

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149306.10 - phase: 0 /pseudo
(138 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
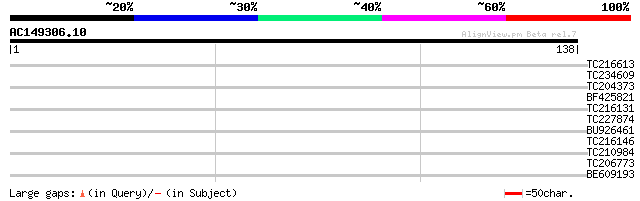
Score E
Sequences producing significant alignments: (bits) Value
TC216613 homologue to UP|Q8VYJ2 (Q8VYJ2) AT4g12080/F16J13_150, p... 28 1.8
TC234609 27 2.4
TC204373 weakly similar to GB|AAM19981.1|20466119|AY098971 AT5g4... 27 4.1
BF425821 26 5.3
TC216131 similar to UP|Q9M3V1 (Q9M3V1) Protein phpsphatase 2C (P... 26 5.3
TC227874 weakly similar to UP|O22670 (O22670) Ag13 protein precu... 26 6.9
BU926461 26 6.9
TC216146 similar to UP|Q7Y256 (Q7Y256) Beta-cyanoalanine synthas... 25 9.0
TC210984 25 9.0
TC206773 similar to UP|CHKA_HUMAN (P35790) Choline kinase alpha ... 25 9.0
BE609193 25 9.0
>TC216613 homologue to UP|Q8VYJ2 (Q8VYJ2) AT4g12080/F16J13_150, partial (44%)
Length = 1790
Score = 27.7 bits (60), Expect = 1.8
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = +2
Query: 88 FLFQGSYMYNLFP*IFFFF 106
F F +++N+FP IFFFF
Sbjct: 122 FKFHSFFLHNIFPSIFFFF 178
>TC234609
Length = 722
Score = 27.3 bits (59), Expect = 2.4
Identities = 13/27 (48%), Positives = 19/27 (70%)
Frame = -3
Query: 102 IFFFFLNIRGVVKFLRKLLGSIGIQYV 128
IFF FL + VK L+KLL +G++Y+
Sbjct: 453 IFFLFLKL*SGVKPLKKLLT*VGLKYL 373
>TC204373 weakly similar to GB|AAM19981.1|20466119|AY098971 AT5g44130/MLN1_5
{Arabidopsis thaliana;} , partial (60%)
Length = 1283
Score = 26.6 bits (57), Expect = 4.1
Identities = 17/52 (32%), Positives = 29/52 (55%)
Frame = +3
Query: 67 IFWWPSGSFEICPILSSGLLSFLFQGSYMYNLFP*IFFFFLNIRGVVKFLRK 118
IF + G E+C +L S +LS Y++NL ++ F N+ ++ F+RK
Sbjct: 870 IFLFFYGQKELCWVLLSLILSI*CPLFYVFNLILVLYLLF-NLDHMILFMRK 1022
>BF425821
Length = 387
Score = 26.2 bits (56), Expect = 5.3
Identities = 13/38 (34%), Positives = 20/38 (52%), Gaps = 4/38 (10%)
Frame = +3
Query: 8 TLWYMGCQTVVFSV----EL*NRYYSVCLFGVAYWWR* 41
TLW + C+ V + +L R S C++ +WWR*
Sbjct: 141 TLWVLQCEWRV*DI**R*KLHARLQSCCIYLCIWWWR* 254
>TC216131 similar to UP|Q9M3V1 (Q9M3V1) Protein phpsphatase 2C (PP2C) ,
partial (47%)
Length = 1320
Score = 26.2 bits (56), Expect = 5.3
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -1
Query: 88 FLFQGSYMYNLFP*IFFFFLNIR 110
F F S + N+ P FFFFLN++
Sbjct: 1188 FFFSFSKVNNVLPLSFFFFLNLK 1120
>TC227874 weakly similar to UP|O22670 (O22670) Ag13 protein precursor,
partial (21%)
Length = 894
Score = 25.8 bits (55), Expect = 6.9
Identities = 19/52 (36%), Positives = 26/52 (49%)
Frame = -1
Query: 79 PILSSGLLSFLFQGSYMYNLFP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYL 130
P + LL+FL +G ++ + F FF FLNI F L IGI + L
Sbjct: 507 PFCLNLLLTFLCRGLFLLS-FSLSFFLFLNINIFCSFWCLNLSRIGIFFCSL 355
>BU926461
Length = 425
Score = 25.8 bits (55), Expect = 6.9
Identities = 8/34 (23%), Positives = 16/34 (46%)
Frame = +3
Query: 67 IFWWPSGSFEICPILSSGLLSFLFQGSYMYNLFP 100
+ WW F++ P++S + F + +FP
Sbjct: 54 LIWWDYPKFQLLPLISIAISCFTHHMEKLIRMFP 155
>TC216146 similar to UP|Q7Y256 (Q7Y256) Beta-cyanoalanine synthase, partial
(96%)
Length = 1431
Score = 25.4 bits (54), Expect = 9.0
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +3
Query: 35 VAYWWR**ET*FLETCC*KNC 55
++ W T*FL++CC KNC
Sbjct: 1290 ISLLWLHGAT*FLQSCCSKNC 1352
>TC210984
Length = 1085
Score = 25.4 bits (54), Expect = 9.0
Identities = 17/36 (47%), Positives = 18/36 (49%), Gaps = 2/36 (5%)
Frame = +2
Query: 77 ICPILSSGL--LSFLFQGSYMYNLFP*IFFFFLNIR 110
I +L GL LSFLF YNLF F F N R
Sbjct: 821 IMTLLLKGLKFLSFLFSLFKHYNLFIIFFIHFCNFR 928
>TC206773 similar to UP|CHKA_HUMAN (P35790) Choline kinase alpha (CK)
(CHETK-alpha) , partial (7%)
Length = 1606
Score = 25.4 bits (54), Expect = 9.0
Identities = 13/37 (35%), Positives = 20/37 (53%), Gaps = 8/37 (21%)
Frame = -2
Query: 107 LNIRGVVKFLRKLLGSIGIQ--------YVYLWIVVV 135
LNI+ FLR LLGS Q ++++W+ V+
Sbjct: 1023 LNIQSFKIFLRHLLGSFRFQVMPEKMILFIHIWVQVI 913
>BE609193
Length = 458
Score = 25.4 bits (54), Expect = 9.0
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 90 FQGSYMYNLFP*IFFFF 106
F+ SY+YNL +FFFF
Sbjct: 235 FKPSYIYNLLHFLFFFF 285
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.368 0.170 0.704
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,221,874
Number of Sequences: 63676
Number of extensions: 132116
Number of successful extensions: 2213
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 2197
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2211
length of query: 138
length of database: 12,639,632
effective HSP length: 87
effective length of query: 51
effective length of database: 7,099,820
effective search space: 362090820
effective search space used: 362090820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC149306.10


