMWSDMLQWRKEFGADTIVEDFDFKEIDEVVKYY 434
Query: 184 PHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASK 243
PHG+HGVDK+GRPV+IE ++D KLMQVTT+DRY+KYH + E +KF AC+IA+K
Sbjct: 435 PHGHHGVDKDGRPVYIENIGQVDATKLMQVTTMDRYIKYHVKEFERTFDLKFAACSIAAK 614
Query: 244 RHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRS 303
+HID S TILD+QG+G N + RE++ R KI DNYP+T + FIIN G R L S
Sbjct: 615 KHIDQSTTILDVQGVGLKNFNKHARELITRLQKIDGDNYPETLNRMFIINAGSGFRMLWS 794
Query: 304 ICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPWNN 363
+ F+DPK SK+HV+G++YQ KLL+VIDAS+LP FLGGTCTCA+QGGC+RSDKGPW +
Sbjct: 795 TVKSFLDPKTTSKIHVLGNKYQSKLLEVIDASQLPEFLGGTCTCADQGGCMRSDKGPWKD 974
Query: 364 PEILKV 369
PE++++
Sbjct: 975 PELVRM 992
>TC14753 similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial (57%)
Length = 1472
Score = 62.4 bits (150), Expect = 1e-10
Identities = 61/231 (26%), Positives = 104/231 (44%), Gaps = 10/231 (4%)
Frame = +1
Query: 123 LLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKY 182
LL + D ++L+FL+AR F + A M + + WRKEFG D ++++ +E ++V+
Sbjct: 61 LLADERSDV-VLLKFLRARDFKVKDAFTMIKNTVRWRKEFGIDALVDEDLGSEWDKVVFT 237
Query: 183 NPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTID-----RYVKYHAQRCEE-MHAIKFP 236
H DK G V F + + +L Q T D +++K+ Q E+ + + F
Sbjct: 238 QGH-----DKLGHAVCYNVFGEFEDKELYQKTFSDDEKRQKFIKWRIQVLEKSVRKLDFS 402
Query: 237 ACTIASKRHIDSSITILDLQ---GIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIIN 293
A I + + + DL+ G+G L +A + L++L DNYP+ + IN
Sbjct: 403 PTPTA----ISTIVQVNDLKNSPGLGKWELRQATNQA----LQLLQDNYPEFVAKQVFIN 558
Query: 294 VGLELRSLRSICEYFMDPKVASKVHVIG-DRYQRKLLKVIDASELPTFLGG 343
V + + F+ + SK G + L K I +P GG
Sbjct: 559 VPWWYLAFSRMISPFLTQRTKSKFVFAGPSKSAETLFKYIAPELVPVQYGG 711
>TC17702 weakly similar to GB|AAN73306.1|25141223|BT002309
At3g51670/T18N14_50 {Arabidopsis thaliana;}, partial
(20%)
Length = 574
Score = 48.1 bits (113), Expect = 3e-06
Identities = 28/66 (42%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Frame = +2
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIM-EDFEFNELNEVIKYNPHGYHGVD 191
++L+FL+AR F + A +M L WRKEFGAD I+ E+ F EL V+ D
Sbjct: 374 LLLKFLRARDFRVCDALNMLLKCLAWRKEFGADNIVDEELGFKELEGVVACT----QVYD 541
Query: 192 KEGRPV 197
+EG PV
Sbjct: 542 REGHPV 559
>AW720085
Length = 524
Score = 38.5 bits (88), Expect = 0.002
Identities = 30/99 (30%), Positives = 44/99 (44%), Gaps = 7/99 (7%)
Frame = +1
Query: 134 MLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNE--LNEVIKYNPHGYHGVD 191
++RFLKAR+++ KA M D L WR + D+I+ E V G G
Sbjct: 235 LMRFLKAREWNASKAHKMLVDCLNWRVQSEIDSILSKPIIPEDLYRAVRDSQLIGLSGFS 414
Query: 192 KEGRPVF-----IERFEKLDRNKLMQVTTIDRYVKYHAQ 225
+EG PVF + F+K ++ YV+ H Q
Sbjct: 415 REGLPVFAIGVGLSTFDK---------ASVHYYVQSHIQ 504
>TC14873 weakly similar to GB|BAB08982.1|9758453|AB008271 selenium-binding
protein-like {Arabidopsis thaliana;} , partial (7%)
Length = 763
Score = 37.0 bits (84), Expect = 0.006
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Frame = +2
Query: 275 LKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVI--GDRYQRKLLK-V 331
+ IL+ +YP+ +F+ N ++ +YF+DPK A KV + ++ +L+K +
Sbjct: 62 IHILLYHYPERLAIAFLFNPPSIFQAFYKAVKYFLDPKTAQKVKFVYPNNKDSVELMKSL 241
Query: 332 IDASELPTFLGGTCT 346
D LP+ GG T
Sbjct: 242 FDIDNLPSEFGGKAT 286
>AV428489
Length = 386
Score = 36.6 bits (83), Expect = 0.008
Identities = 20/96 (20%), Positives = 45/96 (46%)
Frame = +1
Query: 245 HIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSI 304
+I + + +LD+ G+ F L + ++ + NYP+ +I+N + +
Sbjct: 1 YIGTCVKVLDMSGLKFSALNQL--RLLTAISTVDDLNYPEKTDTYYIVNAPYVFSACWKV 174
Query: 305 CEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTF 340
+ + + K+ V+ + +LLK++D + LP F
Sbjct: 175 VKPLLQERTRRKIQVLQGSGKEELLKIMDYASLPHF 282
>AV409080
Length = 423
Score = 36.6 bits (83), Expect = 0.008
Identities = 22/62 (35%), Positives = 34/62 (54%), Gaps = 13/62 (20%)
Frame = +2
Query: 67 SDENRKERRSDFKKKLLNAFSKFKHSFKKK-------------RVDELQAVNVVPAVVDA 113
S++ R+ R KKK +NA SKF+HS +KK +++++ V + A VDA
Sbjct: 239 SEDERRTRIGSLKKKAINASSKFRHSLRKKSSRRKNASRSNSVSIEDIRDVKELQA-VDA 415
Query: 114 FR 115
FR
Sbjct: 416 FR 421
>TC13447
Length = 554
Score = 33.5 bits (75), Expect = 0.067
Identities = 21/71 (29%), Positives = 34/71 (47%)
Frame = +2
Query: 129 DDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYH 188
DD M+ FLK RKF + +A ++WR++F + E+ + K H +
Sbjct: 173 DDEDMIQWFLKDRKFSVDEAVSKLTKAIKWRQDFEVSKLTEE-AVKDAARSGKAYVHDF- 346
Query: 189 GVDKEGRPVFI 199
+D GRPV +
Sbjct: 347 -LDINGRPVLV 376
>TC13909 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial
(20%)
Length = 399
Score = 32.0 bits (71), Expect = 0.19
Identities = 22/60 (36%), Positives = 33/60 (54%), Gaps = 6/60 (10%)
Frame = +3
Query: 109 AVVDAFRKLLIMDEL----LPQKHDDYH--MMLRFLKARKFDIGKAKHMWADMLEWRKEF 162
AV D + L + DE +P DD ++L+FL+AR+F + A M + L WRK+F
Sbjct: 213 AVQDLKQLLTVTDEPSIWGVPLLKDDRTDVILLKFLRAREFKVKDALLMINNTLRWRKDF 392
>BF177744
Length = 251
Score = 30.4 bits (67), Expect = 0.57
Identities = 14/22 (63%), Positives = 15/22 (67%)
Frame = +1
Query: 364 PEILKVKGSDTLTAESSSEAED 385
P +KGSDT TAES SE ED
Sbjct: 34 PRCPMIKGSDTSTAESGSETED 99
>TC11278 similar to UP|NFS1_ARATH (O49543) Cysteine desulfurase,
mitochondrial precursor , partial (20%)
Length = 557
Score = 29.3 bits (64), Expect = 1.3
Identities = 14/49 (28%), Positives = 26/49 (52%)
Frame = -3
Query: 90 KHSFKKKRVDELQAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFL 138
K S K K+ + Q + + +++ A+ ++I LP KH YH ++ L
Sbjct: 522 KDSAKHKKTTQNQNLIKLTSIMSAYSTIIIYFR*LPMKHT*YHKVINLL 376
>TC17251 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2C) ,
partial (30%)
Length = 715
Score = 28.1 bits (61), Expect = 2.8
Identities = 15/62 (24%), Positives = 30/62 (48%)
Frame = +3
Query: 162 FGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVK 221
F +D + E F + ++++ +PH HG+ K + ++ K + + IDR V+
Sbjct: 21 FASDGLWEHFTNQDAVDIVQNHPH--HGIAKRLVKMALQEAAKKREMRYSDLNKIDRGVR 194
Query: 222 YH 223
H
Sbjct: 195 RH 200
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.140 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,131,171
Number of Sequences: 28460
Number of extensions: 94232
Number of successful extensions: 613
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 609
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 611
length of query: 418
length of database: 4,897,600
effective HSP length: 93
effective length of query: 325
effective length of database: 2,250,820
effective search space: 731516500
effective search space used: 731516500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC121244.7


