

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147429.6 - phase: 0 /pseudo
(974 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
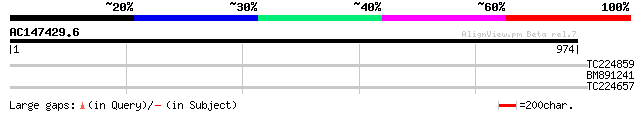
Score E
Sequences producing significant alignments: (bits) Value
TC224859 weakly similar to UP|Q7TP79 (Q7TP79) Aa2-245, partial (3%) 33 0.61
BM891241 33 0.80
TC224657 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich gly... 31 3.0
>TC224859 weakly similar to UP|Q7TP79 (Q7TP79) Aa2-245, partial (3%)
Length = 843
Score = 33.1 bits (74), Expect = 0.61
Identities = 19/41 (46%), Positives = 25/41 (60%), Gaps = 1/41 (2%)
Frame = -2
Query: 272 LCIVSKLLLFMLSLMVNLLR-LFPPERSKARRSPVSLPLHH 311
LC+ + LLL +L L+ +LL L PP SP +LPLHH
Sbjct: 755 LCLCALLLLLLLLLLHSLLLFLLPPPLPFFVASPCALPLHH 633
>BM891241
Length = 407
Score = 32.7 bits (73), Expect = 0.80
Identities = 23/62 (37%), Positives = 32/62 (51%)
Frame = -3
Query: 439 RKFIG*KDLSFLSLSLRGVWISEL*GSSIWPCLLNKFGAFTLNLNLLSLGVSRQSISPIL 498
RKF+G* F LRGVW S++ +SI C N +G L + L + QS+ L
Sbjct: 387 RKFLG*NGRLFACPRLRGVWGSKISPNSI*LCWANGYGP*LLISSSYGLELLIQSMEAGL 208
Query: 499 MF 500
+F
Sbjct: 207 IF 202
>TC224657 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich glycoprotein
(Fragment), partial (50%)
Length = 586
Score = 30.8 bits (68), Expect = 3.0
Identities = 30/111 (27%), Positives = 52/111 (46%), Gaps = 4/111 (3%)
Frame = +1
Query: 684 TVLYFALDAMLK*KTLHIPF*PVLISLRFGLVLAWLLDLQTPLTQTAST--GYMKSYPTR 741
T+ + + +L T IP V I L+ + L++++ST + S+P+
Sbjct: 1 TISFSSSSLLLPLSTTTIPITAVSILLQIPSTSISISTTSLLLSKSSSTISHFSSSFPSS 180
Query: 742 TNLPLFKLHLLSIIFGIQETYACSKTNTYLMLTSS--SVLKDASRTFTVLK 790
+ ++HL I Y C T+TYL ++S+ S + D S FT+LK
Sbjct: 181 CS----QVHLQ--ITSTSGLYLCFPTSTYLQVSSNCESTIHDESCKFTILK 315
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.363 0.165 0.602
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,399,258
Number of Sequences: 63676
Number of extensions: 657724
Number of successful extensions: 6596
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3695
Number of HSP's successfully gapped in prelim test: 240
Number of HSP's that attempted gapping in prelim test: 2822
Number of HSP's gapped (non-prelim): 4113
length of query: 974
length of database: 12,639,632
effective HSP length: 106
effective length of query: 868
effective length of database: 5,889,976
effective search space: 5112499168
effective search space used: 5112499168
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC147429.6


